Abstract
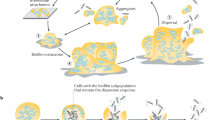
Biofilm dispersion has become widely recognized as a natural phenomenon associated with the terminal stage of biofilm development. As a behavioral characteristic of bacteria, this process is of major significance due to its importance in regulating biofilm structure. Biofilm dispersion is also important as a potential control point for the manipulation of biofilm development and persistence. Over the past decade, an increasing number of laboratories have focused their efforts on the study of biofilm dispersion, resulting in findings on the intracellular and extracellular mechanisms involved in this process in selected species of bacteria. The following is an overview of our current understanding of the biofilm dispersion process and the mechanisms involved in its regulation.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Allison DG, Ruiz B, SanJose C, Jaspe A, Gilbert P (1998) Extracellular products as mediators of the formation and detachment of Pseudomonas fluorescens biofilms. FEMS Microbiol Lett 167:179–184
Allwood A, Walter MR, Burch IW, Kamber BS (2007) 3.43 billion-year-old stromatolite reef from the Pilbara Craton of Western Australia: ecosystem-scale insights to early life on Earth. Precambrian Res 158:198–227
Barber CE et al (1997) A novel regulatory system required for pathogenicity of Xanthomonas campestris is mediated by a small diffusible signal molecule. Mol Microbiol 24:555–566
Barraud N et al (2006) Involvement of nitric oxide in biofilm dispersal of Pseudomonas aeruginosa. J Bacteriol 188:7344–7353
Boles BR, Horswill AR (2008) Agr-mediated dispersal of Staphylococcus aureus biofilms. PLoS Pathog 4:e1000052
Boles BR, Thoendel M, Singh PK (2005) Rhamnolipids mediate detachment of Pseudomonas aeruginosa from biofilms. Mol Microbiol 57:1210–1223
Boon C et al (2007) A novel DSF-like signal from Burkholderia cenocepacia interferes with Candida albicans morphological transition. ISME J 2:27–36
Bowden GH, Li YH (1997) Nutritional influences on biofilm development. Adv Dent Res 11:81–99
Boyd A, Chakrabarty AM (1994) Role of alginate lyase in cell detachment of Pseudomonas aeruginosa. Appl Environ Microbiol 60:2355–2359
Breyers JD (1988) Modeling biofilm accumulation. In: Bazin MJ, Prosser JI (eds) Physiology models in microbiology, vol 2. Boca Raton, FL, pp 109–144
Chatterjee S, Newman KL, Lindow SE (2008) Cell-to-cell signaling in Xylella fastidiosa suppresses movement and xylem vessel colonization in grape. Mol Plant-Microbe Interact 21:1309–1315
Chen X, Stewart PS (2000) Biofilm removal caused by chemical treatments. Water Res 34:4229–4233
Christen M, Christen B, Folcher M, Schauerte A, Jenal U (2005) Identification and characterization of a cyclic di-GMP-specific phosphodiesterase and its allosteric control by GTP. J Biol Chem 280:30829–30837
Costerton JW, Lewandowski Z, Caldwell DE, Korber DR, Lappin-Scott HM (1995) Microbial biofilms. Annu Rev Microbiol 49:711–745
Danhorn T, Hentzer M, Givskov M, Parsek MR, Fuqua C (2004) Phosphorus limitation enhances biofilm formation of the plant pathogen Agrobacterium tumefaciens through the PhoR-PhoB regulatory system. J Bacteriol 186:4492–4501
Davey ME, Caiazza NC, O’Toole GA (2003) Rhamnolipid surfactant production affects biofilm architecture in Pseudomonas aeruginosa PAO1. J Bacteriol 185:1027–1036
David GA, Begoña R, Carmen S, Almudena J, Peter G (1998) Extracellular products as mediators of the formation and detachment of Pseudomonas fluorescens biofilms. FEMS Microbiol Lett 167:179–184
Davies DG, Marques CNH (2009) A fatty acid messenger is responsible for inducing dispersion in microbial biofilms. J Bacteriol 191:1393–1403
Davies DG (1999) Regulation of matrix polymer in biofilm formation and dispersion. In: Wingender J, Neu TR, Flemming H-C (eds) Microbial extrapolymeric substances, characterization, structure and function. Springer, Berlin, pp 93–112
Delaquis PJ, Caldwell DE, Lawrence JR, McCurdy AR (1989) Detachment of Pseudomonas fluorescens from biofilms on glass surfaces in response to nutrient stress. Microb Ecol 18:199–210
Dewanti R, Wong ACL (1995) Influence of culture conditions on biofilm formation by Escherichia coli O157:H7. Int J Food Microbiol 26:147–164
Dow JM et al (2003) Biofilm dispersal in Xanthomonas campestris is controlled by cell-cell signaling and is required for full virulence to plants. Proc Natl Acad Sci 100:10995–11000
Ferreira RBR, Antunes LCM, Greenberg EP, McCarter LL (2008) Vibrio parahaemolyticus ScrC modulates cyclic dimeric GMP regulation of gene expression relevant to growth on surfaces. J Bacteriol 190:851–860
Fouhy Y et al (2007) Diffusible signal factor-dependent cell-cell signaling and virulence in the nosocomial pathogen Stenotrophomonas maltophilia. J Bacteriol 189:4964–4968
Friedman L, Kolter R (2004a) Two genetic loci produce distinct carbohydrate-rich structural components of the Pseudomonas aeruginosa biofilm matrix. J Bacteriol 186:4457–4465
Friedman L, Kolter R (2004b) Genes involved in matrix formation in Pseudomonas aeruginosa PA14 biofilms. Mol Microbiol 51:675–690
Gacesa P (1987) Alginate-modifying-enzymes: a proposed unified mechanism of action for the lyases and epimerases. FEBS Lett 212:199–202
Güvener ZT, Harwood CS (2007) Subcellular location characteristics of the Pseudomonas aeruginosa GGDEF protein, WspR, indicate that it produces cyclic-di-GMP in response to growth on surfaces. Mol Microbiol 66:1459–1473
Hickman JW, Tifrea DF, Harwood CS (2005) A chemosensory system that regulates biofilm formation through modulation of cyclic diguanylate levels. Proc Natl Acad Sci USA 102:14422–14427
Hisatsuka K, Nakahara T, Sano N, Yamada K (1971) Formation of rhamnolipid by Pseudomonas aeruginosa and its function in hydrocarbon fermentation. Agric Biol Chem 35:686–692
Hofmann HJ, Grey K, Hickman AH, Thorpe RI (1999) Origin of 3.45 Ga coniform stromatolites in Warrawoona group, Western Australia. Geol Soc Am Bull 111:1256–1262
Huang T-P, Wong ACL (2007) A cyclic AMP receptor protein-regulated cell-cell communication system mediates expression of a FecA homologue in Stenotrophomonas maltophilia. Appl Environ Microbiol 73:5034–5040
Hunt SM, Werner EM, Huang B, Hamilton MA, Stewart PS (2004) Hypothesis for the role of nutrient starvation in biofilm detachment. Appl Environ Microbiol 70:7418–7425
Itoh Y, Wang X, Hinnebusch BJ, Preston JF III, Romeo T (2005) Depolymerization of {beta}-1,6-N-acetyl-D-glucosamine disrupts the integrity of diverse bacterial biofilms. J Bacteriol 187:382–387
Jackson DW, Simecka JW, Romeo T (2002) Catabolite repression of Escherichia coli biofilm formation. J Bacteriol 184:3406–3410
Jackson KD, Starkey M, Kremer S, Parsek MR, Wozniak DJ (2004) Identification of psl, a locus encoding a potential exopolysaccharide that is essential for Pseudomonas aeruginosa PAO1 biofilm formation. J Bacteriol 186:4466–4475
James GA, Korber DR, Caldwell DE, Costerton JW (1995) Digital image analysis of growth and starvation responses of a surface-colonizing Acinetobacter sp. J Bacteriol 177:907–915
Kaplan JB, Fine DH (2002) Biofilm dispersal of neisseria subflava and other phylogenetically diverse oral bacteria. Appl Environ Microbiol 68:4943–4950
Kaplan JB, Ragunath C, Ramasubbu N, Fine DH (2003) Detachment of Actinobacillus actinomycetemcomitans biofilm cells by an endogenous {beta}-hexosaminidase activity. J Bacteriol 185:4693–4698
Karatan E, Watnick P (2009) Signals, regulatory networks, and materials that build and break bacterial biofilms. Microbiol Mol Biol Rev 73:310–347
Kim Y-K, McCarter LL (2007) ScrG, a GGDEF-EAL protein, participates in regulating swarming and sticking in Vibrio parahaemolyticus. J Bacteriol 189:4094–4107
Lamed R, Bayer EA (1986) Contact and cellulolysis in Clostridium thermocellum via extensive surface organelles. Experientia 42:72–73
Lee J, Bansal T, Jayaraman A, Bentley WE, Wood TK (2007a) Enterohemorrhagic Escherichia coli biofilms are inhibited by 7-hydrozyindole and stimulated by isatin. Appl Environ Microbiol 73:4100–4109
Lee J, Jayaraman A, Wood TK (2007b) Indole is an inter-species biofilm signal mediated by SdiA. BMC Microbiol 7:1–15
Lynch MJ et al (2002) The regulation of biofilm development by quorum sensing in Aeromonas hydrophila. Environ Microbiol 4:18–28
Ma L, Jackson KD, Landry RM, Parsek MR, Wozniak DJ (2006) Analysis of Pseudomonas aeruginosa conditional Psl variants reveals roles for the Psl polysaccharide in adhesion and maintaining biofilm structure postattachment. J Bacteriol 188:8213–8221
Ma L, Lu H, Sprinkle A, Parsek MR, Wozniak D (2007) Pseudomonas aeruginosa Psl is a galactose- and mannose-rich exopolysaccharide. J Bacteriol. doi:10.1128/JB.00620-07
Marshall JC (1988) Adhesion and growth of bacteria at surfaces in oligotrophic habitats. Can J Microbiol 34:503–506
Matsukawa M, Greenberg EP (2004) Putative exopolysaccharide synthesis genes influence Pseudomonas aeruginosa biofilm development. J Bacteriol 186:4449–4456
May TB et al (1991) Alginate synthesis by Pseudomonas aeruginosa: a key pathogenic factor in chronic pulmonary infections of cystic fibrosis patients. Clin Microbiol Rev 4:191–206
Merritt JH, Brothers KM, Kuchma SL, O’Toole GA (2007) SadC reciprocally influences biofilm formation and swarming motility via modulation of exopolysaccharide production and flagellar function. J Bacteriol 189(22):8154–8164
Morgan R, Kohn S, Hwang S-H, Hassett DJ, Sauer K (2006) BdlA, a chemotaxis regulator essential for biofilm dispersion in Pseudomonas aeruginosa. J Bacteriol 188:7335–7343
Ohashi A, Harada H (1994a) Adhesion strength of biofilm developed in an attached-growth reactor. Water Sci Technol 20:10–11
Ohashi A, Harada H (1994b) Characterization of detachment mode of biofilm developed in an attached-growth reactor. Water Sci Technol 30:35–45
Ohashi A, Koyama T, Syutsubo K, Harada H (1999) A novel method for evaluation of biofilm tensile strength resisting erosion. Water Sci Technol 39:261–268
O’Toole G, Kaplan HB, Kolter R (2000) Biofilm formation as microbial development. Annu Rev Microbiol 54:49–79
Peyton BM, Characklis WG (1993) A statistical analysis of the effect of substrate utilization and shear stress on the kinetics of biofilm detachment. Biotechnol Bioeng 41:728–735
Pratt LA, Kolter R (1999) Genetic analyses of bacterial biofilm formation. Curr Opin Microbiol 2:598–603
Purevdorj-Gage B, Costerton WJ, Stoodley P (2005) Phenotypic differentiation and seeding dispersal in non-mucoid and mucoid Pseudomonas aeruginosa biofilms. Microbiology 151:1569–1576
Puskas A, Greenberg EP, Kaplan S, Schaefer AL (1997) A quorum-sensing system in the free-living photosynthetic bacterium Rhodobacter sphaeroides. J Bacteriol 179:7530–7537
Rice SA et al (2005) Biofilm formation and sloughing in Serratia marcescens are controlled by quorum sensing and nutrient cues. J Bacteriol 187:3477–3485
Rittman BR (1982) The effect of shear stress on biofilm loss rate. Biotechnol Bioeng 24:501–506
Rochex A, Lebeault JM (2007) Effects of nutrients on biofilm formation and detachment of a Pseudomonas putida strain isolated from a paper machine. Water Res 41:2885–2992
Roger S, Michael M, Abdul K, Manfred N, Ute R (2004) GGDEF and EAL domains inversely regulate cyclic di-GMP levels and transition from sessility to motility. Mol Microbiol 53:1123–1134
Romeo T (1998) Global regulation by the small RNA-binding protein CsrA and the non-coding RNA molecule CsrB. Mol Microbiol 29:1321–1330
Romeo T, Gong M, Liu MY, Brun-Zinkernagel AM (1993) Identification and molecular characterization of csrA, a pleiotropic gene from Escherichia coli that affects glycogen biosynthesis, gluconeogenesis, cell size, and surface properties. J Bacteriol 175:4744–4755
Ross P et al (1990) The cyclic diguanylic acid regulatory system of cellulose synthesis in Acetobacter xylinum. Chemical synthesis and biological activity of cyclic nucleotide dimer, trimer, and phosphothioate derivatives. J Biol Chem 265:18933–18943
Ryan RP et al (2006) Cell-cell signaling in Xanthomonas campestris involves an HD-GYP domain protein that functions in cyclic di-GMP turnover. Proc Natl Acad Sci USA 103:6712–6717
Sabnis NA, Yang H, Romeo T (1995) Pleiotropic regulation of central carbohydrate metabolism in Escherichia coli via the gene csrA. J Biol Chem 270:29096–29104
Sauer K, Camper AK, Ehrlich GD, Costerton JW, Davies DG (2002) Pseudomonas aeruginosa displays multiple phenotypes during development as a biofilm. J Bacteriol 184:1140–1154
Sauer K et al (2004) Characterization of nutrient-induced dispersion in Pseudomonas aeruginosa PAO1 biofilm. J Bacteriol 186:7312–7326
Sawyer LK, Hermanowicz SW (1998) Detachment of biofilm bacteria due to variations in nutrient supply. Water Sci Technol 37:211–214
Simm R, Morr M, Kader A, Nimtz M, Romling U (2004) GGDEF and EAL domains inversely regulate cyclic di-GMP levels and transition from sessility to motility. Mol Microbiol 53:1123–1134
Slater H, Alvarez-Morales A, Barber CE, Daniels MJ, Dow JM (2000) A two-component system involving an HD-GYP domain protein links cell-cell signalling to pathogenicity gene expression in Xanthomonas campestris. Mol Microbiol 38:986–1003
Stanley NR, Britton RA, Grossman AD, Lazazzera BA (2003) Identification of catabolite repression as a physiological regulator of biofilm formation by Bacillus subtilis by use of DNA microarrays. J Bacteriol 185:1951–1957
Stewart PS (1993) A model of biofilm detachment. Biotechnol Bioeng 41:111–117
Stoodley P et al (2001) Growth and detachment of cell clusters from mature mixed-species biofilms. Appl Environ Microbiol 67:5608–5613
Tal R et al (1998) Three cdg operons control cellular turnover of cyclic di-GMP in Acetobacter xylinum: genetic organization and occurrence of conserved domains in isoenzymes. J Bacteriol 180:4416–4425
Thormann KM, Saville RM, Shukla S, Spormann AM (2005) Induction of rapid detachment in Shewanella oneidensis MR-1 biofilms. J Bacteriol 187:1014–1021
Thormann KM et al (2006) Control of formation and cellular detachment from Shewanella oneidensis MR-1 biofilms by cyclic di-GMP. J Bacteriol 188:2681–2691
Tolker-Nielsen et al T (2000) Development and dynamics of Pseudomonas sp. biofilms. J Bacteriol 182:6482–6489
Trulear MG, Characklis WG (1982) Dynamics of biofilm processes. J Water Pollut Control Fed 9:1288–1301
van Loosdrecht MCM, Picioreanu C, Heijnen JJ (1997) A more unifying hypothesis for the structure of microbial biofilms. FEMS Microbiol Ecol 24:181–183
Vats N, Lee SF (2000) Active detachment of Streptococcus mutans cells adhered to epon-hydroxylapatite surfaces coated with salivary proteins in vitro. Arch Oral Biol 45:305–314
Wang LH et al (2004) A bacterial cell-cell communication signal with cross-kingdom structural analogues. Mol Microbiol 51:903–912
Whitchurch CB, Tolker-Nielsen T, Ragas PC, Mattick JS (2002) Extracellular DNA required for bacterial biofilm formation. Science 295:1487
Xun L, Mah RA, Boone DR (1990) Isolation and characterization of disaggregatase from Methanosarcina mazei LYC. Appl Environ Microbiol 56:3693–3698
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Davies, D.G. (2011). Biofilm Dispersion. In: Flemming, HC., Wingender, J., Szewzyk, U. (eds) Biofilm Highlights. Springer Series on Biofilms, vol 5. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-19940-0_1
Download citation
DOI: https://doi.org/10.1007/978-3-642-19940-0_1
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-19939-4
Online ISBN: 978-3-642-19940-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)