Abstract
The SARS coronavirus (CoV) spike protein is a target of intensive research, as it is a major virulence factor. Transfection of SARS-CoV spike into Vero E6, HEK293T and HL-CZ cells leads to strong expression of the glycosylated spike protein, as shown by Western blot analyses and immunofluorescent imaging using spike-specific human monoclonal antibodies, indicating the potential utility of these antigens and antibodies as diagnostic reagents. Furthermore, we employed cDNA microarray analysis to probe the changes in host gene transcription attributed to transfection of a codon-optimized spike construct into the HL-CZ cell line of monocyte lineage that is linked to immunological responses. A diverse representation of 100 genes displayed altered transcriptional patterns in response to SARS-CoV spike expression, with 61 upregulated and 39 downregulated genes. Genes involved in intracellular trafficking, signaling, modulation or transcription were generally upregulated. In contrast, genes involved in cell metabolism and cytoskeleton formation were notably downregulated. The transcripts of other functional categories exhibited varied responses to SARS-CoV spike transfection. Collectively, our analyses elucidate numerous and complex transcriptomic events that occur in response to spike protein expression and that contribute towards SARS-CoV pathogenesis.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
- Severe Acute Respiratory Syndrome
- Human Monoclonal Antibody
- Severe Acute Respiratory Syndrome
- Severe Acute Respiratory Syndrome Patient
- Spike Protein
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 SARS Coronavirus: A Lethal Zoonotic Virus with Future Potential for Reemergence
In November 2002, the outbreak of the newly emerging severe acute respiratory syndrome (SARS) in many parts of the world, especially in South-East Asia and China, resulted in more than 8,000 cases, with a high fatality rate of about 10%. Infection is characterized by acute flu-like symptoms, causing severe atypical pneumonia with diffuse alveolar damage, and death within two weeks in severely infected patients. This global outbreak of SARS wreaked havoc on public health and socioeconomic stability, but was finally brought under control in July 2003.
Intensive collaboration identified the etiological agent as a new member of the genus Coronavirus. Analyses of the complete nucleotide sequence of the novel SARS-associated coronavirus (SARS-CoV) reveal significant similarities to the other coronaviruses, with the genes encoding the spike (S), envelope (E), membrane (M) and nucleocapsid (N) proteins possessing similar sequence frames. In addition, the novel SARS-CoV contains several unique genes (Poon et al. 2004).
Similar viruses with considerable homology to human SARS-CoV have been isolated from animals such as palm civets and horseshoe bats, suggesting zoonotic reservoirs and the possible reemergence of this deadly virus (Shi and Hu 2008). Hence, the mechanisms of SARS-CoV pathogenicity are the subject of extensive scientific research (Satija and Lal 2007). Furthermore, it is highly imperative that effective and safe vaccines are developed to counter a potential SARS epidemic in the future (He et al. 2004a, b). Candidate SARS-CoV vaccines have been successfully tested in animal models and in clinical trials. The lack of specific therapies against SARS necessitates molecular approaches for selecting attractive targets to facilitate the design of novel vaccines and antiviral agents against SARS-CoV infections (Groneberg et al. 2005; Han et al. 2006; van den Brink et al. 2005; ter Meulen et al. 2004, 2006; Poh et al. 2009).
2 SARS-CoV Spike Glycoprotein Contributes to Virulence and Pathogenesis
Comprising 1,255 amino acid residues, the SARS-CoV spike is the first protein to interact with the host membrane, culminating in a cascade of infection events. Given that this glycoprotein plays a crucial role in viral virulence and pathogenesis, its functions have been extensively characterized, and it is probably the most important viral target for vaccine design (Du et al. 2009). Containing intracellular, extracellular and transmembrane domains, the spike protein is classified as a type 1 transmembrane glycoprotein that is responsible for virus binding, fusion and entry (Simmons et al. 2004; Zhang et al. 2004; Zhou et al. 2004). Of particular interest and importance is the S1 unit which constitutes one of the two subunits of the extracellular domain. This domain is responsible for binding of the virus to its main receptor, angiotensin-converting enzyme 2 (ACE2), present on susceptible cells (Li et al. 2003; Babcock et al. 2004). Moreover, CD209L (L-SIGN) and DC-SIGN also facilitate cell entry of SARS-CoV (Jeffers et al. 2004; Shih et al. 2006). SARS-CoV spike is selectively recognized by pulmonary surfactant D and activates macrophages (Leth-Larsen et al. 2007).
Vero E6 cells are derived from the African green monkey kidney clone E6, and are the most permissive for SARS-CoV infection and replication. Human cell lines that are susceptible to SARS-CoV infection include LoVo, HepG2 and Huh-7. The lytic infection produced in Vero E6 cells mimics the apoptosis observed in the respiratory epithelial cells of severely ill SARS patients (Leong et al. 2005; Chow and Leong 2008). In contrast, persistent infection of LoVo cells does not exhibit visible cytopathic effect (CPE).
3 Generation of Human Monoclonal Antibodies by Selection from Recombinant Single-Chain Antibody-Phage Libraries Constructed from the Lymphocytes of a Convalescent SARS Patient
Antibody phage display is an established technology for the isolation of antibodies from large libraries of these molecules (Pini and Bracci 2000). Antibody-binding regions (Fab or scFv) are cloned as fusions of the phage M13 coat proteins. The procedure results in the isolation of genes encoding antibody-binding regions specific for the target antigen. These genes can be used for the construction of fully human immunoglobulin molecules, i.e., phage libraries can be either naïve or immune. The latter library is generated using antibody genes extracted from the B cells of an immunized donor or from a convalescent patient. Both methods may yield antibodies that can be employed for therapeutic applications (Hoogenboom 2002).
An immune scFv phage library was constructed from the lymphocytes of a convalescent Singapore SARS patient with high neutralizing antibody titers. Briefly, several hundred scFv clones were checked for specific binding to inactivated SARS-CoV. After further selection, binding affinities of monovalent scFv were estimated, and scFv-binding epitopes were mapped. Finally, V regions were cloned into IgG expression vectors for the expression of human IgG1 molecules in the human PER.C6 cell line.
Thus, human monoclonal antibodies against the SARS-CoV spike glycoprotein have been successfully generated, and shown to completely prevent lung pathology and abolish pharyngeal shedding of SARS-CoV in infected ferrets (ter Meulen et al. 2004, 2006).
4 Expression of SARS-CoV Spike Protein in Mammalian Cell Lines and Specific Detection by Human Monoclonal Antibodies: Potential Application as Diagnostic Reagents
The human monoclonal antibodies (16 designated pgG103-014C03P01, pgG103-019C03P01, pgG103-020C03P01, pgG103-022C03P01, pgG103-023C03P01, pgG103-023aC03P01, pgG103-024C03P01, pgG103-031C03P01, pgG103-046C03P01, pgG103-050C03P01, pgG103-053C03P01, pgG103-054C03P01, pgG103-055C03P01, pgG103-056C03P01, pgG103-057C03P01 and pgG103-059C03P01) were analyzed by Western blotting and immunocytochemistry on human cell lines (HL-CZ and HEK293T) transfected with spike construct or with the pcDNA3.1 vector alone as control.
4.1 Western Blot Characterization of HL-CZ Cells Transfected with Codon-Optimized SARS-CoV Spike Construct
For expression of the spike protein, the pCon-S-Harv-Myc-His plasmid (incorporating the extracellular domain of the spike protein) was constructed on a pcDNA3.1 vector backbone with a double tag of Myc and His (kindly provided by Dr Michael Farzan, Harvard University). The construction of this codon-optimized SARS-CoV spike ectodomain is more effective for spike protein expression during transfection in mammalian cells by replacing the natural codons with optimum codons (Babcock et al. 2004).
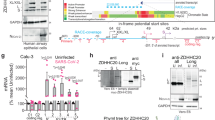
HL-CZ is a suspension cell line derived from human promonocytic leukemia that possesses CD15, CD34 and CD11b markers, and is permissive for the propagation of dengue virus, influenza virus type A and HIV-1 (Lai et al. 2007). Twenty-four hours after cell seeding, one batch of HL-CZ cells was transiently transfected with pCon-S-Harv-Myc-His using TransFectin (Bio-Rad), while another batch was mock-transfected with pcDNA3.1 only as control. Protein samples of both batches were then analyzed by SDS-PAGE and immunoblotting using the human monoclonal antibodies to SARS-CoV spike. Among the 16 antibodies, the antibody designated pgG103-057C03P01 detected a single distinct band of ~180 kDa compatible with the glycosylated spike ectodomain in the total cell lysate and pellet of HL-CZ cells transfected with SARS-CoV spike, but not in the corresponding samples of cells transfected with pcDNA3.1 control. With the pgG103-057C03P01 antibody, Western blotting also revealed intense SARS-CoV spike bands in both the lysate and supernatant of spike-transfected human embryonic kidney HEK293T cells. These immunoblot results were further confirmed by using spike polyclonal antibody IMG-541 (Imgenex) which detected a weaker spike protein band (Fig. 18.1).
Western blot analyses of SARS-CoV spike protein in transfected HL-CZ and HEK293T cells. Lanes 1–7 were probed with anti-spike human monoclonal antibody pgG103-057C03P01 followed by secondary antihuman antibody. SARS-CoV spike bands of ~180 kDa were detected in cells transfected with spike construct, i.e., lanes 2 (HL-CZ lysate), 4 (HL-CZ pellet), 6 (HEK293T lysate) and 7 (HEK293T supernatant). A similar but weak band was observed in lane 8 (pellet of spike-transfected HL-CZ cells but probed with commercial antibody). This protein was absent in cells transfected with pcDNA3.1 vector alone, i.e., lanes 1 (HL-CZ lysate), 3 (HL-CZ pellet) and 5 (HEK293T lysate). The blots were developed using an ECL kit (Amersham Pharmacia). Lane M depicts the prestained Kaleidoscope Protein ladder markers (Bio-Rad)
4.2 Microscopic Analysis of SARS-CoV Spike Expression in HL-CZ Cells by Indirect Immunofluorescence Labeling
Prior to transfection, HL-CZ cells were cultured on glass coverslips coated with Cell-Tak (BD Biosciences). To visualize the intracellular distribution of the spike protein, immunofluorescence assay (IFA) was performed on cells transfected for 24 h with either pCon-S-Harv-Myc-His or control cells transfected with the pcDNA3.1 vector alone. Monoclonal antibody pgG103-057C03P01 exhibited the strongest staining of spike protein with predominant cytoplasmic localization (Fig. 18.2a–c). Antibodies pgG103-023C03P01 and pgG103-054C03P01 also displayed positive but relatively less staining, mainly in the cytoplasm. However, antibodies such as pgG103-055C03P01 did not demonstrate any significant fluorescence (data not shown).
Intracellular distribution of spike protein in HL-CZ cells transiently transfected for 24 h with (a–c) pCon-Spike-Harv-Myc-His construct or (d) pcDNA3.1 vector only, and incubated with anti-spike human monoclonal antibody pgG103-057C03P01 followed by secondary FITC-labeled antihuman antibody. (e) Vero and (f) HL-CZ cells were stained with DAPI to display nuclei, and with anti-spike polyclonal antibody followed by secondary antibody labeled with rhodamine. The superimposed images localize the spike protein to the cytoplasm giving rise to characteristic “ring” structures (arrows)
Transfected cells were also incubated with primary rabbit polyclonal antibody against SARS-CoV spike and secondary antirabbit antibody labeled with rhodamine. DAPI counterstaining was performed to highlight cell nuclei. Confocal microscopy revealed the wide distribution of spike in the cytoplasm of Vero E6 and HL-CZ cells transfected with pCon-S-Harv-Myc-His. The intense immunostaining in the cytoplasm of HL-CZ cells may be attributed to spike localization in smooth membrane vesicles or smooth endoplasmic reticulum. Spike staining is prominent in Vero E6 cells, which may partly be explained by their adherent property (Fig. 18.2e–f).
While spike expression has also been localized to the cell membrane, confocal microscopy studies show that SARS-CoV spike protein subunit C localizes in the cytoplasm of Sf-9 cells. The detection sensitivity of the spike protein-based IFA is reflected by its ability to detect SARS-CoV specific antibody at a lower dilution compared to commercial IFAs (Manopo et al. 2005).
Our results indicate that the pgG103-057C03P01 human monoclonal antibody is highly specific for detecting SARS-CoV spike antigen. Furthermore, transfected cell lines expressing the SARS-CoV spike extracellular domain can also serve as highly specific reagents for serodiagnosis of clinical specimens by Western blotting or immunocytochemistry.
5 Transcriptomic Analysis Reveals Differentially Regulated Genes in HL-CZ Cells Following SARS-CoV Spike Transfection
The interactions between host and virus involve an elaborate and intricate interplay between molecular pathways initiated by the host in response to the infection and the mechanisms triggered by the virus to successfully infect the cell. Unraveling the transcriptomic events that ensue from the host–virus interactions can enhance our understanding of viral pathogenesis and disease manifestations, and can provide clues for designing preventive and therapeutic strategies to counter the infection (Leong and Chow 2006; Liew and Chow 2006). For example, multifarious and intriguing changes in genetic expression occur during infection of Vero E6 cells with whole live SARS-CoV (Leong et al. 2005). Compared with HL-CZ cells, Vero E6 cells are highly permissive to SARS-CoV infection and exhibit extensive CPE.
In order to gain insights into the role of the SARS-CoV spike protein in the human genetic responses and pathophysiology of SARS, we employed cDNA microarray to investigate gene expression changes in HL-CZ promonocytic cells following spike transfection. The various downstream transcriptomic changes that occur following this interaction will offer a clearer understanding of the role of spike during SARS-CoV pathogenesis.
Total cellular RNAs were extracted from HL-CZ cells transfected with pCon-S-Harv-Myc-His construct or with pcDNA3.1 alone using the SV Total RNA isolation kit (Promega). Labeling of first-strand cDNA with amino allyl-dUTP and purification of the CyDye-labeled cDNA prior to hybridization were performed with the CyScribe Post-Labeling kit and CyScribe GFX Purification kit (Amersham Biosciences). In both slides of set A of the gene array, Cy3 dye was incorporated into the spike-transfected cellular RNA, and Cy5 dye into vector-transfected cellular RNA. For set B, Cy5 and Cy3 dyes were incorporated into total RNA from cells transfected with spike and vector alone, respectively. This allows comparison of Cy-dye uptake efficiency as well as authentication of the microarray data.
Gene expression studies were performed using the Atlas Human 3.8I and 3.8II cDNA microarray panel (BD Biosciences) which comprises 7,600 human genes spotted onto two slides each consisting of 3,800 genes. Following microarray hybridization, the slides were scanned for Cy3 and Cy5 with a Genepix 4000B dual color image scanner (Axon Instruments).
Using the Acuity analysis software, low quality spots (e.g., extreme unevenness in intensity) were flagged off, but spots that exhibited sufficient signals were retained. Log values obtained from the scanning of labeled spike-transfected cellular cDNA were then normalized against values from labeled vector-transfected cDNA using locally weighted linear regression (LOWESS) analysis. Significant scan log ratios (spike versus vector transfection) of transcripts were then translated to actual fold changes in gene expression. By defining the threshold to be equivalent to greater than 1.96 standard deviations from the local mean based on the duplicated set, a total of 100 genes were identified at the 95% confidence level to be differentially expressed greater than two-fold. Of these, 61 transcripts were significantly upregulated, while 39 genes displayed reduced expression. Table 18.1 classifies the transcripts that were significantly altered at 24 h after spike transfection according to their known functional roles, including mRNAs of unknown function.
The most distinguishable genes to be downregulated by spike transfection were those involved in cell metabolism and cytoskeletal framework, while genes that mediate protein trafficking were generally upregulated. This was accompanied by a mixed response of transcripts that encode cell cycle and other proteins, suggesting an interesting interplay of host transcriptional alterations in response to spike protein expression.
5.1 Immune and Inflammatory Genes
Patients infected by SARS-CoV exhibit high plasma levels of IL1, IL6, IL8, IL12, monocyte chemoattractant protein-1 and IFN-γ inducible protein. High levels of IL8 and IL6 are associated with lung lesions in SARS patients at the acute stage. Elevations of IL8 and Th1-related cytokines induce hyperinflammatory innate responses following SARS-CoV invasion of the respiratory tract (Wang et al. 2007; Thiel and Weber 2008). Recombinant baculovirus expressing amino acids 17–688 of the spike protein induces AP-1 activation through MAP kinases and AP-1, with the domains spanning residues 324–488 and 609–688 being crucial for the induction of IL8 release in lung cells (Chang et al. 2004). IL8 induction can be specifically inhibited by antibody against ACE2. The IL-8 receptor binds to IL8 with high affinity, also binds to chemokine (C–X–C motif) ligand 1 (CXCL1/MGSA), and mediates neutrophil migration to sites of inflammation.
From our microarray data, the sharp upregulation of IL8 receptor transcript β (~12-fold increase) implies that spike actively induces expression of this receptor which has a stronger affinity for IL8. This finding concurs with Versteeg et al. (2007) who documented IL8 receptor induction by SARS-CoV spike. Interestingly, our previous study showed that the expression of IL8 receptor A or IL8RA (which is a low affinity receptor for IL8) is also upregulated in Vero E6 cells infected with SARS-CoV (Leong et al. 2005). Overproduction of specific inflammatory cytokines is a typical hallmark of SARS-CoV infections, and overactivity of the IL-8 receptor may be partly mediated by spike protein as an inflammatory response.
Another upregulated gene is the major histocompatibility complex (MHC) class I A. MHC class I molecules are expressed in nearly all cells, and play a central role in the immune system by presenting peptides derived from the endoplasmic reticulum lumen. Given that HL-CZ cells are derived from adult T cells on which MHC class I is typically expressed, upregulation of this gene is not surprising. In addition, the B-cell-specific transcription factor mRNA is increased ~9-fold upon spike transfection, indicating B-lymphocyte activation in response to SARS-CoV infection.
5.2 Cell Cycle Genes
Infectious bronchitis coronavirus infection induces cell cycle perturbations with downstream effects on viral replication (Dove et al. 2006). SARS-CoV spike expression in HL-CZ cells is also accompanied by differential responses of transcripts that encode cell cycle-related proteins. Following spike transfection, the cyclin A1 transcript is upregulated ~4-fold. Cyclin A1 binds to and regulates cdk1 and cdk2 which are critical factors during the S and G2 phases of the cell cycle. Consequently, modification of cyclin A1 transcription has an indirect impact on virus maturation and life cycle. In HIV-1 progeny formation, cyclin A1 binds Rb family members, and the p21/waf1 family of endogenous cdk inhibitors, as well as the E2F-1 transcriptional factor, all of which are important in regulating cell cycle progression (Liang et al. 2005).
Also upregulated is the retinoblastoma-like 1 (p107) (RBL1) transcript variant 1 whose protein is similar in sequence and possibly function to the retinoblastoma 1 (RB1) gene product. RB1 is a tumor suppressor protein involved in cell cycle regulation, being phosphorylated in the S to M phase transition, but is dephosphorylated in the G1 phase.
Prothymosin α is an abundant acidic nuclear protein with a role in supporting cell proliferation. It is present in all mammalian tissues, and is usually proportional to the proliferative activity of the tissue. Although this protein does not directly regulate cell division, it is required for entry from the G2 to M phase, and its mRNA is induced in normal human lymphocytes when stimulated by mitogens (Gómez-Márquez and Rodríguez 1998). Spike transfection upregulates prothymosin α (6.4-fold), compatible with extensive studies establishing that viral infections transmit proliferation signals to otherwise terminally differentiated cells (Vareli et al. 1995). Cells that overexpress this transcript also exhibit higher levels of histone-H1 depleted chromatin, by interacting with histone H1, leading to the decondensation and remodeling of chromatin fibers. Such changes in chromatin organization are important in essential cellular processes such as recombination, replication, transcription and chromosome packaging. Microarray analysis demonstrated upregulation of the histone H1 transcript concurrent with that of prothymosin α.
Another notable upregulated transcript (~2.2-fold) is histone deacetylase 5 (HDAC5) which also plays an important role in chromatin remodeling. Whether a cell is permissive for viral infection may be dependent on the state of cellular differentiation. For example, permissiveness for human cytomegalovirus (HCMV) infection is linked to repression of the viral major immediate early promoter (MIEP). In HCMV, monocytes represent a site of viral latency in HCMV carriers, and reactivated virus is only observed upon differentiation into macrophages. Histone deacetylases (HDACs) are involved in MIEP repression in nonpermissive cells, since inhibition of HDACs induces viral permissiveness and increases MIEP activity (Murphy et al. 2002). HDACs modulate MIEP transcriptional activity by affecting the acetylation of transcriptional factors as well as histones. A related protein, HDAC6, regulates formation of the HIV-mediated fusion pore. Binding of the HIV gp120 protein to CD4-positive permissive cells increases the level of acetylated α-tubulin. Overexpression of active HDAC6 inhibits tubulin acetylation, thus preventing HIV-1 envelope-dependent cell fusion as well as infection without affecting the expression of other HIV-1 receptors (Valenzuela-Fernández et al. 2005).
5.3 Cytoskeletal Genes
SARS-CoV-infected cells exhibit diminished expression of genes associated with the maintenance of cytoskeletal structure. Many cellular processes depend on cytoskeletal rearrangements. Villin 1 is a structural protein regulated by increased intracellular calcium that in turn binds to actin, and contributes to the formation of microvilli in the small intestine. Cytokeratins are part of the family of keratins consisting of about 20 different proteins that comprise intermediate filaments in almost all epithelial cells. These filaments form the internal infrastructure of cells and are vital for maintaining cellular integrity. Disruption of such features affects cellular flexibility, tensile strength and interaction with other cellular components (Glass et al. 2006). Infection of animal cells by a number of viruses generally results in CPE. In lytic viral infections, there is a profound impairment of cell structural integrity, and inhibition of host protein synthesis related to widespread destruction of the intermediate filament network, e.g. proteolysis of cytokeratins K7 and K18 by the adenovirus late-acting L3 proteinase (Zhang and Schneider 1994). The collapse of this network (including vimentin and lamin), coupled with suppression of new keratin synthesis to restore the damaged filament network, contributes to CPE. In SARS-CoV infection, these microtubule networks may become disrupted, making it conducive for subsequent viral release, partially attributed to the spike protein.
Following SARS-CoV spike transfection, the mRNA encoding zyxin is greatly upregulated ~45-fold. Zyxin enhances cell motility through rearrangement of the actin cytoskeleton, and it facilitates the formation of the molecular complex that promotes the assembly of actin-rich structures targeted particularly to the inner face of the plasma membrane (Beckerle 1997). Correct positioning of zyxin within the cell is also critical for its physiological function. Mislocation of zyxin affects cell migration and spreading, influences behavior of the cell edge, and causes irregular distribution of important proteins that promote actin assembly (Drees et al. 1999). Furthermore, the β-actin transcript is upregulated ~7.7-fold, suggesting the complex interplay between the various elements that contribute to structural integrity within the cell.
5.4 Trafficking and Transport Genes
The vacuolar protein sorting 4A gene transcript is upregulated upon spike transfection. In many viral infections, viral proteins must recruit a variety of cellular factors, including members of the vacuolar protein sorting family, for subsequent viral budding and egress, failing which the viral particles would be nearly assembled but remain tethered to the cell surface (Sherer et al. 2003).
Another noteworthy upregulated transcript (~5-fold) is ICln mRNA which encodes a 42-kDa chloride channel regulatory protein essential in regulating cell volume, and which interacts with the specific platelet integrin αIIbβ3 through the KVGFFKR motif on platelet membranes (Larkin et al. 2004). Treatment with the specific channel protein inhibitory agent, acyclovir, culminates in inhibition of platelet aggregation and integrin activation. It is also noteworthy that the SARS-CoV protein 3a upregulates fibrinogen expression in lung epithelial cells, congruent with observations of SARS patients with thrombocytopenia, and suggesting dysfunctional coagulation and activation of the fibrinogen pathway (Tan et al. 2005).
6 Conclusions and Future Prospects
This is the first report documenting SARS-CoV expression in the human HL-CZ cell line that triggers a cascade of host transcriptional responses involving a wide repertoire of genes belonging to key functional classes including inflammation, immunity, cell signaling and trafficking. Intriguingly, most of the transcriptional factors upregulated in response to spike transfection are zinc finger proteins. In contrast to live SARS-CoV infection, this study provides comparative insights into the host transcriptomic effects of the spike protein when expressed alone in relevant human cells, thus shedding light on its specific roles during viral infection. These data indicate that SARS-CoV spike expression significantly modifies the regulation of numerous genes in HL-CZ cells, reiterate its importance in SARS pathogenesis, and further justify it as a key target for vaccine development. Such studies can enhance our understanding of the “infectomics” of viral infections in different human and mammalian cell types. More in-depth analyses of the individual genes may be conducted by quantitative real-time RT-PCR to verify the microarray data, and by using gene-knockout animal models to further explore their specific roles in SARS-CoV infection in vivo.
References
Babcock GJ, Esshaki DJ, Thomas WD, Ambrosino DM (2004) Amino acids 270 to 510 of the severe acute respiratory syndrome coronavirus spike protein are required for interaction with receptor. J Virol 78:4552–4560
Beckerle MC (1997) Zyxin: zinc fingers at sites of cell adhesion. Bioessays 19:949–957
Chang YJ, Liu CY, Chiang BL, Chao YC, Chen CC (2004) Induction of IL-8 release in lung cells via activator protein-1 by recombinant baculovirus displaying severe acute respiratory syndrome-coronavirus spike proteins: identification of two functional regions. J Immunol 173:7602–7614
Chow VT, Leong WF (2008) Severe acute respiratory syndrome coronavirus induces differential host gene expression responses associated with pathogenesis. In: Yang D (ed) RNA viruses: host gene response to infection. World Scientific, New Jersey, pp 295–320
Dove B, Brooks G, Bicknell K, Wurm T, Hiscox JA (2006) Cell cycle perturbations induced by infection with the coronavirus infectious bronchitis virus and their effect on virus replication. J Virol 80:4147–4156
Drees EB, Andrews KM, Beckerle MC (1999) Molecular dissection of zyxin function reveals its involvement in cell motility. J Cell Biol 147:1549–1559
Du L, He Y, Zhou Y, Liu S, Zheng BJ, Jiang S (2009) The spike protein of SARS-CoV – a target for vaccine and therapeutic development. Nat Rev Microbiol 7:226–236
Glass C, Kim KH, Fuchs E (2006) Sequence and expression of a human type II mesothelial keratin. J Cell Biol 101:2366–2373
Gómez-Márquez J, Rodríguez P (1998) Prothymosin α is a chromatin-remodelling protein in mammalian cells. Biochem J 333:1–3
Groneberg DA, Poutanen SM, Low DE, Lode H, Welte T, Zabel P (2005) Treatment and vaccines for severe acute respiratory syndrome. Lancet Infect Dis 5:147–155
Han DP, Penn-Nicholson A, Cho MW (2006) Identification of critical determinants on ACE2 for SARS-CoV entry and development of a potent entry inhibitor. Virology 350:15–25
He Y, Zhou Y, Liu S, Kou Z, Li W, Farzan M, Jiang S (2004a) Receptor-binding domain of SARS-CoV spike protein induces highly potent neutralizing antibodies: implication for developing subunit vaccine. Biochem Biophys Res Commun 324:773–781
He Y, Zhou Y, Siddiqui P, Jiang S (2004b) Inactivated SARS-CoV vaccine elicits high titers of spike protein-specific antibodies that block receptor binding and virus entry. Biochem Biophys Res Commun 325:445–452
Hoogenboom HR (2002) Overview of antibody phage-display technology and its applications. Methods Mol Biol 178:1–37
Jeffers SA, Tusell SM, Gillim-Ross L, Hemmila EM, Achenbach JE, Babcock GJ, Thomas WD Jr, Thackray LB, Young MD, Mason RJ, Ambrosino DM, Wentworth DE, Demartini JC, Holmes KV (2004) CD209L (L-SIGN) is a receptor for severe acute respiratory syndrome coronavirus. Proc Natl Acad Sci USA 101:15748–15753
Lai CC, Jou MJ, Huang SY, Li SW, Wan L, Tsai FJ, Lin CW (2007) Proteomic analysis of up-regulated proteins in human promonocyte cells expressing severe acute respiratory syndrome coronavirus 3C-like protease. Proteomics 7:1446–1460
Larkin D, Murphy D, Reilly DF, Cahill M, Sattler E, Harriott P, Cahill DJ, Moran N (2004) ICln, a novel integrin αIIbβ3-associated protein, functionally regulates platelet activation. J Biol Chem 279:27286–27293
Leong WF, Chow VT (2006) Transcriptomic and proteomic analyses of rhabdomyosarcoma cells reveal differential cellular gene expression in response to enterovirus 71 infection. Cell Microbiol 8:565–580
Leong WF, Tan HC, Ooi EE, Koh DR, Chow VT (2005) Microarray and real time RT-PCR analyses of differential human gene expression patterns induced by severe acute respiratory syndrome (SARS) coronavirus infection of Vero E6 cells. Microbes Infect 7:248–259
Leth-Larsen R, Zhong F, Chow VT, Holmskov U, Lu J (2007) The SARS coronavirus spike glycoprotein is selectively recognized by lung surfactant protein D and activates macrophages. Immunobiology 212:201–211
Li W, Moore MJ, Vasilieva N, Sui J, Wong SK, Berne MA, Somasundaran M, Sullivan JL, Luzuriaga K, Greenough TC, Choe H, Farzan M (2003) Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 426:450–454
Liang WS, Maddukuri A, Teslovich TM, de la Fuente C, Agbottah E, Dadgar S, Kehn K, Hautaniemi S, Purnfery A, Stephan DA, Kashanchi F (2005) Therapeutic targets for HIV-1 infection in the host proteome. Retrovirology 2:20
Liew KJ, Chow VT (2006) Microarray and real-time RT-PCR analyses of a novel set of differentially expressed human genes in ECV304 endothelial-like cells infected with dengue virus type 2. J Virol Methods 131:47–57
Manopo I, Lu L, He Q, Chee LL, Chan SW, Kwang J (2005) Evaluation of a safe and sensitive Spike protein-based immunofluorescence assay for the detection of antibody responses to SARS-CoV. J Immunol Methods 296:37–44
Murphy JC, Fischle W, Verdin E, Sinclair JH (2002) Control of cytomegalovirus lytic gene expression by histone acetylation. EMBO J 21:1112–1120
Pini A, Bracci L (2000) Phage display of antibody fragments. Curr Protein Pept Sci 1:155–169
Poh WP, Narasaraju T, Pereira NA, Zhong F, Phoon MC, Macary PA, Wong SH, Lu J, Koh DR, Chow VT (2009) Characterization of cytotoxic T-lymphocyte epitopes and immune responses to SARS coronavirus spike DNA vaccine expressing the RGD-integrin-binding motif. J Med Virol 81:1131–1139
Poon LL, Guan Y, Nicholls JM, Yuen KY, Peiris JS (2004) The aetiology, origins, and diagnosis of severe acute respiratory syndrome. Lancet Infect Dis Rev 4:663–671
Satija N, Lal SK (2007) The molecular biology of SARS coronavirus. Ann N Y Acad Sci 1102:26–38
Sherer NM, Lehmann MJ, Jemenez-Soto LF, Ingmundson A, Horner SM, Cicchetti G, Allen PG, Pypaert M, Cunningham JM, Mothes W (2003) Visualization of retroviral replication in living cells reveals budding into multivesicular bodies. Traffic 4:785–801
Shi Z, Hu Z (2008) A review of studies on animal reservoirs of the SARS coronavirus. Virus Res 133:74–87
Shih YP, Chen CY, Liu SJ, Chen KH, Lee YM, Chao YC, Chen YM (2006) Identifying epitopes responsible for neutralizing antibody and DC-SIGN binding on the spike glycoprotein of the severe acute respiratory syndrome coronavirus. J Virol 80:10315–10324
Simmons G, Reeves JD, Rennekamp AJ, Amberg SM, Piefer AJ, Bates P (2004) Characterization of severe acute respiratory syndrome-associated coronavirus (SARS-CoV) spike glycoprotein-mediated viral entry. Proc Natl Acad Sci 101:4240–4245
Tan YJ, Tham PY, Chan DZ, Chou CF, Shen S, Fielding BC, Tan TH, Lim SG, Hong W (2005) The severe acute respiratory syndrome coronavirus 3a protein up-regulates expression of fibrinogen in lung epithelial cells. J Virol 79:10083–10087
ter Meulen J, Bakker AB, van den Brink EN, Weverling GJ, Martina BE, Haagmans BL, Kuiken T, de Kruif J, Preiser W, Spaan W, Gelderblom HR, Goudsmit J, Osterhaus AD (2004) Human monoclonal antibody as prophylaxis for SARS coronavirus infection in ferrets. Lancet 363:2139–2141
ter Meulen M, van den Brink EN, Poon LL, Marissen WE, Leung CS, Cox F, Cheung CY, Bakker AQ, Bogaards JA, van Deventer E, Preiser W, Doerr HW, Chow VT, de Kruif J, Peiris JS, Goudsmit J (2006) Human monoclonal antibody combination against SARS coronavirus: synergy and coverage of escape mutants. PLoS Med 3:e237
Thiel V, Weber F (2008) Interferon and cytokine responses to SARS-coronavirus infection. Cytokine Growth Factor Rev 19:121–132
Valenzuela-Fernández A, Álvarez S, Gordon-Alonso M, Barrero M, Ursa Á, Cabrero JR, Fernández G, Naranjo-Suárez S, Yáñez-Mo M, Serrador JM, Muñoz-Fernández MÁ, Sánchez Madrid F (2005) Histone deacetylase 6 regulates human immunodeficiency virus type 1 infection. Mol Biol Cell 16:5445–5454
van den Brink EN, Ter Meulen J, Cox F, Jongeneelen MA, Thijsse A, Throsby M, Marissen WE, Rood PM, Bakker AB, Gelderblom HR, Martina BE, Osterhaus AD, Preiser W, Doerr HW, de Kruif J, Goudsmit J (2005) Molecular and biological characterization of human monoclonal antibodies binding to the spike and nucleocapsid proteins of severe acute respiratory syndrome coronavirus. J Virol 79:1635–1644
Vareli K, Lazaridis MF, Tsolas O (1995) Prothymosin α mRNA levels vary with c-myc expression during tissue proliferation, viral infection and heat shock. FEBS Lett 371:337–340
Versteeg GA, van de Nes PS, Bredenbeek PJ, Spaan WJ (2007) The coronavirus spike protein induces endoplasmic reticulum stress and upregulation of intracellular chemokine mRNA concentrations. J Virol 81:10981–10990
Wang W, Ye L, Ye L, Li B, Gao B, Zeng Y, Kong L, Fang X, Zheng H, Wu Z, She Y (2007) Up-regulation of IL-6 and TNF-alpha induced by SARS-coronavirus spike protein in murine macrophages via NF-kappaB pathway. Virus Res 128:1–8
Zhang Y, Schneider RJ (1994) Adenovirus inhibition of cell translation facilitates release of virus particles and enhances degradation of the cytokeratin network. J Virol 68:2544–2555
Zhang H, Wang G, Li J, Nie Y, Shi X, Lian G, Wang W, Yin X, Zhao Y, Qu X, Ding M, Deng H (2004) Identification of an antigenic determinant on the S2 domain of the severe acute respiratory syndrome coronavirus spike glycoprotein capable of inducing neutralizing antibodies. J Virol 78:6938–6945
Zhou T, Wang H, Luo D, Rowe T, Wang Z, Hogan RJ, Qiu S, Bunzel RJ, Huang G, Mishra V, Voss TG, Kimberly R, Luo M (2004) An exposed domain in the severe acute respiratory syndrome coronavirus spike protein induces neutralizing antibodies. J Virol 78:7217–7226
Acknowledgments
The authors thank J. de Kruif, H.N. Leong, B. Liu and W.M. Yeo for their valuable assistance. This study was funded by the Biomedical Research Council, Singapore and the Microbiology Vaccine Initiative, National University of Singapore.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2010 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Narasaraju, T., Soong, P.L., ter Meulen, J., Goudsmit, J., Chow, V.T.K. (2010). SARS Coronavirus Spike Protein Expression in HL-CZ Human Promonocytic Cells: Monoclonal Antibody and Cellular Transcriptomic Analyses. In: Lal, S. (eds) Molecular Biology of the SARS-Coronavirus. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-03683-5_18
Download citation
DOI: https://doi.org/10.1007/978-3-642-03683-5_18
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-03682-8
Online ISBN: 978-3-642-03683-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)