Abstract
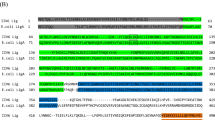
The analysis on codon usage bias of IFN-a gene of giant panda (Ailuropoda melanoleuca) may provide a basis for understanding the evolution relationship of giant panda and for selecting appropriate host expression systems to improve the expression of target genes. In this paper, the codon usage bias in the mature IFN-a sequence of giant panda and 15 reference species have been analyzed. The results showed that the synonymous codons with G and C at the third codon position were widely used and the ENC-GC3S plot revealed that the genetic heterogeneity in IFN-a gene was main constrained by mutational bias. Contrastive analysis revealed that there were 40 codons showing distinct usage differences between GpIFN-a and Escherichia coli, 38 codons between GpIFN-a and yeast. and only 30 between GpIFN-a and Homo sapiens. Therefore the Homo expression system may be more suitable for the expression of GpIFN-a genes.
Chapter PDF
Similar content being viewed by others
References
Gustafsson, C., Govindarajan, S., Minshull, J.: Codon bias and heterologous protein expression. Trends in Biotechnology 22(7), 346–353 (2004)
Hershberg, R., Petrov, D.: Selection on codon bias. Annual Review of Genetics 42, 287–299 (2008)
Ikemura, T.: Codon usage and tRNA content in unicellular and multicellular organisms. Molecular Biology and Evolution 2(1), 13 (1985)
Sharp, P.: Codon usage in regulatory genes in Escherichia coli does not reflect selection for’rare’codons. Nucleic Acids Research 14(19), 7737 (1986)
Sharp, P., Li, W.: An evolutionary perspective on synonymous codon usage in unicellular organisms. Journal of Molecular Evolution 24(1), 28–38 (1986)
Shields, D., Sharp, P., Higgins, D., Wright, F.: " Silent" sites in Drosophila genes are not neutral: evidence of selection among synonymous codons. Molecular Biology and Evolution 5(6), 704 (1988)
Smith, N., Eyre-Walker, A.: Synonymous codon bias is not caused by mutation bias in G+ C-rich genes in humans. Molecular Biology and Evolution 18(6), 982 (2001)
Bains, W.: Codon distribution in vertebrate genes be used to predict gene length. Journal of Molecular Biology 197(3), 379–388 (1987)
D’Onofrio, G., Ghosh, T., Bernardi, G.: The base composition of the genes is correlated with the secondary structures of the encoded proteins. Gene 300(1-2), 179–187 (2002)
Bernardi, G.: Compositional constraints and genome evolution. Journal of Molecular Evolution 24(1), 1–11 (1986)
Gouy, M., Gautier, C.: Codon usage in bacteria: correlation with gene expressivity. Nucleic Acids Research 10(22), 7055 (1982)
Tao, X., Dafu, D.: The relationship between synonymous codon usage and protein structure. FEBS Letters 434(1-2), 93–96 (1998)
Angellotti, M., Bhuiyan, S., Chen, G., Wan, X.: CodonO: codon usage bias analysis within and across genomes. Nucleic Acids Research 35(Web Server issue), W132 (2007)
Epstein, R., Lin, K., Tan, T.: A functional significance for codon third bases. Gene 245(2), 291–298 (2000)
Ma, J., Zhou, T., Gu, W., Sun, X., Lu, Z.: Cluster analysis of the codon use frequency of MHC genes from different species. Biosystems 65(2-3), 199–207 (2002)
Coghlan, A., Wolfe, K.: Relationship of codon bias to mRNA concentration and protein length in Saccharomyces cerevisiae. Yeast-Chichester 16(12), 1131–1146 (2000)
Marais, G., Duret, L.: Synonymous codon usage, accuracy of translation, and gene length in Caenorhabditis elegans. Journal of Molecular Evolution 52(3), 275–280 (2001)
Tao, Y., Zeng, B., Xu, L., Yue, B., Yang, D., Zou, F.: Interferon-γ of the Giant Panda (Ailuropoda melanoleuca): Complementary DNA Cloning, Expression, and Phylogenetic Analysis. DNA and Cell Biology 29(1), 41–45
Xu, L., Zeng, B., Peng, R., Zou, F., Yue, B.: Molecular cloning and sequence analysis of the gene encoding interleukin-6 of the giant panda (Ailuropoda melanoleuca). Journal of Natural History 42(39), 2585–2591 (2008)
Isaacs, A., Lindenmann, J.: “Virus interference: I”. The interferon. CA: A Cancer Journal for Clinicians 38(5), 280 (1988)
Bin, F., Bu-feng, L., Guang-yuan, H.: Synonymous codon usage bias and overexpression of a synthetic gene encoding Interferon α2b in yeast. Virologica Sinica 22(3), 226–232 (2007)
Duret, L., Mouchiroud, D.: Expression pattern and, surprisingly, gene length shape codon usage in Caenorhabditis, Drosophila, and Arabidopsis. Proceedings of the National Academy of Sciences of the United States of America 96(8), 4482 (1999)
Wright, F.: The effective number of codons’ used in a gene. Gene 87(1), 23–29 (1990)
Sakai, H., Washio, T., Saito, R., Shinagawa, A., Itoh, M., Shibata, K., et al.: Correlation between sequence conservation of the 5’untranslated region and codon usage bias in Mus musculus genes. Gene 276(1-2), 101–105 (2001)
Lu, H., Zhao, W., Zheng, Y., Wang, H., Qi, M., Yu, X.: Analysis of synonymous codon usage bias in Chlamydia. Acta Biochimica et Biophysica Sinica 37(1), 1 (2005)
Gu, W., Zhou, T., Ma, J., Sun, X., Lu, Z.: Analysis of synonymous codon usage in SARS Coronavirus and other viruses in the Nidovirales. Virus Research 101(2), 155–161 (2004)
Wu, X., Wu, S., Ren, D., Zhu, Y., He, F.: The analysis method and progress in the study of codon bias. Yi Chuan= Hereditas/Zhongguo Yi Chuan Xue Hui Bian Ji 29(4), 420 (2007)
Tjalsma, H., Bolhuis, A., Jongbloed, J., Bron, S., Van Dijl: Signal peptide-dependent protein transport in Bacillus subtilis: a genome-based survey of the secretome. Microbiology and Molecular Biology Reviews 64(3), 515 (2000)
von Heijne, G.: The signal peptide. Journal of Membrane Biology 115(3), 195–201 (1990)
Zhang, S., Cheng, A., Wang, M.: Characterization of Codon Usage Bias in the Newly Identified DEV UL53 Gene, pp. 1–9. IEEE
Shen, F., Cheng, A., Wang, M., Huang, H., Li, C., Jiang, J., et al.: Factors affecting synonymous codon usage bias in the gC gene of DPV CHv strain. African Journal of Microbiology Research (5), 343–353
Xiang, J., Cheng, A., Wang, M., Zhang, S., Chang, H., Zhu, D., et al.: Analysis of Synonymous Codon Usage in the Capsid Gene UL38 of Duck Enteritis Virus, pp. 1–8. IEEE
Jia, R., Cheng, A., Wang, M., Xin, H., Guo, Y., Zhu, D., et al.: Analysis of synonymous codon usage in the UL24 gene of duck enteritis virus. Virus Genes 38(1), 96–103 (2009)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer-Verlag GmbH Berlin Heidelberg
About this chapter
Cite this chapter
Yi, Y. et al. (2012). Analysis of Codon Usage Bias in Interferon Alpha Gene of the Giant Panda (Ailuropoda Melanoleuca). In: Zhu, E., Sambath, S. (eds) Information Technology and Agricultural Engineering. Advances in Intelligent and Soft Computing, vol 134. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-27537-1_37
Download citation
DOI: https://doi.org/10.1007/978-3-642-27537-1_37
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-27536-4
Online ISBN: 978-3-642-27537-1
eBook Packages: EngineeringEngineering (R0)