Abstract
Self-assembled organic compounds which form ribbon-like micellar clusters may attach themselves to proteins, penetrating in areas of low stability. Such complexation involves regions other than the protein’s natural binding site. The supramolecular ligand adheres to beta folds or random coils which become susceptible to complexation as a result of function-related structural changes – e.g. antibodies engaged in immune complexes or acute phase proteins. However, even seemingly unsusceptible helical proteins may bind Congo red if they include chameleon sequences (short peptide fragments capable of adopting different secondary conformations depending on environmental conditions). Examples of such proteins include hemoglobin and albumin. Complexation of supramolecular Congo red is often associated with increased fluorescence, indicating breakdown of ligand micelles in the complex. This phenomenon may be used in diagnostic tests.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
- Congo red binding to haemoglobin
- Acute phase
- Congo red binding to albumin
- Fluorescence of Congo red
- Cell-cell interaction and Congo red binding
- Proteins susceptibility for protein complexation
- Chameleon sequences and Congo red binding
3.1 Natural Susceptibility of Proteins to Bind Congo Red
As remarked in Chaps. 1 and 2, supramolecular micellar systems may form complexes with proteins by penetrating in areas other than the active site . This process is conditioned by local instabilities in the protein structure [1, 2].
As a rule not all parts of a protein molecule are equally stable. Local instabilities are usually not significant enough to enable direct penetration of a large ligand consisting of many associated molecules. Such instabilities can, however, be artificially exacerbated, e.g. through heating [3]. Under natural conditions Congo red (CR ) is spontaneously bound by partly unfolded proteins capable of forming aggregations – such as amyloids and some abnormal [4,5,6,7,8,9]. While aberrant (unstable) proteins are usually eliminated before they can leave their parent cell [10,11,12,13], under certain circumstances – such as mass synthesis of light chains associated with multiple myeloma – they can be detected in circulation (Fig. 3.1) [14,15,16,17,18,19].
On the other hand, proteins destabilized partly through complexation of their “intended” natural ligands are commonly found in bodily fluids, especially in blood serum. Such proteins usually acquire the ability to bind CR . Examples include acute-phase proteins, conditioned to capture and eliminate other proteins whose presence in the bloodstream is harmful – e.g. proteolytic enzymes (which – due to their specific mechanism of action – may penetrate from the digestive tract or be released by necrotic cells ), as well as hemoglobin , which has undesirable catalytic properties and should not be present in the bloodstream in its unbound form. These acute phase complexes are similar to antibody /ligand complexes – by binding its natural ligand the protein undergoes structural deformations which facilitate penetration of an additional supramolecular ligand [20]. Acute phase complexes are short-lived, since they are recognized and removed by liver enzymes and phagocytes. Nevertheless, as long as the underlying cause persists, the blood serum will always contain proteins capable of association with CR . The specific composition of acute phase proteins present depend on the ongoing pathological processes. Acute phase proteins include serpins , haptoglobin , ceruloplasmin , ferritin , complement factors C3 and C4, as well as albumin, prealbumin and transferrin [21] (although note that in the last three cases pathological processes do not increase the concentration of the corresponding protein, but decrease it instead).
The ability of CR to bind to serum proteins is readily evidenced by agarose gel electrophoresis where the supramolecular ligand is added to the column at a certain stage of the process, accelerating migration of the affected proteins. Figure 3.2 presents a typical scenario where serum proteins are subjected to two-stage electrophoresis on agarose gel, with the second stage carried out in a perpendicular direction to the first, in the presence of CR (red band in Fig. 3.2D) [3, 21].
Two-directional agarose electrophoresis with exposed CR binding fractions dislocated over the diagonal line which includes non-binding serum proteins (A, B, C - three independent examples). (D) Presents the basis for this method. The red band indicates CR , which is spread on the agarose plate before the second electrophoretic step
The dye quickly migrates towards the anode and bypasses most proteins, except those structurally conditioned to bind the dye. As a result, migration of the affected proteins is accelerated and those proteins are found above the diagonal line which collects non bonding proteins. The mobility imparted by CR upon the target protein depends on the number of associated dye molecules and the molecular weight of the protein itself – hence the variable results observed under electrophoresis . The presented technique could have important diagnostic applications; here, however, our goal is merely to present the interaction of CR with serum proteins in order to explain the mechanism of supramolecular complexation. Regarding acute phase proteins, CR complexation is particularly evident in the case of proteins which always exhibit some form of activity, but whose activity in pathological conditions is significantly increased and/or altered.
Some proteins can bind supramolecular ligands even in the absence of a complexation partner which would account for structural rearrangement and reduction in stability. This includes albumin. Owing to its function, albumin is capable of binding various anionic compounds, of which the dye is an example. CR is furthermore capable of associating with amyloidogenic apolipoproteins [22,23,24,25,26].
3.2 Interaction of Congo Red with Helical Structures of Polypeptides
With regard to CR complexation capabilities, haptoglobin represents another interesting research target. The protein binds free hemoglobin , dissociating it into its alpha/beta subunits, but persists as a bridge between both halves [27]. The resulting structure can form complexes with CR . The dye itself also causes dissociation of hemoglobin into identical alpha/beta subunits, but releases them without forming a bridge. Hemoglobin is a typical allosteric protein, with the associated instability most likely concentrated in its alpha/beta interface region. This instability promotes complexation of supramolecular ligands . It appears that CR induces local changes in the alpha-helix structure of the subunit alpha of hemoglobin, transforming it into a beta fold or a random coil , which aids complexation. The process is reversible – adsorption of CR (e.g. on the P15 gel, which strongly binds the dye) yields normal hemoglobin tetramers. Increasing concentrations of the dye produce stronger dissociation – see Fig. 3.3 for electrophoretic images of ½ Hb/CR complexation activity. The effect is independently confirmed by DLS measurements, revealing in the mixture the reaction products of lower molecular weight (Fig. 3.4).
Complexation of dog hemoglobin with CR . Bars representing decreasing amount of dog hemoglobin (slow moving electrophoretic fraction) caused by its transfer to the fast-migrating complex with CR (inset). The CR dye concentrations in Hb were: 1 1.0 × 10−6 M/ml; 2 2 × 10−6 M/ml; 3 4 × 10−6 M/ml; 4 5 × 10−6 M/ml; 5 7.5 × 10−6 M/ml
Dissociation of hemoglobin can also be observed when treating hemoglobin crystals (horse) with CR . The resulting dissolution proves that, as a result of binding the dye, the protein undergoes structural changes which prevent crystallization (Fig. 3.5) [28,29,30,31].
Probable complexation loci in hemoglobin subunits have been identified by locating structurally unstable alpha folds, which also exhibit measurable propensity towards adopting beta or random coil conformations. Such folds may favor penetration of the supramolecular ligand , altering their own conformation in the process and producing finally stable bond. Comparative analysis has been performed by querying a database of chameleon sequences (i.e. sequences which may adopt either alpha or beta conformations, depending on local conditions) [32]. For each tetrapeptide, the database lists the corresponding probabilities of encountering alpha, beta and random coil conformations in actual proteins – this enables identification of fragments of the hemoglobin chain which may potentially transit to beta folds or random coil.
Results are presented in Fig. 3.6 as a sequence of bar charts illustrating the structural propensities of each residue sequence.
Bars representing the tendency of successive polypeptide chain fragments (numbers listed on the horizontal axis) of the human hemoglobin alpha subunit to adopt alpha, beta and random coil conformations respectively. The presented values are derived from the chameleon sequence database [32] (A) The same concerning neighbor amino acids (B)
While chameleon sequences are found in both subunits of the hemoglobin chain, their placement in the alpha subunit appears to be more favorable for binding CR – they are located close to one another and provide a convenient pocket with beta or random folds found on either side of the ligand. Experimental evidence suggests that CR penetrates the protein as a supramolecular ligand , wedging itself between parallel folds as long as such folds are not tightly packed and may be induced to adopt beta or random coil conformations. In contrast, helical folds do not support complexation of CR due to steric clashes. Regarding the alpha subunit, chameleon fragments are found in areas referred to as the G, H and FG helices, all located in close proximity of the heme. Structural rearrangement caused by complexation of CR therefore causes dissociation of the protein.
Figure 3.7 provides a space-filling depiction of the abovementioned fragments, while Fig. 3.8 illustrates the transition of unstable helices into beta folds or random coils , using albumin as an example. While albumin is ordinarily a helical protein, complexation of CR produces a notable conformational shift in favor of beta folds and random coils. Interestingly, similar interaction with EB and TB does not have a similar effect on the secondary conformation of the albumin chain [26] (Fig. 3.8).
It should be noted that the CR micelle is much more cohesive than its EB counterpart and therefore exerts a greater structural influence upon its surroundings. As a result, this relatively stable supramolecular ligand may force the target protein to adapt to its conformational preferences. This contrasts with EB , which relies on natural complexation sites present in the protein and does not induce conformational changes in its alpha chains. As already remarked, such “forcing” action of CR affects chameleon sequences which nominally appear as helices, but may also adopt beta or random coil conformations. Stable alpha folds do not yield to the presence of CR – although the stability threshold beyond which conformational changes occur is not well defined. It appears that increasing the concentration of CR results in more cohesive micelles with greater protein penetration ability. This is suggested by electrophoretic migration rate which increases as the acidity of CR in solution grows, indicating changes in pK of sulfonic group (Fig. 3.9).
Fluorescent albumin spot seen on an electrophoretic plate, showing that positively charged rhodamine B molecules with no affinity to albumin may be introduced to it anyway via CR which binds rhodamine B by intercalation . Agarose electrophoresis . 1 albumin combined with CR and rhodamine B (complex); 2 albumin + rhodamine B ; 3 rhodamine B ; 4 albumin + CR ; 5 CR . (A) UV image. (B) Visible light image following reduction and protein staining
Having induced structural changes in the protein, CR forms a complex with the newly produced beta fold or random coil , stabilizing the change. The dye penetration limit is determined by mutual alignment between the protein’s secondary folds and the dye micelle. An important factor facilitating penetration is the cohesiveness of the ligand itself, which – unlike the polypeptide – is not a polymer but rather an associate, stabilized by noncovalent interactions [33,34,35].
Albumin is a typical helical protein with a particularly vital role in blood serum. The importance of albumin is underscored by its natural concentrations and deep involvement in energy management processes. Albumin represents a source of amino acids in cases of malnutrition, but it also serves as a carrier of fatty acids. It is further capable of binding and transporting a variety of dyes and drugs [27]. Its complexation affinity for supramolecular CR and EB [26] makes it a useful study subject. Clearly, albumin may play a role in immunotargeting , since it is capable of forming complexes with supramolecular ligands doped with therapeutic agents.
The binding of supramolecular CR and EB by albumin has been confirmed by synthesizing co-micellar structures, i.e. supramolecular structures consisting of either dye mixed with a foreign compound a positively charged dye not normally complexed by albumin, such as rhodamine B or Janus Green , which can only be bound to the protein as an intercalant. In the case of rhodamine B, this effect is revealed by UV imaging of electrophoretic plates. The characteristic fluorescence of rhodamine B coincides with the location of the albumin stain, proving that the dye enters the protein as a component of a co-micellar structure formed by CR or EB (Fig. 3.10). The supramolecular nature of the ligand bound to albumin is also confirmed by counting the number of dye molecules attached to each albumin molecule (16–20 unit molecules in the case of CR ) [26]. The link between self-association properties and protein complexation potential further suggests that the ligand functions as a supramolecular entity.
Additional insight into the specifics of CR and EB complexation with albumin is provided by spectro-polarimetric analysis. The complex with EB is strongly chiral, while the corresponding complex with CR does not exhibit chirality [26]. This suggests that EB favors cis binding , while CR binds in a trans (alternating) alignment. The effect can be explained by referring to the structure of both dyes. In EB , all sulfonic groups are adjacent and bound to aromatic rings at either end of the molecule and therefore aligned with its long axis. This results in mutual modification of the dissociation constant and focuses electrostatic interactions on the polar regions. Consequently, rotation of the molecule about its long axis does not yield any benefits for self-association and can be considered irrelevant. Cis binding is likely related to the properties of other binding-capable substituents: the −OH and −NH2 groups. In contrast, in CR the location of sulfonic groups clearly favors trans binding, i.e. alternating alignment of symmetrical halves (Fig. 3.10).
The structure of albumin suggests that the supramolecular ligand is attracted to the gap between the protein’s two lobes [26]. This is where the longest non-helical folds can be found and where oscillatory structural changes (RMS-F) are revealed by molecular dynamic studies (depending on temperature), suggesting limited stability of the native secondary conformation. All these factors enable the anchoring of a supramolecular ligand (Fig. 3.11).
The most unstable area of albumin (200–400 aa) estimated according to secondary structure predisposition. (A) Profile indicating the predisposition to adopt helical (H), beta (B) and random coil (RC) conformations of each chain fragment. Fragments particularly likely to adopt beta forms are marked on the horizontal axis (shaded areas). (B) 3D presentation of albumin, with highlighted fragments (red) exhibiting the greatest predisposition to adopt a beta form. (C) 200–400 aa fragment of Hb polypeptide chain. (D) Flexibility along the polypeptide chain in albumin as expressed by the RMS-F parameter
3.3 Fluorescence Property of Congo Red
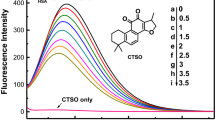
Interesting conclusions may be drawn from fluorescence analysis. Free CR is generally not fluorescent; however, fluorescence appears when the dye forms complexes with cellulose or with certain proteins. Theoretical analysis indicates that fluorescence should be inhibited by self-association which increases the mobility of Π-electrons and besides does not produce dipoles of equal length. In turn, fluorescence should be expected to emerge when the micelle dissociates into individual molecules or (possibly) oligomers. The exact conditions which favor fluorescence are not known with certainty. Supramolecular CR resembles a twisted tape. In the absence of a complementary surface (which mirrors its twists), the ribbon may only adhere to other molecules locally. This disrupts the micelle and favors retention of individual molecules or oligomers, explaining the fluorescence observed when CR interacts with cellulose. Another evidence of the proposed mechanism is induced fluorescence which occurs when CR is dissociated by a detergent, such as cholate , which can be intercalated into the micelle due to its planar structure, but which lacks aromatic rings (Fig. 3.12).
Calculations [36] based on the assumption that the bond between CR and beta polysaccharides such as cellulose is mediated by the polar fragments of the dye, appear to suggest that the optimal arrangement would involve clustering of individual dye molecules perpendicular to the polysaccharide chain. Although this argument does not acknowledge the self-association tendencies of CR , both theories do account for the retention of individual CR molecules, which would explain the observed fluorescence . Similar dissociation of the micelle may also be observed when CR forms complexes with certain proteins or protein aggregates – e.g. immunoglobulins or amyloids . It may also be caused by mechanical strain and dislocations in proteins which form complexes with the supramolecular dye. Such a situation may arise e.g. when cells react via their surface receptors – mostly cadherins and integrins , which have immunoglobulin -like conformations and become susceptible to CR penetration as a result of structural strain [37]. Subsequent motion of the cells exacerbates tension and promotes structural rearrangements, which may fragment the attached supramolecular ligand [38]. An example is provided by the reaction between monocytes and cancer cells . Monocytes recognize cancer cells as alien and attack them. Figure 3.13 presents visible-light and UV images of this process. Fluorescent patches correspond to the specific areas where cancer cells are attacked by monocytes. The images reveal free monocytes with no signs of fluorescence, as well as aggregations of monocytes clustered around cancer cells, with CR fluorescence clearly visible. The presented interpretation is also supported by induction of fluorescence through structural strain and rearrangements in antibodies forming immune complexes. In order to reveal this effect, the rosetting technique has been applied [39].
This process involves monocytes which bind antibodies via their Fc receptors. Sheep red blood cells have been added to a solution of anti-SRBC antibodies. Subsequent images show monocyte rosettes entirely covered by erythrocytes , suggesting even distribution of the immune complex. This indicates that conditions which favor CR complexation may emerge anywhere on the cell surface. When analyzing UV images, it becomes evident that fluorescence appears where the structural strain is greatest, fragmenting the ligand micelle into smaller units and/or individual molecules (Fig. 3.14).
The fluorescence of CR induced in a rosette system indicates that the supramolecular ligand is destabilized and partly dissociated by structural strain in its attached protein which does not provide a close match for the ligand’s own preferred conformation [40]. Similar results are obtained when heating complexes of CR with IgG light chains, which usually form two distinct fractions – the slow-moving and the fast-moving fraction. The former fraction is represented by ligands comprised of four dye molecules, whereas the latter fraction includes ligands with 5–8 dye molecules per micelle. There is only minimal smearing between the pure light chain stain and the slow-moving fraction stain, which indicates that slow-moving complexes are produced with ease. This fraction exhibits weak fluorescence. In contrast, the fast-moving fraction is formed slowly (preferentially at higher temperatures – 45–55 °C) and its electrophoretic image is smeared, indicating that the complex grows steadily from 5 to 8 dye molecules conquering some resistance – a process which also progressively increases its fluorescence (Fig. 3.15). The proposed mechanism is therefore validated. At higher dye concentrations, the distinction between both fractions is less dependent on temperature. This is likely due to simultaneous formation of both types of complexes in an environment characterized by abundance of CR .
As already discussed, supramolecular CR attaches itself to beta folds or random coils . This process is promoted by instabilities in the protein molecule, and, additionally, the complexation capabilities of the dye increase along with its concentration. By binding a supramolecular ligand , the protein adapts its tertiary conformation to the micelle; however most of the structural rearrangements which enable binding occur in the dye itself, and are facilitated by the noncovalent interactions between individual dye molecules.
References
Roterman I, No KT, Piekarska B, Kaszuba J, Pawlicki R, Rybarska J, Konieczny L (1993) Bis azo dyes –studies on the mechanism of complex formation with IgG modulated by heating or antigen binding. J Physiol Pharmacol 44(3):213–232
Rybarska J, Piekarska B, Stopa B, Zemanek G, Konieczny L, Nowak M, Król M, Roterman I, Szymczakiewicz-Multanowska A (2001) Evidence that supramolecular Congo red is the sole ligation form of this dye for L chain lambda derived amyloid proteins. Folia Histochem Cytobiol 39(4):307–314
Piekarska B, Konieczny L, Rybarska J, Stopa B, Zemanek G, Szneler E, Król M, Nowak M, Roterman I (2001) Heat-induced formation of a specific binding site for self-assembled Congo red in the V domain of immunoglobulin L chain lambda. Biopolymers 59(6):446–456
Lendel C, Bolognesi B, Wahlström A, Dobson CM, Gräslund A (2010) Detergent-like interaction of Congo red with the amyloid beta peptide. Biochemistry 49(7):1358–1360
Frid P, Anisimov SV, Popovic N (2007) Congo red and protein aggregation in neurodegenerative diseases. Brain Res Rev 53(1):135–160
Wang Y, Liu Y, Deng X, Cong Y, Jiang X (2016) Peptidic β-sheet binding with Congo Red allows both reduction of error variance and signal amplification for immunoassays. Biosens Bioelectron 86:211–218
Buell AK, Dobson CM, Knowles TP, Welland ME (2010) Interactions between amyloidophilic dyes and their relevance to studies of amyloid inhibitors. Biophys J 99(10):3492–3497
Howie AJ, Brewer DB (2009) Optical properties of amyloid stained by Congo red: history and mechanisms. Micron 40(3):285–301
Khurana R, Gillespie JR, Talapatra A, Minert LJ, Ionescu-Zanetti C, Millett I, Fink AL (2001) Partially folded intermediates as critical precursors of light chain amyloid fibrils and amorphous aggregates. Biochemistry 40(12):3525–3535
Ding Q, Cecarini V, Keller JN (2007) Interplay between protein synthesis and degradation in the CNS: physiological and pathological implications. Trends Neurosci 30(1):31–36
van Galen P, Kreso A, Mbong N, Kent DG, Fitzmaurice T, Chambers JE, Xie S, Laurenti E, Hermans K, Eppert K, Marciniak SJ, Goodall JC, Green AR, Wouters BG, Wienholds E, Dick JE (2014) The unfolded protein response governs integrity of the haematopoietic stem-cell pool during stress. Nature 510(7504):268–272
Lin JH, Li H, Yasumura D, Cohen HR, Zhang C, Panning B, Shokat KM, Lavail MM, Walter P (2007) IRE1 signaling affects cell fate during the unfolded protein response. Science 318(5852):944–949
Kawaguchi S, Ng DT (2011) Cell biology. Sensing ER stress. Science 333(6051):1830–1831
Edelman GM, Gally JA (1962) The nature of Bence-Jones proteins. Chemical similarities to polypetide chains of myeloma globulins and normal gamma-globulins. J Exp Med 116:207–227
Nakano T, Matsui M, Inoue I, Awata T, Katayama S, Murakoshi T (2011) Free immunoglobulin light chain: its biology and implications in diseases. Clin Chim Acta 412(11–12):843–849
Leitzgen K, Knittler MR, Haas IG (1997) Assembly of immunoglobulin light chains as a prerequisite for secretion. A model for oligomerization-dependent subunit folding. J Biol Chem 272(5):3117–3123
Kaplan B, Livneh A, Sela BA (2011) Immunoglobulin free light chain dimers in human diseases. Sci World J 11:726–735
Charafeddine KM, Jabbour MN, Kadi RH, Daher RT (2012) Extended use of serum free light chain as a biomarker in lymphoproliferative disorders: a comprehensive review. Am J Clin Pathol 137(6):890–897
Stevens FJ, Myatt EA, Chang CH, Westholm FA, Eulitz M, Weiss DT, Murphy C, Solomon A, Schiffer M (1995) A molecular model for self-assembly of amyloid fibrils: immunoglobulin light chains. Biochemistry 34(34):10697–10702
Raynes JG, Eagling S, McAdam KP (1991) Acute-phase protein synthesis in human hepatoma cells: differential regulation of serum amyloid A (SAA) and haptoglobin by inteleukin-1 and interlekin-6. Clin. Exp Immunol 83:488–491
Spólnik P, Piekarska B, Stopa B, Konieczny L, Zemanek G, Rybarska J, Król M, Nowak M, Roterman I (2003) The structural abnormality of myeloma immunoglobulins tested by Congo red binding. Med Sci Monit 9(4):BR145–BR153
Molloy TP, Wilson CW (1980) Protein-binding of ascorbic acid: 1. Binding of bovine serum albumin. Int J Vitam Nutr Res 50:380–386
Perrin JH, Nelson DA (1972) Induced optical activity following the binding of warfarin, indomethacin, 4-hydroxycoumarin and salicylic acid to human serum albumin. Life Sci I 11(6):277–283
Raghupathy E, Peterson NA, Estey SJ, Peters T Jr, Reed RG (1978) Serum albumin stimulation of synaptosomal proline uptake: partial identification on the active site. Biochem Biophys Res Commun 85(2):641–646
Ray A, Reynolds JA, Polet H, Steinhardt J (1966) Binding of large organic anions and neutral molecules by native bovine serum albumin. Biochemistry 5(8):2606–2616
Stopa B, Rybarska J, Drozd A, Konieczny L, Król M, Lisowski M, Piekarska B, Roterman I, Spólnik P, Zemanek G (2006) Albumin binds self-assembling dyes as specific polymolecular ligands. Int J Biol Macromol 40(1):1–8
Peters T Jr (1996) The albumin molecule: its structure and chemical properties. In: Peters T Jr (ed) All about albumin. Academic, San Diego/New York/Boston/London/Sydney/Tokyo/Toronto, pp 9–78
Dobryszycka W (1997) Biological functions of haptoglobin--new pieces to an old puzzle. Eur J Clin Chem Clin Biochem 35(9):647–654
Kurosky A, Barnett DR, Lee TH, Touchstone B, Hay RE, Arnott MS, Bowman BH, Fitch WM (1980) Covalent structure of human haptoglobin: a serine protease homolog. Proc Natl Acad Sci U S A 77(6):3388–3392
McCormick DJ, Atassi MZ (1990) Hemoglobin binding with haptoglobin: delineation of the haptoglobin binding site on the alpha-chain of human hemoglobin. J Protein Chem 9(6):735–742
Andersen CB, Torvund-Jensen M, Nielsen MJ, de Oliveira CL, Hersleth HP, Andersen NH, Pedersen JS, Andersen GR, Moestrup SK (2012) Structure of the haptoglobin-haemoglobin complex. Nature 489(7416):456–459
Ghozlane A, Joseph AP, Bornot A, de Brevern AG (2009) Analysis of protein chameleon sequence characteristics. Bioinformation 3(9):367–369
Spólnik P, Konieczny L, Piekarska B, Rybarska J, Stopa B, Zemanek G, Król M, Roterman I (2004) Instability of monoclonal myeloma protein may be identified as susceptibility to penetration and binding by newly synthesized Congo red derivatives. Biochimie 86(6):397–401
Konieczny L, Piekarska B, Rybarska J, Skowronek M, Stopa B, Tabor B, Dabroś W, Pawlicki R, Roterman I (1997) The use of Congo red as a lyotropic liquid crystal to carry stains in a model immunotargeting system--microscopic studies. Folia Histochem Cytobiol 35(4):203–210
Skowronek M, Stopa B, Konieczny L, Rybarska J, Piekarska B, Szneler E, Bakalarski G, Roterman I (1998) Self-assembly of Congo red – a theoretical approach to identify its supramolecular organization In water and salt solutions. Biopolymers 46:267–281
Woodcock S, Henrissat B, Sugiyama J (1995) Docking of Congo red to the surface of crystalline cellulose using molecular mechanics. Biopolymers 36(2):201–210
Burdick MM, McCarty OJ, Jadhav S, Konstantopoulos K (2001) Cell-cell interactions in inflammation and cancer metastasis. IEEE Eng Med Biol Mag 20(3):86–91
Zembala M, Siedlar M, Ruggiero I, Wieckiewicz J, Mytar B, Mattei M, Colizzi V (1994) The MHC class-II and CD44 molecules are involved in the induction of tumor necrosis factor (TNF) gene expression by human monocytes stimulated with tumour cells. Int J Cancer 56(2):269–274
Zupanska B (1990) Phenotypic markers for the feline monocyte: rosette which binds myeloid cells. Proc Soc Exp Biol Med 197:317–325
Roterman I, Rybarska J, Konieczny L, Skowronek M, Stopa B, Piekarska B, Bakalarski G (1998) Congo red bound to α-1-proteinase inhibitor as a model of supramolecular ligand and protein complex. Comput Chem 22:61–70
Acknowledgements
We acknowledge the financial support from the National Science Centre, Poland (grant no. 2016/21/D/NZ1/02763) and from the project Interdisciplinary PhD Studies “Molecular sciences for medicine” (co-financed by the European Social Fund within the Human Capital Operational Programme) and Ministry of Science and Higher Education (grant no. K/DSC/001370).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
This chapter is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
Copyright information
© 2018 The Author(s)
About this chapter
Cite this chapter
Zemanek, G., Jagusiak, A., Rybarska, J., Piwowar, P., Chłopaś, K., Roterman, I. (2018). Protein Conditioning for Binding Congo Red and Other Supramolecular Ligands. In: Roterman, I., Konieczny, L. (eds) Self-Assembled Molecules – New Kind of Protein Ligands. Springer, Cham. https://doi.org/10.1007/978-3-319-65639-7_3
Download citation
DOI: https://doi.org/10.1007/978-3-319-65639-7_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-65638-0
Online ISBN: 978-3-319-65639-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)