Abstract
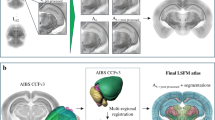
With the recent advance in 3D microscopy such as Single Plane Illumination Microscopy (SPIM) it is possible to obtain high resolution image volumes of the entire mouse brain. These data can be used to study the access of several peptides such as the glucagon-like peptide-1 (GLP-1) analogue Exendin-4, into the brain with the aim of developing medication for obesity. To investigate mode of action of the medication it is important to identify the specific anatomical brain nuclei that are targeted by the compound. Such segmentations can be obtained using an annotated digital brain atlas. We construct a SPIM brain atlas based on the Allen mouse brain 3D reference model and use it to analyze the access of peripherally injected Exendin-4 into the brain compared to a negative control group. The constructed atlas consists of an average SPIM volume obtained from eight C57BL mouse brains using group-wise registration. A cross-modality registration is performed between the constructed average volume and the Allen mouse brain reference model to allow propagation of annotations to the SPIM average brain. Finally, manual corrections of the annotations are performed and validated by visual inspection. The study shows that Exendin-4 have access to brain regions such as the arcuate hypothalamic nucleus and the nucleus of the solitary tract, which are areas involved in regulating food intake.
Chapter PDF
Similar content being viewed by others
Keywords
References
Allen institute for brain science. allen brain atlas [internet]. http://www.brain-map.org
Balci, S.K., Golland, P., Shenton, M., Wells, W.M.: Free-form b-spline deformation model for groupwise registration. In: MICCAI... International Conference on Medical Image Computing and Computer-Assisted Intervention, vol. 10, p. 23. NIH Public Access (2007)
Balland, E., Dam, J., Langlet, F., Caron, E., Steculorum, S., Messina, A., Rasika, S., Falluel-Morel, A., Anouar, Y., Dehouck, B., et al.: Hypothalamic tanycytes are an erk-gated conduit for leptin into the brain. Cell Metabolism 19(2), 293–301 (2014)
Castro-González, C., Ledesma-Carbayo, M.J., Peyriéras, N., Santos, A.: Assembling models of embryo development: Image analysis and the construction of digital atlases. Birth Defects Research Part C: Embryo Today 96(2), 109–120 (2012)
Dodt, H.U., Leischner, U., Schierloh, A., Jährling, N., Mauch, C.P., Deininger, K., Deussing, J.M., Eder, M., Zieglgänsberger, W., Becker, K.: Ultramicroscopy: three-dimensional visualization of neuronal networks in the whole mouse brain. Nature Methods 4(4), 331–336 (2007)
Dorr, A., Lerch, J.P., Spring, S., Kabani, N., Henkelman, R.M.: High resolution three-dimensional brain atlas using an average magnetic resonance image of 40 adult c57bl/6j mice. Neuroimage 42(1), 60–69 (2008)
Greger, K., Swoger, J., Stelzer, E.: Basic building units and properties of a fluorescence single plane illumination microscope. Review of Scientific Instruments 78(2), 023705 (2007)
Larsen, C.T., Iglesias, J.E., Van Leemput, K.: N3 bias field correction explained as a bayesian modeling method. In: Cardoso, M.J., Simpson, I., Arbel, T., Precup, D., Ribbens, A. (eds.) BAMBI 2014. LNCS, vol. 8677, pp. 1–12. Springer, Heidelberg (2014)
Lein, E.S., et al.: Genome-wide atlas of gene expression in the adult mouse brain. Nature 445(7124), 168–176 (2006)
Paxinos, G., Watson, C.: The rat brain in stereotaxic coordinates: hard cover edition. Academic press (2006)
Rueckert, D., Sonoda, L.I., Hayes, C., Hill, D.L.G., Leach, M.O., Hawkes, D.J.: Non-rigid registration using free-form deformations: Application to breast mr images. IEEE Transactions on Medical Imaging 18, 712–721 (1999)
Schnabel, J.A., et al.: A generic framework for non-rigid registration based on non-uniform multi-level free-form deformations. In: Niessen, W.J., Viergever, M.A. (eds.) MICCAI 2001. LNCS, vol. 2208, pp. 573–581. Springer, Heidelberg (2001)
Secher, A., et al.: The arcuate nucleus mediates glp-1 receptor agonist liraglutide-dependent weight loss. The Journal of Clinical Investigation 124(10), 4473 (2014)
Studholme, C., Hill, D.L., Hawkes, D.J.: An overlap invariant entropy measure of 3d medical image alignment. Pattern Recognition 32(1), 71–86 (1999)
Yushkevich, P.A., Avants, B.B., Ng, L., Hawrylycz, M., Burstein, P.D., Zhang, H., Gee, J.C.: 3D mouse brain reconstruction from histology using a coarse-to-fine approach. In: Pluim, J.P.W., Likar, B., Gerritsen, F.A. (eds.) WBIR 2006. LNCS, vol. 4057, pp. 230–237. Springer, Heidelberg (2006)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this paper
Cite this paper
Jensen, C.B., Secher, A., Hecksher-Sørensen, J., Conradsen, K., Larsen, R. (2015). Quantification of Brain Access of Exendin-4 in the C57BL Mouse Model by SPIM Fluorescence Imaging and the Allen Mouse Brain Reference Model. In: Paulsen, R., Pedersen, K. (eds) Image Analysis. SCIA 2015. Lecture Notes in Computer Science(), vol 9127. Springer, Cham. https://doi.org/10.1007/978-3-319-19665-7_38
Download citation
DOI: https://doi.org/10.1007/978-3-319-19665-7_38
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-19664-0
Online ISBN: 978-3-319-19665-7
eBook Packages: Computer ScienceComputer Science (R0)