Abstract
Ontologies are a basic tool to formalize and share knowledge. However, very often the conceptualization of a specific domain depends on the particular user’s needs. We propose a methodology to perform user-centric ontology population that efficiently includes human-in-the-loop at each step. Given the existence of suitable target ontologies, our methodology supports the alignment of concepts in the user’s conceptualization with concepts of the target ontologies, using a novel hierarchical classification approach. Our methodology also helps the user to build, alter and grow their initial conceptualization, exploiting both the target ontologies and new facts extracted from unstructured data. We evaluate our approach on a real-world example in the healthcare domain, in which adverse phrases for drug reactions, as extracted from user blogs, are aligned with MedDRA concepts. The evaluation shows that our approach has high efficacy in assisting the user to both build the initial ontology (\({{\mathrm{{\textit{HITS}\,@10}}}}\) up to 99.5%) and to maintain it (\({{\mathrm{{\textit{HITS}\,@10}}}}\) up to 99.1%).
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
Maintaining data in a structured and machine-readable form allows easy data sharing between humans and software agents, and also enables other tasks related to data handling, including data analysis and data reuse to name a few. In some domains where the majority of data is only available as unstructured text, extracting such structured knowledge constitutes a crucial step.
Many available tools extract items of interest (mainly in the form of named entities) from free text. The extracted instances can be maintained at varying degrees of complexity: as simple as flat dictionaries, or as rich as a structured concept organization in the form of an ontology.
While ontologies are an excellent means to formalize and share knowledge, it is rare to have a single unique conceptualization of a domain: depending on the field, on the task at hand, and on the specific user, the best representation can vary, in some cases extensively. It is well known that human annotation tasks intrinsically carry a level of disagreement among annotators, regardless of their level of domain expertise [1, 43]. While it is important to maintain the user conceptualization of the domain, connecting it to any existing and well-defined ontology in the field is one of the paramount principles of the Semantic Web movement.
The challenge is to achieve the right balance between the user conceptualization and available knowledge, enabling the population and maintenance of the user ontology with both relevant facts already available in structured form (e.g. other ontologies), as well as the new extracted facts from unstructured data.
Most of the ontology population solutions proposed in the literature focus on solutions for open domain problems, where it is not crucial to achieve perfect performance. However, in many domains near-perfection is required. For example, many biomedical applications have near 0% error tolerance, despite datasets full of uncertainty, incompleteness and noise. Furthermore, some problems in the medical domain are quite challenging, making the application of fully automated models difficult, or at least raising questions on the quality of results. Consequently, efficiently including a domain expert as an integral part of the system not only greatly enhances the knowledge discovery process pipeline [14, 15], but can in certain circumstances be legally or ethically required.
We propose a methodology to perform user-centric ontology population that efficiently includes human-in-the-loop at each step: the user is assisted in building, connecting and maintaining their conceptualization of the domain, while taking advantage of any already available ontology.
Given initial user data comprising a number of concepts and their initial instances, and assuming the existence of candidate ontologies for the alignment, available either publicly (the Linked Open Data cloud) or within the enterprise, our methodology supports three main steps: (i) selecting the relevant ones (target ontologies); (ii) aligning the concepts in the user’s conceptualization with concepts of the target ontologies, using a novel hierarchical classification approach; (iii) assisting the user to build, change, and grow their initial ontology, by (respectively) creating new concepts, splitting or merging concepts, and adding new instances to each concept, all via exploitation of both target ontologies and new facts extracted from unstructured data. Each step includes human-in-the-loop. That is, the methodology is designed to efficiently assist the user rather than fully automate the process.
The contribution of this work is threefold. First, our approach does not require the user to have any expertise with the Semantic Web:Footnote 1 the input data is a set of coherent concepts defined with only some initial instances that can be provided as a simple populated taxonomy, or even as disconnected groups. These instances are used to identify available target ontologies. Second, we propose a novel hierarchical classification method that allows mapping the user data to the target ontology. To the best of our knowledge, this is the first method for ontology population that builds hierarchical classification models that are dynamically refined and based on user interaction. Finally, the method does not require any training material (since it only exploits the target ontology), nor any NLP processing or linguistic features. Therefore, the method is also potentially flexible with respect to different domains and languages.
The main advantage of our approach is that the user has full control of their level of involvement, with a trade-off on the accuracy of results, so that the more precise and granular the representation needs to be, the more they can be in the loop. We test the approach on a real-world example in the setting of Adverse Drug Reactions. Starting from a concepts representation extracted from user medical blogs,Footnote 2 we identify an available ontology, namely MedDRA [2], within the enterprise knowledge base, and map the user’s initial concepts to the target ontology. In the experiment, a user concept is a group of coherent phrases, e.g. teeth grinding, teeth clenching, clench my teeth, jaw clenching, clinching my jaw, which we help to align to concepts in MedDRA, in this case “Bruxism”. We show that we can assist the user with the alignment with \({{\mathrm{{\textit{HITS}\,@10}}}}= 99.5\%\) on the most general level of the ontology and \({{\mathrm{{\textit{HITS}\,@10}}}}= 86.5\%\) on the most granular level of the ontology. We also evaluate the approach for adding new instances, achieving \({{\mathrm{{\textit{HITS}\,@10}}}}= 99.1\%\) on the most general level of the ontology and \({{\mathrm{{\textit{HITS}\,@10}}}}= 91.27\%\) on the most granular level of the ontology.
In the following, we give an overview of related work in Sect. 2; we formally define the problem of user-centric ontology population and describe our solution in Sect. 3; and we test our solution in the medical domain, Sect. 4.
2 State of the Art
There is a vast literature devoted to ontology population from text, with many established initiatives to foster research on the topic, such as the Knowledge Base Population task at TAC,Footnote 3 the TREC Knowledge Base Acceleration track,Footnote 4 and the Open Knowledge Extraction Challenge [24]. In these initiatives, systems are compared on the basis of recognizing individuals belonging to a few selected ontology classes, spanning from the common Person, Place and Organization [36], to more specific classes such as Facility, Weapon, Vehicle [8], Role [24] or Drug [31], among others.
FRED and Framester [11] and [10] are an established example of a comprehensive solution to the problem. The tools transform text in an internal ontology representation and then attempt to align it with available Linked Data. FRED is a general purpose machine reader, mostly based on core NLP tools, which can potentially process text from any domain and in many different languages (bounded to the availability of NLP components). In the same direction, there is a plethora of tools for automatically detecting named entities in free text and aligning them to a predefined knowledge base, i.e., Spotlight [19], X-Lisa [44], Babelfy [21]. However, all these tools are able to identify only instances that already exist in a knowledge base.
Some of the earliest approaches for ontology population from text are based on pattern matching, string similarity functions, and external glossaries and knowledge bases. Velardi et al. [37, 38] develop OntoLearn which is one of the first tools for learning and populating ontologies from text. The approach heavily uses NLP parsers, pattern matching, and external glossaries, in combination with human assistance. Similar approaches are presented in [3, 18]. Cimiano and Völker [4] describe an unsupervised approach, called Class-Word, for ontology population based on vector-feature similarity between each concept and a term to be classified. The feature vectors are generated from the text corpus. The approach assumes that the entity and the concept usually appear together in the same sentences. The approach is extended in Tanev and Magnini [35], called Class-Example, which learns a classification model from a set of classified terms, exploiting lexico-syntactic features. They upgrade the previous approach by adding features extracted from dependency parse trees. Giuliano and Gliozzo [12] propose an approach that is based on the assumption that entities that occur in similar contexts belong to the same concept(s), and so it counts the shared n-grams in the context of the entities. An overview of pattern-based approaches is given in a survey by Petasise et al. [25].
Several works use machine learning for ontology population. HYENA [42] and FIGER [17] are two examples of fine-grained multi-label classifiers for named entity types based on hierarchical taxonomies derived from YAGO. Ling and Weld [17] also release the benchmark dataset annotated with 112 classes from YAGO. Typically, the models use standard NLP features extracted from text, or more sophisticated features such as type relational phrases: either their type signatures and disjointness constraints [23], or type correlation based on co-occurring entities [27].
Many approaches for ontology population are based on word and graph embedding models. WSABIE [41] adopts weighted approximate pairwise loss to learn embeddings of features and types in a common feature space. Entities that share the same type appear close to each other in the embedded space. Similarly, FIGMENT [40] proposes a combination of global and context model, where the global model performs global embedding over the whole corpus using multilayer perceptron, while the context model focuses on small context windows sizes. Ristoski et al. [29] use standard word embeddings and graph embeddings to align instances extracted from the Common CrawlFootnote 5 to the DBpedia ontology.
The use of deep learning models has also been explored for this task. Dong et al. [9] propose the first deep learning architecture for entity typing. The architecture consist of two models. The mention model uses recurrent neural networks to recursively obtain the vector representation of an entity mention from the words it contains. The context model, on the other hand, employs multilayer perceptrons to obtain the hidden representation for contextual information of a mention. The approach is evaluated on 22 general types from DBpedia. Shimoaka et al. [32, 33] propose a very simple neural network, using averaging encoder, LSTM encoder, and attentive encoder, for computing context representations. Similarly, Yaghoobzadeh et al. [39] propose a convolutional neural network for entity typing. Both approaches are evaluated on 112 entity types. Murty et al. [22] present TypeNet, a dataset of entity types consisting of over 1941 types organized in a hierarchy, on which they train several neural models for entity typing.
None of these methods take into consideration the hierarchical structure of the ontology, and for all of them the number of types is relatively small and within a general open domain. In this paper, we present an approach that exploits such hierarchical structure, which we evaluate on an ontology with significantly more concepts than related work.
3 Approach
Input. The input of our approach is a set of example entities within a particular domain, usually extracted from a coherent textual corpus. Given a textual corpus, we assume there is a domain entity extractor (specifically we used SPOT [5]) that produces the set of relevant entities in the corpus \(I_U={i_1, i_2, \ldots , i_n}\). The user refines this set by organizing all instances in \(I_U\) in concepts. The result is a finite set of user-defined concepts \(C_u={c_{u1}, c_{u2},\ldots , c_{un}}\) where each concept contains at least one instance. Using the user-defined conceptualization \(C_u\), we scout for ontology candidates that can fit the user data.
Alignment. After a target ontology \(C_T\) is selected, our goal is to align \(C_u\) to \(C_T\). The alignment can be performed at different granularity: given the depth L (or number of levels from root to leaves) of the target ontology, the user can specify the desired level for the alignment, and our method will use as target concepts all concepts \(l_{CT} = {c_{ct_1}, c_{ct_2}, \ldots , c_{ct_n}}\)at level l.
Maintenance. Once the initial alignment is done, we support the maintenance of user knowledge by providing functions for adding new instances, splitting or merging concepts and creating new concepts.
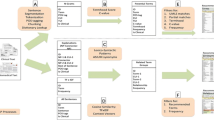
Figure 1 illustrates the overall design of the proposed system.
3.1 Aligning User’s Conceptualization with a Target Ontology
We identify available knowledge using simple collective instance matching between user data and a repository of ontologies. From the repository, either publicly available (such as the Linked Open Data cloud) or proprietary, the top N matching ontologies are presented to the user, who chooses a target ontology \(C_T\). This step can be performed using many alternative state-of-the-art methods [19, 28, 30]; in this work we consider this step as given, and focus on the alignment.
Three novel machine learning approaches are proposed for hierarchical classification, inspired by existing top-down hierarchical classification methods [34]. Considering the user data as “new instances,” the approaches try to identify the concepts in the target ontologies that represent the best match. To do so, we build machine learning models that use the instances of the target ontology as training data (completely unsupervised), and exploit domain-specific word embeddings as features.
In the first solution, we perform a flat hierarchical classification. Given \(C_u\), and considering user-chosen level l of the ontology, we build one classifier with as many classes as concepts at level l, using the leaves of each concept as instances for training the classifier. The architecture is shown in Fig. 2a. This model is rather simple, and it achieves high performance in the upper levels of the hierarchy. However, in the lower levels of the ontology, when the number of classes rapidly increases, the complexity of the model rises, and the performance drops.
The second solution is a top-down model, where we build a local classifier for each parent node. Given \(C_u\), and considering user-chosen level l of the ontology, the approach builds a classifier for each parent node, starting from the top of the hierarchy to level \(l-1\), using all children nodes as classes, and their corresponding leaves as instances to train the model. The architecture is shown in Fig. 2b. This approach can easily cope with a large number of classes in the lower levels of the hierarchy; however, the errors are propagated from the top to the bottom of the ontology.
To circumvent this drawback, we propose a third hierarchical architecture (Fig. 2c), which is a combination of the previous two. Given \(C_u\), and considering user-chosen level l of the ontology, the approach builds (i) a flat classifier for level \(l-1\) of the ontology, and (ii) a classifier for each parent node at level \(l-1\) using the concepts of the l level as classes. This approach is very effective when there is significant difference in the number of nodes between the l and \(l-1\) level of the ontology, for two reasons: (i) the flat classifier performs well on level \(l-1\), which has smaller number of classes; (ii) the per-parent node classifier will only be affected by the errors propagated from the previous level, rather than from the top of the hierarchy.
As classification methods we use standard machine-learning models, i.e., Support Vector Machines, Logistic Regression, and Random Forests, and state-of-the-art deep learning models, i.e., Convolutional Neural Networks.
To perform the final alignment for each user concept \(c_u={i_{u1}, i_{u2},\ldots , i_{un}}\), we classify each instance of \(c_u\) into concepts at target level l of the ontology, and choose the final assignment by majority vote on all instances of \(c_u\), weighting each of them by the class probability distribution returned by the classifier.
The user can define their level of involvement by defining a confidence threshold for each level in the hierarchy: whenever the confidence of the approach is below the given threshold, the system displays top-N candidates to the user who can manually select the desired alignment.
3.2 Ontology Maintenance
Once the alignment has been completed, we provide functions for maintaining the created knowledge base, such as adding instances, adding new concepts and merging/splitting concepts. These functions have been shown to be of a high importance, because of the continuous need to add new data as well as to take into account changes in the user conceptualization over time.
Adding New Instances. When new instances appear, we use the same approaches proposed in Sect. 3.1 to align them to the user’s conceptualization. In this case, the models only consider the concepts defined by the user. When an instance doesn’t fit any of the user-defined concepts, a new concept is added to the user’s conceptualization (with the “Adding New Concepts” function), which is then aligned to the target ontology.
Adding New Concepts. To decide if there is a need for a new concept in the user representation, we follow an approach similar to the one presented in [7], i.e., using entropy as uncertainty measure for the classifier’s predictions. Given the class probability distribution \([P(C_1|x)\ldots P(C_k|x)]\) of existing classes k, for a new instance x, for a given machine learning approach, we decide that we need to generate a new class if the class probabilities entropy is larger than 1.0:
Whenever the entropy is high, we inform the user that there might be the need to introduce a new concept. Using the hierarchical classification models, we suggest potential new candidate concepts retrieved from the target ontology.
Merging Concepts. The action of merging concepts is trivial: if two user-defined concepts are aligned to the same target ontology concept, then the user concepts are merged.
Reassigning Instances. As the user conceptualization grows by adding new instances and new concepts, the user’s view is also evolving, so reorganization of the instances might be needed. To assist the user in this step, we train the hierarchical classification model on all the instances in the user’s conceptualization data, and then we use the model to classify all the instances. Analyzing the class prediction distribution, we can identify two types of candidates for reassigning: (i) Misclassifying an instance indicates that the instance might be an outlier in the currently assigned concept, implying that the instance is assigned in the current concept because of a user error; (ii) High entropy (see Eq. 1) indicates that the instance might fit better in a different concept than the current one. The system presents the suggestions to the user to decide if the instances need to be reassigned. When instances are reassigned, the model is retrained on the updated conceptualization. The stopping criterion for reassignment is that there are no more updates in the concepts.
4 Experiments
The goal of the experiments is to (i) test the performance of the novel alignment strategy (Sect. 4.1), and (ii) test the effectiveness of the ontology maintenance steps: adding new instances to existing user concepts (Sect. 4.2), detecting when a new concept should be added to the user model (Sect. 4.2), and suggesting when a concept should be split (Sect. 4.2).
All the experiments were carried out in the medical domain, specifically tackling the problem of Adverse Drug Reactions, for which we worked with a medical doctor to create a manually annotated gold standard dataset. Starting from user blogs extracted from http://www.askapatient.com (a forum where patients report their experience with medication drugs), we extracted all instances referring to adverse drug events, grouped the instances referring to the same adverse event into concepts, and aligned them to the MedDRA ontology [2]. MedDRA is a rich and standardized medical terminology organized in 5 levels, arranged from very general to very specific concepts: the fifth level contains 95, 061 leaf instances. The user data contains 203 concepts (adverse drug reactions), each of them containing several different phrases to refer to each concept, for a total of 3, 262 instances. The 203 concepts have been manually aligned to MedDRA, using a total of 169 concepts at the lowest level (some of the user concepts are aligned to the same MedDRA concept). The details for the user’s dataset and the MedDRA dataset per level are shown in Table 1.
4.1 Aligning User’s Conceptualization with a Target Ontology
Given each user concept, the task is to identify, if it exists, a concept in the target ontology that identifies it. We assess the performance of our proposed methods against the gold standard dataset, and compare them against different baseline methods. To evaluate the approaches, we use the metric HITS@K, which measures if the correct alignment is in the top-K ranked results of the approach.
We implemented three baselines for comparison:
- String-based average-link matching. :
-
Given a user concept \(c_u\), we calculate the similarity to each concept \(c_t\) at a given level of the hierarchy (Eq. 2), using a LuceneFootnote 6 token-based similarity score with edit distance of 2 and tf-idf weighting. We then rank the results and select the top-N classes.
$$\begin{aligned} sim(c_u,c_t)=\frac{1}{|c_u||c_t)|}\sum _{i=1}^{|c_u|}\sum _{j=1}^{|c_t|}sim(x_{ci},x_{cj}) \end{aligned}$$(2) - Word embeddings. :
-
To build the word embedding we first collected a domain-specific text, i.e., patient reports about adverse drug reaction for more than 2, 000 drugs, retrieved from www.askapatient.com, the ADE corpus [13], and the EMEA datasetFootnote 7 (European Medicines Agency documents). We use the corpora of sentences to build both CBOW and Skip-Gram models with the default parameters proposed in [20].Footnote 8 Given a user-defined group \(c_u\), we calculate the similarity to each concept \(c_t\) on a given level of the hierarchy, using Eq. 2, where the similarity between two instances is calculated as a cosine similarity between the averaged vectors of all the words in the instances.
- LDA. :
-
We use the Stanford Labeled LDA tool [26] to build a supervised topic model, using the nodes in each level of the hierarchy as labels. To select the top-N classes for each user-defined group, we perform majority vote using the topic probabilities as weights.
Our three methods (Sect. 3.1) are not bound to the choice of the specific classifier. We use the instances of the target ontology to train each classifier, which is then used to classify the user’s concepts. We report the performances for the following classifiers: Support Vector Machines (SVM) with RBF kernel, Random Forests (RF), Logistic Regression (LR), and Convolutional Neural Network (CNN). All classifiers use the domain-specific word2vec word embedding as features, the same as the baseline method. The architecture of the CNN model is inspired by Collobert et al. [6] and Kim [16], which has shown high performances in many NLP tasks.Footnote 9
Figure 3 shows the results for HITS@1 to HITS@10 for each level of the hierarchy. For each of the three hierarchical approaches we report the best classifier.Footnote 10 The LDA approach performs rather poorly, therefore we exclude it from the plots. The HITS@1 results for all approaches are shown in Table 2. As the curves show (Fig. 3), while HITS@1 results (fully automated approach) are very encouraging, including human-in-the-loop (proposing the 10 most likely options) increases the performance up to 99.5% accuracy (on level 1 of the hierarchy) which is desirable in this domain.
We can observe that all three approaches outperform the baseline methods, with a larger margin as we move down in the hierarchy. The word embeddings approach outperforms the string-based baseline approach on all levels. The CNN classifier outperforms the standard classifiers, although Logistic Regression achieves comparable performance. It is noteworthy that the flat hierarchical classification approach performs rather well on the first 3 levels, however the performance drops at level 4, where the number of classes is significantly higher. Furthermore, we were not able to build flat SVM, RF and LR models for the lowest level of the hierarchy, as the number of class labels is rather high, the models ran out of memory. The top-down per parent node approach shows comparable results for HITS@1, however the propagation of errors from the previous levels leads to poor performances for HITS@10, i.e., if an instance is incorrectly classified in level \(l-1\), in level l the HITS will not increase when k increases because the model cannot find the correct concept in the ontology. The combined hierarchical classification approach outperforms all the others on all levels of the hierarchy.
4.2 Ontology Maintenance
Adding New Instances. In this experiment the goal was to add new instances to the user’s knowledge base. To do so, we first built a CNN model for each level of the already aligned user hierarchy. Then we retrieved additional 298 instances of Adverse Drug Events from www.askapatient.com, which were not included in the initial data, and used the previously built model to assign each of them in the user’s knowledge base.
The results for HITS@10 at each level of the hierarchy are shown in Fig. 4. The results show that we were able to classify the instances in the correct user concepts with HITS@10 = 99.1% on the most general level, and HITS@10 = 91.27% on the lowest level of the hierarchy.
Adding New Concepts. In this experiment we evaluated the model’s ability to notify the user that a new concept should be introduced, i.e., a new instance doesn’t fit in any of the defined concepts, therefore a new concept should be added. To do so, we selected 500 instances from the MedDRA ontology that don’t belong to any of the user’s concepts, i.e., positive instances for which the model is expected to create a new concept, and 500 instances that belonged to some of the user’s concepts, i.e., negative instances for which the model shouldn’t create a new concept.Footnote 11 Then we used the previously built CNN model for the last level of the hierarchy to classify the set of instances. We used the approach for adding new concepts (shown in Sect. 3.2) to decide if for each instance we need to add new concept. We expect the approach to notify the user that a new concept should be added for the first 500 instances.
For this task we measure precision (P), recall (R) and F-score(F). The approach achieved \(P=73.8\%\), \(R=84.6\%\) and \(F=78.83\%\).
Reassigning Instances. In this experiment we evaluated the model’s ability to reassign instances to other concepts. We try to identify (i) mistakes made by the user in the conceptualization or (ii) alternative and potentially better concepts for a given instance (if any is found). Also, the user’s view is evolving over time, so reorganization of the instances might be needed.
The model was able to identify 82 instances that needed to be reassigned. The instances were reviewed by a medical doctor, who accepted 67 instances to be reassigned, yielding precision \(P=81.7\%\). For those instances, we used the model to assign new concepts, achieving \(HITS@1=76.11\%\) and \(HITS@5=91.05\%\). Using our approach we were able to easily identify misclassifications caused by user error. For example, “stomach aches” was initially assigned in the “Emotional disorder” concept, which was identified by our model as a mistake and was reassigned to “Abdominal distension”. Beside the trivial cases, the model proposes to the user to review instances that might fall in different concepts. For example, “sensitivity to light” was initially assigned in the “Visually impairment” concept, but after the growth of the concepts, the model suggested to move the instance to “Photophobia”, which was accepted by the user.
5 Conclusions and Future Work
In this paper we introduce a methodology to perform user-centric ontology population that efficiently includes human-in-the-loop at each step: the user is assisted in building, connecting and maintaining their conceptualization of the domain, while taking advantage of already available ontologies. We design a novel hierarchical classification method for ontology population, which builds hierarchical classification models that are dynamically refined based on user interaction. Our main objective is not to fully automate the process but rather to assist the user in achieving their goals more efficiently and effectively. The experiments confirm that the approach supports the user to achieve nearly perfect performance. The user has full control on their level of involvement in the process, depending on the requirements for quality and precision of the data, and her time/cost limit.
As future work, we are performing experiments on a broader task in the medical domain and using UMLSFootnote 12 for the alignment. Furthermore, we will analyze to which extent our approach can be applied to different languages, and perform cross-lingual alignment.
Notes
- 1.
It is expected the user to be able to perform simple browsing and navigation through data, but no knowledge of Semantic Web Technologies is needed, e.g., RDF, SPARQL etc.
- 2.
- 3.
- 4.
- 5.
- 6.
- 7.
- 8.
Additionally we fix window size = 5; dimensions = 200; number of iterations = 15; negative sampling for optimization; negative samples = 25; average input vector for CBOW.
- 9.
We selected the following parameters for the CNN model: an input embedding layer, 4 convolutional layers followed by max-pooling layers, a fully connected softmax layer, rectified linear units, filter windows of 2, 3, 4, 5 with 100 feature maps each, dropout rate of 0.2 and mini-batch size of 50. For the embedding layer we use the word2vec embeddings used in the baseline approach to initialize the weighing matrix. We train 100 epochs with early stopping.
- 10.
Note that for the first level we report only the results for the flat hierarchical classification approach, because the results are the same for all three approaches.
- 11.
Note that these examples were not used in the training phase.
- 12.
References
Aroyo, L., Welty, C.: Crowd Truth: harnessing disagreement in crowdsourcing a relation extraction gold standard. Web Science 2013, 25371, pp. 1–6 (2013)
Brown, E.G., Wood, L., Wood, S.: The medical dictionary for regulatory activities (MedDRA). Drug Saf. 20(2), 109–117 (1999)
Castano, S., Peraldi, I.S.E., Ferrara, A., Karkaletsis, V., Kaya, A., Möller, R., Montanelli, S., Petasis, G., Wessel, M.: Multimedia interpretation for dynamic ontology evolution. J. Log. Comput. 19(5), 859–897 (2008)
Cimiano, P., Völker, J.: Towards large-scale, open-domain and ontology-based named entity classification. In: RANLP (2005)
Coden, A., Gruhl, D., Lewis, N., Tanenblatt, M., Terdiman, J.: SPOT the drug! An unsupervised pattern matching method to extract drug names from very large clinical corpora. In: Proceedings of the 2012 IEEE 2nd Conference on Healthcare Informatics, Imaging and Systems Biology, HISB 2012, pp. 33–39 (2012)
Collobert, R., Weston, J., Bottou, L., Karlen, M., Kavukcuoglu, K., Kuksa, P.: Natural language processing (almost) from scratch. J. Mach. Learn. Res. 12(Aug), 2493–2537 (2011)
Dalvi, B., Mishra, A., Cohen, W.W.: Hierarchical semi-supervised classification with incomplete class hierarchies. In: Proceedings of the Ninth ACM International Conference on Web Search and Data Mining, pp. 193–202. ACM (2016)
Doddington, G.R., Mitchell, A., Przybocki, M.A., Ramshaw, L.A., Strassel, S., Weischedel, R.M.: The automatic content extraction (ACE) program-tasks, data, and evaluation. In: LREC (2004)
Dong, L., Wei, F., Sun, H., Zhou, M., Xu, K.: A hybrid neural model for type classification of entity mentions. In: IJCAI, pp. 1243–1249 (2015)
Gangemi, A., Alam, M., Asprino, L., Presutti, V., Recupero, D.R.: Framester: a wide coverage linguistic linked data hub. In: Blomqvist, E., Ciancarini, P., Poggi, F., Vitali, F. (eds.) EKAW 2016. LNCS (LNAI), vol. 10024, pp. 239–254. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-49004-5_16
Gangemi, A., Presutti, V., Reforgiato Recupero, D., Nuzzolese, A.G., Draicchio, F., Mongiovì, M.: Semantic web machine reading with FRED. Semantic Web (Preprint), pp. 1–21 (2016)
Giuliano, C., Gliozzo, A.: Instance-based ontology population exploiting named-entity substitution. In: ACL 2008, pp. 265–272. ACL (2008)
Gurulingappa, H., Rajput, A.M., Roberts, A., Fluck, J., Hofmann-Apitius, M., Toldo, L.: Development of a benchmark corpus to support the automatic extraction of drug-related adverse effects from medical case reports. J. Biomed. Inform. 45(5), 885–892 (2012)
Holzinger, A.: Interactive machine learning for health informatics: when do we need the human-in-the-loop? Brain Inform. 3(2), 119–131 (2016)
Holzinger, A., Jurisica, I.: Knowledge discovery and data mining in biomedical informatics: the future is in integrative, interactive machine learning solutions. In: Holzinger, A., Jurisica, I. (eds.) Interactive Knowledge Discovery and Data Mining in Biomedical Informatics. LNCS, vol. 8401, pp. 1–18. Springer, Heidelberg (2014). https://doi.org/10.1007/978-3-662-43968-5_1
Kim, Y.: Convolutional neural networks for sentence classification. arXiv preprint arXiv:1408.5882 (2014)
Ling, X., Weld, D.S.: Fine-grained entity recognition. In: AAAI 2012, pp. 94–100. AAAI Press (2012). http://dl.acm.org/citation.cfm?id=2900728.2900742
McDowell, L.K., Cafarella, M.: Ontology-driven, unsupervised instance population. Web Semant. Sci. Serv. Agents World Wide Web 6(3), 218–236 (2008)
Mendes, P.N., Jakob, M., García-Silva, A., Bizer, C.: DBpedia spotlight: shedding light on the web of documents. In: Proceedings of the 7th International Conference on Semantic Systems, pp. 1–8. ACM (2011)
Mikolov, T., Sutskever, I., Chen, K., Corrado, G.S., Dean, J.: Distributed representations of words and phrases and their compositionality. In: Advances in Neural Information Processing Systems, pp. 3111–3119 (2013)
Moro, A., Raganato, A., Navigli, R.: Entity linking meets word sense disambiguation: a unified approach. Trans. Assoc. Comput. Linguist. 2, 231–244 (2014)
Murty, S., Verga, P., Vilnis, L., McCallum, A.: Finer grained entity typing with typenet. arXiv preprint arXiv:1711.05795 (2017)
Nakashole, N., Tylenda, T., Weikum, G.: Fine-grained semantic typing of emerging entities. In: ACL, vol. 1, pp. 1488–1497 (2013)
Nuzzolese, A.G., Gentile, A.L., Presutti, V., Gangemi, A., Garigliotti, D., Navigli, R.: Open knowledge extraction challenge. In: Gandon, F., Cabrio, E., Stankovic, M., Zimmermann, A. (eds.) SemWebEval 2015. CCIS, vol. 548, pp. 3–15. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-25518-7_1
Petasis, G., Karkaletsis, V., Paliouras, G., Krithara, A., Zavitsanos, E.: Ontology population and enrichment: state of the art. In: Paliouras, G., Spyropoulos, C.D., Tsatsaronis, G. (eds.) Knowledge-Driven Multimedia Information Extraction and Ontology Evolution. LNCS (LNAI), vol. 6050, pp. 134–166. Springer, Heidelberg (2011). https://doi.org/10.1007/978-3-642-20795-2_6
Ramage, D., Hall, D., Nallapati, R., Manning, C.D.: Labeled LDA: a supervised topic model for credit attribution in multi-labeled corpora. In: Empirical Methods in Natural Language Processing (EMNLP), pp. 248–256. Association for Computational Linguistics, Singapore, August 2009
Ren, X., He, W., Qu, M., Huang, L., Ji, H., Han, J.: AFET: automatic fine-grained entity typing by hierarchical partial-label embedding. In: Proceedings of the Conference on Empirical Methods in Natural Language Processing (EMNLP) (2016)
Ristoski, P., Bizer, C., Paulheim, H.: Mining the web of linked data with rapidminer. Web Semant. Sci. Serv. Agents World Wide Web 35, 142–151 (2015)
Ristoski, P., Faralli, S., Paolo Ponzetto, S., Paulheim, H.: Large-scale taxonomy induction using entity and word embeddings. In: Proceedings of the International Conference on Web Intelligence (2017)
Ristoski, P., Paulheim, H.: Semantic web in data mining and knowledge discovery: a comprehensive survey. Web Semant. Sci. Serv. Agents World Wide Web 36, 1–22 (2016)
Segura-Bedmar, I., Martínez, P., Herrero Zazo, M.: Semeval-2013 task 9: extraction of drug-drug interactions from biomedical texts (DDIExtraction 2013). In: SemEval 2013, pp. 341–350. ACL, June 2013
Shimaoka, S., Stenetorp, P., Inui, K., Riedel, S.: An attentive neural architecture for fine-grained entity type classification. arXiv preprint arXiv:1604.05525 (2016)
Shimaoka, S., Stenetorp, P., Inui, K., Riedel, S.: Neural architectures for fine-grained entity type classification. arXiv preprint arXiv:1606.01341 (2016)
Silla Jr., C.N., Freitas, A.A.: A survey of hierarchical classification across different application domains. Data Min. Knowl. Discov. 22(1–2), 31–72 (2011)
Tanev, H., Magnini, B.: Weakly supervised approaches for ontology population. Citeseer (2008)
Tjong Kim Sang, E.F., De Meulder, F.: Introduction to the CoNLL-2003 shared task: language-independent named entity recognition. In: HLT-NAACL 2003, pp. 142–147. CONLL, Stroudsburg (2003)
Velardi, P., Faralli, S., Navigli, R.: Ontolearn reloaded: a graph-based algorithm for taxonomy induction. Comput. Linguist. 39(3), 665–707 (2013)
Velardi, P., Navigli, R., Cuchiarelli, A., Neri, R.: Evaluation of ontolearn, a methodology for automatic learning of domain ontologies. In: Ontology Learning from Text: Methods, Evaluation and Applications, vol. 123, p. 92 (2005)
Yaghoobzadeh, Y., Adel, H., Schütze, H.: Noise mitigation for neural entity typing and relation extraction. arXiv preprint arXiv:1612.07495 (2016)
Yaghoobzadeh, Y., Schütze, H.: Corpus-level fine-grained entity typing using contextual information. arXiv preprint arXiv:1606.07901 (2016)
Yogatama, D., Gillick, D., Lazic, N.: Embedding methods for fine grained entity type classification. In: ACL, vol. 2, pp. 291–296 (2015)
Yosef, A.M., Bauer, S., Hoffart, J., Spaniol, M., Weikum, G.: HYENA: hierarchical type classification for entity names. In: COLING 2012: Posters, pp. 1361–1370 (2012)
Zhai, H., Lingren, T., Deleger, L., Li, Q., Kaiser, M., Stoutenborough, L., Solti, I.: Web 2.0-based crowdsourcing for high-quality gold standard development in clinical natural language processing. J. Med. Int. Res. 15(4), 1–17 (2013)
Zhang, L., Rettinger, A.: X-LiSA: cross-lingual semantic annotation. VLDB 7(13), 1693–1696 (2014)
Acknowledgement
We would like to thank Dr. Joseph Terdiman MD, a general practitioner with over 50 years of clinical experience, for the manual annotation of the gold standard.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this paper
Cite this paper
Clarkson, K., Gentile, A.L., Gruhl, D., Ristoski, P., Terdiman, J., Welch, S. (2018). User-Centric Ontology Population. In: Gangemi, A., et al. The Semantic Web. ESWC 2018. Lecture Notes in Computer Science(), vol 10843. Springer, Cham. https://doi.org/10.1007/978-3-319-93417-4_8
Download citation
DOI: https://doi.org/10.1007/978-3-319-93417-4_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-93416-7
Online ISBN: 978-3-319-93417-4
eBook Packages: Computer ScienceComputer Science (R0)