Abstract
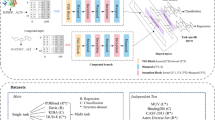
Protein-protein interactions (PPIs) mediate myriad biological functions. Estimating the binding affinity for PPIs help us to understand the underlying molecular recognition mechanism. In this work, we utilized deep learning approach to discriminate protein-protein complexes based on their binding affinity. We setup a database of 464 protein-protein complexes along with their experimental binding affinities and developed a deep learning based binary classification model, which showed an accuracy of 81.75% using 5-fold cross-validation. Furthermore, we refined the method for predicting the binding affinity of protein-protein complexes using a large set of complexes (PPA-Pred2). It could predict the binding affinity of the training set and blind test set with a mean absolute error of 1.24 kcal/mol and 1.31 kcal/mol, respectively. We suggest that our methods could serve as efficient tools to study PPIs and provide crucial insights about the underlying mechanism for the molecular recognition process.
R. Nikam and K. Yugandhar—Contributed equally to this work.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Bahadur, R.P., Chakrabarti, P., Rodier, F., Janin, J.: A dissection of specific and non-specific protein-protein interfaces. J. Mol. Biol. 336, 943–955 (2004)
Yugandhar, K., Gromiha, M.M.: Analysis of protein-protein interaction networks based on binding affinity. Curr. Prot. Pept. Sci. 17, 72–81 (2016)
Moal, I.H., Agius, R., Bates, P.A.: Protein-protein binding affinity prediction on a diverse set of structures. Bioinformatics 27, 3002–3009 (2011)
Yugandhar, K., Michael Gromiha, M.: Feature selection and classification of protein-protein complexes based on their binding affinities using machine learning approaches. Proteins 82(9), 2088–2096 (2014)
Liu, Z., Li, Y., Han, L., Li, J., Liu, J., Zhao, Z., Nie, W., Liu, Y., Wang, R.: PDB-wide collection of binding data: current status of the PDBbind database. Bioinformatics 31, 405–412 (2015)
Kastritis, P.L., Bonvin, A.M.J.J.: On the binding affinity of macromolecular interactions: daring to ask why proteins interact. J. R. Soc. Interface 10, 20120835 (2013)
Chen, J., Sawyer, N., Regan, L.: Protein-protein interactions: general trends in the relationship between binding affinity and interfacial buried surface area. Protein Sci. 22, 510–515 (2013)
Lomax, J.E., Christopher, M.B., Chang, A., George, N.P.J.: Functional evolution of ribonuclease inhibitor: insights from birds and reptiles. J. Mol. Biol. 426, 3041–3056 (2014)
Spencer, A.L., Bagai, I., Becker, D.F., Zuiderweg, E.R., Ragsdale, S.W.: Protein-protein interactions in the mammalian heme degradation pathway heme oxygenase-2, cytochrome p450 reductase, and biliverdin reductase. J. Biol. Chem. 289, 29836–29858 (2014)
Vreven, T., Iain, H.M., Anna, V., Brian, G.P.: Updates to the integrated protein-protein interaction benchmarks: docking benchmark version 5 and affinity benchmark version 2. J. Mol. Biol. 427, 3031–3041 (2015)
Wang, G., Dunbrack, R.L.J.: PISCES: a protein sequence culling server. Bioinformatics 19, 1589–1591 (2003)
Chollet, F.: “Keras” (2015)
Saraboji, K., Gromiha, M.M., Ponnuswamy, M.N.: Average assignment method for predicting the stability of protein mutants. Biopolymers 82, 80–92 (2006)
Yugandhar, K., Michael Gromiha, M.: Protein-protein binding affinity prediction from the amino acid sequence. Bioinformatics 30, 3583–3589 (2014)
Kawashima, S., Pokarowski, P., Pokarowska, M., Kolinski, A., Katayama, T., Kanehisa, M.: AAindex: amino acid index database, progress report 2008. Nucleic Acids Res. 36, D202–D205 (2008)
Ofran, Y., Rost, B.: Interaction sites identified from sequence. Bioinformatics 23, e13–e16 (2007)
Gromiha, M.M.: A statistical model for predicting protein folding rates from amino acid sequence with structural class information. J. Chem. Inf. Model. 45, 494–501 (2005)
Gromiha, M.M., Saranya, N., Selvaraj, S., Jayaram, B., Fukui, K.: Sequence and structural features of binding site residues in protein-protein complexes: comparison with protein-nucleic acid complexes. Proteome Sci. 9, S13 (2011)
Li, W., Hamill, S.J., Hemmings, A.M., Moore, G.R., James, R., Kleanthous, C.: Dual recognition and the role of specificity-determining residues in colicin E9 DNase-immunity protein interactions. Biochemistry 37, 11771–11779 (1998)
Eathiraj, S., Pan, X., Ritacco, C., Lambright, D.G.: Structural basis of family-wide Rab GTPase recognition by rabenosyn-5. Nature 436, 415–419 (2005)
Maenaka, K., van der Merwe, P.A., Stuart, D.I., Jones, E.Y., Sondermann, P.: The human low affinity Fc receptors IIa, IIb, and IIIbind IgG with fast kinetics and distinct thermodynamic properties. J. Biol. Chem. 276, 44898–44904 (2001)
Blanco, M.A., Tatiana, P., Vincenzo, M., Mauro, M., Christopher, J.R.: Protein-protein interactions in dilute to concentrated solutions: α-Chymotrypsinogen in acidic conditions. J. Phys. Chem. B 118, 5817–5831 (2014)
Acknowledgements
We thank Indian Institute of Technology Madras and the High-Performance Computing Environment (HPCE) for computational facilities. The work was partially supported by the Department of Science and Technology, Government of India (DST/INT/SWD/P-05/2016) and of the Swedish Research Council.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this paper
Cite this paper
Nikam, R., Yugandhar, K., Michael Gromiha, M. (2018). Discrimination and Prediction of Protein-Protein Binding Affinity Using Deep Learning Approach. In: Huang, DS., Jo, KH., Zhang, XL. (eds) Intelligent Computing Theories and Application. ICIC 2018. Lecture Notes in Computer Science(), vol 10955. Springer, Cham. https://doi.org/10.1007/978-3-319-95933-7_89
Download citation
DOI: https://doi.org/10.1007/978-3-319-95933-7_89
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-95932-0
Online ISBN: 978-3-319-95933-7
eBook Packages: Computer ScienceComputer Science (R0)