Abstract
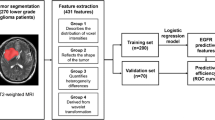
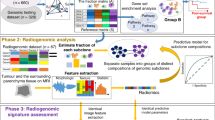
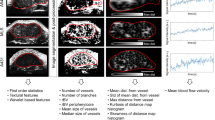
Breast tumors are heterogeneous lesions. Intra-tumor heterogeneity presents a major challenge for cancer diagnosis and treatment. Few studies have worked on capturing tumor heterogeneity from imaging. Most studies to date consider aggregate measures for tumor characterization. In this work we capture tumor heterogeneity by partitioning tumor pixels into subregions and extracting heterogeneity wavelet kinetic (HetWave) features from breast dynamic contrast-enhanced magnetic resonance imaging (DCE-MRI) to obtain the spatiotemporal patterns of the wavelet coefficients and contrast agent uptake from each partition. Using a genetic algorithm for feature selection, and a logistic regression classifier with leave one-out cross validation, we tested our proposed HetWave features for the task of classifying breast cancer recurrence risk. The classifier based on our features gave an ROC AUC of 0.78, outperforming previously proposed kinetic, texture, and spatial enhancement variance features which give AUCs of 0.69, 0.64, and 0.65, respectively.
Chapter PDF
Similar content being viewed by others
Keywords
References
Schnall, M.D., et al.: Diagnostic architectural and dynamic features at breast MR imaging: multicenter study. Radiology 238, 42–53 (2006)
Bhooshan, N., Giger, M.L., Jansen, S.A., Li, H., Lan, L., Newstead, G.M.: Cancerous breast lesions on dynamic contrast-enhanced MR images: computerized characterization for image-based prognostic markers. Radiology 254, 680–690 (2010)
Chen, W., Giger, M.L., Lan, L., Bick, U.: Computerized interpretation of breast MRI: investigation of enhancement-variance dynamics. Med. Phys. 31, 1076–1082 (2004)
Lee, S.H., Kim, J.H., Cho, N., Park, J.S., Yang, Z., Jung, Y.S., Moon, W.K.: Multilevel analysis of spatiotemporal association features for differentiation of tumor enhancement patterns in breast DCE-MRI. Med. Phys. 37, 3940 (2010)
Zheng, Y., Englander, S., Baloch, S., Zacharaki, E.I., Fan, Y., Schnall, M.D., Shen, D.: STEP: spatiotemporal enhancement pattern for MR-based breast tumor diagnosis. Med. Phys. 37(7), 3192–3204 (2009)
Agner, S., Soman, S., Libfeld, E., McDonald, M., Thomas, K., Englander, S., Rosen, M.A., Chin, D., Nosher, J., Madabhushi, A.: Textural kinetics: a novel dynamic contrast-enhanced (DCE)-MRI feature for breast lesion classification. J. Digit. Imaging 24(3), 446–463 (2011)
Marusyk, A., Polyak, K.: Tumor heterogeneity:causes and consequences. Biochim. Biophys. Acta 1805, 105–117 (2010)
Yang, J., Honavar, V.: Feature subset selection using a genetic algorithm. IEEE Intelligent Systems and their Applications 13(2), 44–49 (1998)
Paik, S., Tang, G., Shak, S., Kim, C., Baker, J., Kim, W., Cronin, M., Baehner, F.L., Watson, D., Bryant, J., Constantino, J.P., Geyer, C.E., Wickerham, D.L., Wolmark, N.: Gene expression and benefit of chemotherapy in women with node-negative, estrogen receptor-positive breast cancer. J. Clin. Oncol. 24(23), 3726–3734 (2006)
Gonzalez, R., Woods, R.E.: Digital Image Processing. Prentice Hall (2007)
Hylton, N.: MR imaging for assessment of breast cancer response to neoadjuvant chemotherapy. Magn. Reson. Imaging Clin. N. Am. 14(3), 383–389 (2006)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Mahrooghy, M. et al. (2013). Heterogeneity Wavelet Kinetics from DCE-MRI for Classifying Gene Expression Based Breast Cancer Recurrence Risk. In: Mori, K., Sakuma, I., Sato, Y., Barillot, C., Navab, N. (eds) Medical Image Computing and Computer-Assisted Intervention – MICCAI 2013. MICCAI 2013. Lecture Notes in Computer Science, vol 8150. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-40763-5_37
Download citation
DOI: https://doi.org/10.1007/978-3-642-40763-5_37
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-40762-8
Online ISBN: 978-3-642-40763-5
eBook Packages: Computer ScienceComputer Science (R0)