Abstract
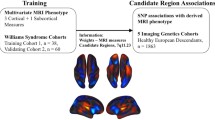
Genetic mapping of hippocampal shape, an under-explored area, has strong potential as a neurodegeneration biomarker for AD and MCI. This study investigates the genetic effects of top candidate single nucleotide polymorphisms (SNPs) on hippocampal shape features as quantitative traits (QTs) in a large cohort. FS+LDDMM was used to segment hippocampal surfaces from MRI scans and shape features were extracted after surface registration. Elastic net (EN) and sparse canonical correlation analysis (SCCA) were proposed to examine SNP-QT associations, and compared with multiple regression (MR). Although similar in power, EN yielded substantially fewer predictors than MR. Detailed surface mapping of global and localized genetic effects were identified by MR and EN to reveal multi-SNP-single-QT relationships, and by SCCA to discover multi-SNP-multi-QT associations. Shape analysis identified stronger SNP-QT correlations than volume analysis. Sparse multivariate models have greater power to reveal complex SNP-QT relationships. Genetic analysis of quantitative shape features has considerable potential for enhancing mechanistic understanding of complex disorders like AD.
Data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database ( www.loni.ucla.edu/ADNI ). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at: www.loni.ucla.edu/ADNI/Collaboration/ADNI_Authorship_list.pdf .
Chapter PDF
Similar content being viewed by others
Keywords
- Mild Cognitive Impairment
- Hippocampal Volume
- Canonical Vector
- Sparse Canonical Correlation Analysis
- AlzGene Database
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
References
Bertram, L., McQueen, M.B., Mullin, K., Blacker, D., Tanzi, R.E.: Systematic meta-analyses of alzheimer disease genetic association studies: the alzgene database. Nat. Genet. 39(1), 17–23 (2007)
Friedman, J., Hastie, T., Tibshirani, R.: Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33(1), 1–22 (2010)
Khan, A.R., Wang, L., Beg, M.F.: Freesurfer-initiated fully-automated subcortical brain segmentation in mri using large deformation diffeomorphic metric mapping. Neuroimage 41(3), 735–746 (2008)
Li, Y., Willer, C.J., Ding, J., Scheet, P., Abecasis, G.R.: MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet. Epidemiol. 34(8), 816–834 (2010)
Liu, J., Pearlson, G., Windemuth, A., Ruano, G., Perrone-Bizzozero, N.I., Calhoun, V.: Combining fmri and snp data to investigate connections between brain function and genetics using parallel ica. Hum. Brain Mapp. 30(1), 241–255 (2009)
Marchini, J., Howie, B., Myers, S., McVean, G., Donnelly, P.: A new multipoint method for genome-wide association studies via imputation of genotypes. Nature Genetics 39, 906–913 (2007)
Shen, L., Kim, S., Risacher, S.L., Nho, K., Swaminathan, S., West, J.D., Foroud, T., Pankratz, N., Moore, J.H., Sloan, C.D., Huentelman, M.J., Craig, D.W., Dechairo, B.M., Potkin, S.G., Jack Jr., C.R., Weiner, M.W., Saykin, A.J.: ADNI: Whole genome association study of brain-wide imaging phenotypes for identifying quantitative trait loci in MCI and AD: A study of the ADNI cohort. Neuroimage 53(3), 1051–1063 (2010)
Stein, J.L., Hua, X., Lee, S., Ho, A.J., Leow, A.D., Toga, A.W., Saykin, A.J., Shen, L., Foroud, T., Pankratz, N., Huentelman, M.J., Craig, D.W., Gerber, J.D., Allen, A.N., Corneveaux, J.J., Dechairo, B.M., Potkin, S.G., Weiner, M.W., Thompson, P.: Voxelwise genome-wide association study (vgwas). Neuroimage 53(3), 1160–1174 (2010)
Tibshirani, R.: Glmnet, http://www-stat.stanford.edu/~tibs/glmnet-matlab/
Vounou, M., Nichols, T.E., Montana, G.: Discovering genetic associations with high-dimensional neuroimaging phenotypes: A sparse reduced-rank regression approach. Neuroimage 53(3), 1147–1159 (2010)
Witten, D.M., Tibshirani, R., Hastie, T.: A penalized matrix decomposition, with applications to sparse principal components and canonical correlation analysis. Biostatistics 10(3), 515–534 (2009)
Worsley, K.J.: Surfst, http://www.math.mcgill.ca/keith/surfstat
Worsley, K.J., Andermann, M., Koulis, T., MacDonald, D., Evans, A.C.: Detecting changes in nonisotropic images. Hum. Brain Mapp. 8(2-3), 98–101 (1999)
Zou, H., Hastie, T.: Regularization and variable selection via the elastic net. J. R. Statist. Soc. 67(2), 301–320 (2005)
Author information
Authors and Affiliations
Consortia
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Wan, J. et al. (2011). Hippocampal Surface Mapping of Genetic Risk Factors in AD via Sparse Learning Models. In: Fichtinger, G., Martel, A., Peters, T. (eds) Medical Image Computing and Computer-Assisted Intervention – MICCAI 2011. MICCAI 2011. Lecture Notes in Computer Science, vol 6892. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-23629-7_46
Download citation
DOI: https://doi.org/10.1007/978-3-642-23629-7_46
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-23628-0
Online ISBN: 978-3-642-23629-7
eBook Packages: Computer ScienceComputer Science (R0)