Abstract
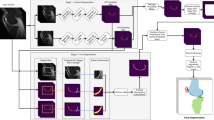
Segmentation of anatomical structures in medical images is often based on a voxel/pixel classification approach. Deep learning systems, such as convolutional neural networks (CNNs), can infer a hierarchical representation of images that fosters categorization. We propose a novel system for voxel classification integrating three 2D CNNs, which have a one-to-one association with the xy, yz and zx planes of 3D image, respectively. We applied our method to the segmentation of tibial cartilage in low field knee MRI scans and tested it on 114 unseen scans. Although our method uses only 2D features at a single scale, it performs better than a state-of-the-art method using 3D multi-scale features. In the latter approach, the features and the classifier have been carefully adapted to the problem at hand. That we were able to get better results by a deep learning architecture that autonomously learns the features from the images is the main insight of this study.
This work has been funded by the Danish Strategic Research Council through the grant Learning Imaging Biomarkers (grant no. 09-065145).
Chapter PDF
Similar content being viewed by others
Keywords
- Articular Cartilage
- Convolutional Neural Network
- Tibial Cartilage
- Convolutional Layer
- Training Data Point
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
References
LeCun, Y., Bottou, L., Bengio, Y., Haffner, P.: Gradient-Based Learning Applied to Document Recognition. Proceedings of the IEEE 86(11), 2278–2324 (1998)
Schulz, H., Behnke, S.: Learning Object-Class Segmentation with Convolutional Neural Networks. In: ESANN (2012)
Turaga, S.C., Murray, J.F., Jain, V., Roth, F., Helmstaedter, M., Briggman, K.L., Denk, W., Seung, H.S.: Convolutional Networks can Learn to Generate Affinity Graphs for Image Segmentation. Neural Computation 22(2), 511–538 (2010)
Jackson, D., Simon, T., Aberman, H.: Symptomatic Articular Cartilage Degeneration: The impact in the New Millenium. Clinical Orthopaedics and Related Research 133, 14–25 (2001)
Folkesson, J., Dam, E., Olsen, O., Pettersen, P., Christiansen, C.: Segmenting Articular Cartilage Automatically Using a Voxel Classification Approach. IEEE TMI 26(1), 106–115 (2007)
Solloway, S., Hutchinson, C., Vaterton, J., Taylor, C.: The Use of Active Shape Models for Making Thickness Measurements of Articular Cartilage from MR Images. Magnetic Resonance in Medicine 37, 943–952 (1997)
Pakin, S.K., Tamez-Pena, J.G., Totterman, S., Parker, K.J.: Segmentation, Surface Extraction and Thickness Computation of Articular Cartilage. In: SPIE Medical Imaging, vol. 4684, pp. 155–166 (2002)
Fripp, J., Crozier, S., Warfield, S., Ourselin, S.: Automatic Segmentation and Quantitative Analysis of the Articular Cartilages from Magnetic Resonance Images of the Knee. IEEE TMI 29(1), 55–64 (2010)
Yin, Y., Xiangmin, Z., Williams, R., Xiaodong, W., Anderson, D., Sonka, M.: LOGISMOS - Layered Optimal Graph Image Segmentation of Multiple Objects and Surfaces: Cartilage Segmentation in the Knee Joint. IEEE TMI 29(12), 2023–2037 (2010)
Ning, F., Delhomme, D., LeCun, Y., Piano, F., Bottou, L., Barbano, P.E.: Toward Automatic Phenotyping of Developing Embryos from Videos. IEEE TIP 14(9), 1360–1371 (2005)
Sermanet, P., LeCun, Y.: Traffic Sign Recognition with Multi-Scale Convolutional Networks. In: IJCNN, pp. 2809–2813 (2011)
Ciresan, D.C., Meier, U., Masci, J., Gambardella, L.M., Schmidhuber, J.: Flexible, High Performance Convolutional Neural Networks for Image Classification. In: IJCAI, 1237–1242 (2011)
Wang, T., Wu, D.J., Coates, A., Ng, A.Y.: End-to-End Text Recognition with Convolutional Neural Networks. In: ICPR, pp. 3304–3308 (2012)
Ji, S., Xu, W., Yang, M., Yu, K.: 3D Convolutional Neural Networks for Human Action Recognition. IEEE TPAMI 35(1), 221–231 (2013)
Le, Q.V., Ngiam, J., Coates, A., Lahiri, A., Prochnow, B., Ng, A.Y.: On Optimization Methods for Deep Learning. In: ICML, 265–272 (2011)
Simard, P.Y., Steinkraus, D., Platt, J.C.: Best Practices for Convolutional Neural Networks Applied to Visual Document Analysis. In: ICDAR, pp. 958–962 (2003)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Prasoon, A., Petersen, K., Igel, C., Lauze, F., Dam, E., Nielsen, M. (2013). Deep Feature Learning for Knee Cartilage Segmentation Using a Triplanar Convolutional Neural Network. In: Mori, K., Sakuma, I., Sato, Y., Barillot, C., Navab, N. (eds) Medical Image Computing and Computer-Assisted Intervention – MICCAI 2013. MICCAI 2013. Lecture Notes in Computer Science, vol 8150. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-40763-5_31
Download citation
DOI: https://doi.org/10.1007/978-3-642-40763-5_31
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-40762-8
Online ISBN: 978-3-642-40763-5
eBook Packages: Computer ScienceComputer Science (R0)