Abstract
This paper describes a novel feature selection algorithm embedded into logistic regression. It specifically addresses high dimensional data with few observations, which are commonly found in the biomedical domain such as microarray data. The overall objective is to optimize the predictive performance of a classifier while favoring also sparse and stable models.
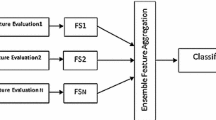
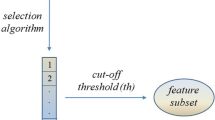
Feature relevance is first estimated according to a simple t-test ranking. This initial feature relevance is treated as a feature sampling probability and a multivariate logistic regression is iteratively reestimated on subsets of randomly and non-uniformly sampled features. At each iteration, the feature sampling probability is adapted according to the predictive performance and the weights of the logistic regression. Globally, the proposed selection method can be seen as an ensemble of logistic regression models voting jointly for the final relevance of features.
Practical experiments reported on several microarray datasets show that the proposed method offers a comparable or better stability and significantly better predictive performances than logistic regression regularized with Elastic Net. It also outperforms a selection based on Random Forests, another popular embedded feature selection from an ensemble of classifiers.
Chapter PDF
Similar content being viewed by others
References
Abeel, T., Helleputte, T., Van de Peer, Y., Dupont, P., Saeys, Y.: Robust biomarker identification for cancer diagnosis with ensemble feature selection methods. Bioinformatics 26, 392–398 (2010)
Bach, F.R.: Bolasso: model consistent lasso estimation through the bootstrap. In: Proceedings of the 25th International Conference on Machine Learning, pp. 33–40. ACM (2008)
Breiman, L.: Random forests. Machine Learning 45, 5–32 (2001)
Chandran, U.R., Ma, C., Dhir, R., Bisceglia, M., Lyons-Weiler, M., Liang, W., Michalopoulos, G., Becich, M., Monzon, F.A.: Gene expression profiles of prostate cancer reveal involvement of multiple molecular pathways in the metastatic process. BMC Cancer 7(1), 64 (2007)
Cox, D.R., Snell, E.J.: Analysis of binary data. Monographs on statistics and applied probability. Chapman and Hall (1989)
Desmedt, C., Piette, F., Loi, S., Wang, Y., Lallemand, F., Haibe-Kains, B., Viale, G., Delorenzi, M., Zhang, Y., D’Assignies, M.S., Bergh, J., Lidereau, R., Ellis, P., Harris, A., Klijn, J., Foekens, J., Cardoso, F., Piccart, M., Buyse, M., Sotiriou, C.: Strong time dependence of the 76-gene prognostic signature for node-negative breast cancer patients in the transbig multicenter independent validation series. Clinical Cancer Research 13(11), 3207–3214 (2007)
Dietterich, T.G.: Ensemble methods in machine learning. In: Kittler, J., Roli, F. (eds.) MCS 2000. LNCS, vol. 1857, pp. 1–15. Springer, Heidelberg (2000)
Guyon, I., Gunn, S., Nikravesh, M., Zadeh, L. (eds.): Feature Extraction. Foundations and Applications. Studies in Fuzziness and Soft Computing. Physica-Verlag, Springer (2006)
Helleputte, T., Dupont, P.: Feature Selection by Transfer Learning with Linear Regularized Models. In: Buntine, W., Grobelnik, M., Mladenić, D., Shawe-Taylor, J. (eds.) ECML PKDD 2009. LNCS, vol. 5781, pp. 533–547. Springer, Heidelberg (2009)
Hoerl, A.E., Kennard, R.W.: Ridge regression: Biased estimation for nonorthogonal problems. Technometrics 12, 55–67 (1970)
Kalousis, A., Prados, J., Hilario, M.: Stability of feature selection algorithms: a study on high-dimensional spaces. Knowledge and Information Systems 12, 95–116 (2007), doi:10.1007/s10115-006-0040-8
Kuncheva, L.I.: A stability index for feature selection. In: Proceedings of the 25th International Multi-Conference Artificial Intelligence and Applications, pp. 390–395. ACTA Press, Anaheim (2007)
Li, Q., Eklund, A.C., Juul, N., Haibe-Kains, B., Workman, C.T., Richardson, A.L., Szallasi, Z., Swanton, C.: Minimising immunohistochemical false negative er classification using a complementary 23 gene expression signature of er status. PLoS ONE 5(12), e15031 (2010)
Nadeau, C., Bengio, Y.: Inference for the generalization error. Machine Learning 52, 239–281 (2003)
Ng, A.Y.: Feature selection, l 1 vs. l 2 regularization, and rotational invariance. In: Proceedings of the Twenty-First International Conference on Machine Learning (ICML), vol. 1, pp. 78–85 (2004)
Roth, V.: The generalized LASSO. IEEE Transactions on Neural Networks 15(1), 16–28 (2004)
Saeys, Y., Inza, I., Larrañaga, P.: A review of feature selection techniques in bioinformatics. Bioinformatics 23(19), 2507–2517 (2007)
Shipp, M., Ross, K., Tamayo, P., Weng, A., Kutok, J., Aguiar, R., Gaasenbeek, M., Angelo, M., Reich, M., Pinkus, G., Ray, T., Koval, M., Last, K., Norton, A., Lister, A., Mesirov, J.: Diffuse large b-cell lymphoma outcome prediction by gene-expression profiling and supervised machine learning. Nature Medicine 8, 68–74 (2002)
Singh, D., Febbo, P.G., Ross, K., Jackson, D.G., Manola, J., Ladd, C., Tamayo, P., Renshaw, A.A., D’Amico, A.V., Richie, J.P., Lander, E.S., Loda, M., Kantoff, P.W., Golub, T.R., Sellers, W.R.: Gene expression correlates of clinical prostate cancer behavior. Cancer Cell 1, 203–209 (2002)
Tibshirani, R.: Regression shrinkage and selection via the lasso. Journal of the Royal Statistical Society, Series B 58, 267–288 (1994)
Witten, D.M., Tibshirani, R.: A comparison of fold-change and the t-statistic for microarray data analysis. Stanford University. Technical report (2007)
Zou, H., Hastie, T.: Regularization and variable selection via the elastic net. Journal of the Royal Statistical Society, Series B 67, 301–320 (2005)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Zakharov, R., Dupont, P. (2011). Ensemble Logistic Regression for Feature Selection. In: Loog, M., Wessels, L., Reinders, M.J.T., de Ridder, D. (eds) Pattern Recognition in Bioinformatics. PRIB 2011. Lecture Notes in Computer Science(), vol 7036. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-24855-9_12
Download citation
DOI: https://doi.org/10.1007/978-3-642-24855-9_12
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-24854-2
Online ISBN: 978-3-642-24855-9
eBook Packages: Computer ScienceComputer Science (R0)