Abstract
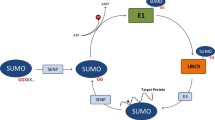
Recent evidence suggests that SUMOylation of proteins plays a key regulatory role in the assembly and dis-assembly of nuclear sub-compartments, and may repress transcription by modifying chromatin. Determining whether a protein contains a SUMOylation site or not thus provides essential clues about a substrate’s intra-nuclear spatial association and function.
Previous SUMOylation predictors are largely based on a degenerate and functionally unreliable consensus motif description, not rendering satisfactory accuracy to confidently map the extent of this essential class of regulatory modifications. This paper embarks on an exploration of predictive dependencies among SUMOylation site amino acids, non-local and structural properties (including secondary structure, solvent accessibility and evolutionary profiles).
An extensive examination of two main machine learning paradigms, Support-Vector-Machine and Bidirectional Recurrent Neural Networks, demonstrates that (1) with careful attention to generalization issues both methods achieve comparable performance and, that (2) local features enable best generalization, with structural features having little to no impact. The predictive model for SUMOylation sites based on the primary protein sequence achieves an area under the ROC of 0.92 using 5-fold cross-validation, and 96% accuracy on an independent hold-out test set. However, similar to other predictors, the new predictor is unable to generalize beyond the simple consensus motif.
Chapter PDF
Similar content being viewed by others
Keywords
- Radial Basis Function
- Consensus Motif
- Radial Basis Function Kernel
- Sequence Logo
- Relative Solvent Accessibility
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
References
Altschul, S.F., Madden, T.L., Schäffer, A.A., Zhang, J., Zhang, Z., Miller, W., Lipman, D.J.: Gapped bLAST and pSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25(17), 3389–3402 (1997)
Baldi, P., Brunak, S., Frasconi, P., Soda, G., Pollastri, G.: Exploiting the past and the future in protein secondary structure prediction. Bioinformatics 15, 937–946 (1999)
Bodén, M., Yuan, Z., Bailey, T.L.: Prediction of protein continuum secondary structure with probabilistic models based on NMR solved structures. BMC Bioinformatics 7, 68 (2006)
Crooks, G.E., Hon, G., Chandonia, J.-M., Brenner, S.E.: WebLogo: a sequence logo generator. Genome Res. 14(6), 1188–1190 (2004)
Dorval, V., Fraser, P.E.: SUMO on the road to neurodegeneration. Biochim Biophys Acta. 1773(6), 694–706 (2007)
Fawcett, T.: Roc graphs: Notes and practical considerations for researchers. Machine Learning (2004)
Haussler, D.: Convolution kernels on discrete structures. Technical Report UCSC-CRL-99-10, Department of Computer Science, University of California, Santa Cruz, CA 95064 (1999)
Hay, R.T.: SUMO: a history of modification. Mol Cell 18(1), 1–12 (2005)
Heun, P.: SUMOrganization of the nucleus. Curr Opin Cell Biol. 19(3), 350–355 (2007)
Nathan, D., Ingvarsdottir, K., Sterner, D.E., Bylebyl, G.R., Dokmanovic, M., Dorsey, J.A., Whelan, K.A., Krsmanovic, M., Lane, W.S., Meluh, P.B., Johnson, E.S., Berger, S.L.: Histone sumoylation is a negative regulator in saccharomyces cerevisiae and shows dynamic interplay with positive-acting histone modifications. Genes Dev. 20(8), 966–976 (2006)
Rodriguez, M.S., Dargemont, C., Hay, R.T.: SUMO-1 conjugation in vivo requires both a consensus modification motif and nuclear targeting. J Biol Chem. 276(16), 12654–12659 (2001)
Saigo, H., Vert, J., Ueda, N., Akutsu, T.: Protein homology detection using string alignment kernels. Bioinformatics 20(11), 1682–1689 (2004)
Schwartz, D., Gygi, S.P.: An iterative statistical approach to the identification of protein phosphorylation motifs from large-scale data sets. Nat Biotechnol. 23(11), 1391–1398 (2005)
Shen, T.H., Lin, H.-K., Scaglioni, P.P., Yung, T.M., Pandolfi, P.P.: The mechanisms of PML-nuclear body formation. Mol. Cell 24(3), 331–339 (2006)
Vapnik, V.: Statistical learning theory. Wiley, Chichester (1998)
Xu, J., He, Y., Qiang, B., Yuan, J., Peng, X., Pan, X.: A novel method for high accuracy sumoylation site prediction from protein sequences. BMC Bioinformatics 9, 8 (2008)
Xue, Y., Zhou, F., Fu, C., Xu, Y., Yao, X.: SUMOsp: a web server for sumoylation site prediction. Nucleic Acids Res. 34, W254–W257 (2006)
Yuan, Z., Huang, B.: Prediction of protein accessible surface areas by support vector regression. Proteins 57(3), 558–564 (2004)
Zhou, F., Xue, Y., Chen, G., Yao, X.: GPS: a novel group-based phosphorylation predicting and scoring method. Biochem. Biophys. Res. Commun. 325(4), 1443–1448 (2004)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2008 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Bauer, D.C., Buske, F.A., Bodén, M. (2008). Predicting SUMOylation Sites. In: Chetty, M., Ngom, A., Ahmad, S. (eds) Pattern Recognition in Bioinformatics. PRIB 2008. Lecture Notes in Computer Science(), vol 5265. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-540-88436-1_3
Download citation
DOI: https://doi.org/10.1007/978-3-540-88436-1_3
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-88434-7
Online ISBN: 978-3-540-88436-1
eBook Packages: Computer ScienceComputer Science (R0)