Abstract
Fibroblast Growth Factors (FGFs), in a complex with their receptors (FGFRs) and heparan sulphate (HS), impact on a wide range of cellular functions, regulating processes from embryogenesis to metabolism. Upon ligand binding and receptor dimerisation, four key downstream pathways are initiated: MAPK, PI3K/AKT, STAT and PLCγ. Regulation of FGF signalling is critical to ensure a balanced response to receptor stimulation. This occurs through negative feedback mechanisms, including internalisation, cleavage and induction of negative regulators. FGF signalling has been studied in depth by developmental biologists, in a variety of model systems, and plays a critical role in developmental patterning and the establishment of paracrine signalling loops. Both germ line and somatic FGFR mutations are known to play a role in a range of diseases, most notably developmentally regulated diseases such as craniosynostosis dysplasias, dwarfism and hearing loss. Because of the ability of FGFR signalling to induce cell proliferation, migration and survival, FGFRs are readily co-opted by cancer cells. Mutations in, and amplifications of, these receptors are found in a range of cancers. Here, we outline the molecular mechanisms of FGFR signalling and discuss the role of this pathway in development and disease. We also address the rationale for therapeutic intervention and the need for FGFR-targeted therapy to selectively target cancer cells in view of the fundamental roles of FGF signalling in normal physiology.
Similar content being viewed by others
Keywords
6.1 Introduction
Fibroblast growth factors (FGFs) exert their cellular effects by interacting with FGF receptors (FGFRs) in a complex with heparan sulphate (HS) [1]. FGFRs, a class of receptor tyrosine kinase (RTK), dimerise and undergo transphosphorylation of the kinase domain upon ligand binding [2], leading to the recruitment of adapter proteins and initiating downstream signalling.
The extended FGF family is composed of 22 members, varying in size from 17 to 34 kDa. All members share a conserved 120 amino acid sequence and show 16–65 % sequence homology [3]. However, only eighteen FGFs signal via FGFR interactions (FGF1–10 and 16–23), while FGF11–14, which lack a signal peptide, act in an intracellular manner. Thus, many consider the FGF family to comprise only 18 members. Furthermore, although they are numbered from 1 to 23, FGF15 is the mouse ortholog of human FGF19. Each ligand binds to FGFRs with varying specificity; some are promiscuous, for example FGF1, and bind to multiple receptors, while others, like FGF7, bind only to one receptor isoform [4] (Fig. 6.1).
There are seven signalling receptors, encoded by four FGFR genes, FGFR1–4 [6]. FGFRs 1–3 have highly conserved intron/exon boundaries [4] (Fig. 6.2).
Schematic representation of FGFR1–4. FGFR1–4 contain a variety of defined structural domains, some of which are highly conserved across the receptors and their individual isoforms. The receptors are encoded by genes found on chromosomes 8, 10, 4 and 5, respectively. Three isoforms, termed a, b and c, exist for FGFR1 and 2; only isoforms b and c function in a signalling capacity. FGFR3 has two isoforms, b and c, while only one isoform of FGFR4 exists. These isoforms are generated through alternative splicing. Only one isoform of each receptor is shown. Amino acid residue numbers are indicated at the top of each panel, including a 21–22 amino acid signal sequence (UniProt accession: P11362, P21802, P22607, P22455, respectively)
Alternative splicing of exons 8 and 9, encoding IgIII of FGFR1–3, results in translation of two distinct isoforms capable of signal transduction. These isoforms are termed IIIb and IIIc, depending on which exons are spliced out (Fig. 6.3). This third Ig loop encodes the ligand binding domain; alternative splicing of this region is responsible for ligand binding specificity (Fig. 6.1). A third isoform exists for FGFR1 and 2, termed IIIa. This variant results in a truncated, secreted protein, which is unable to transduce a signal and may have an autoinhibitory role in FGF signalling, possibly by sequestering ligands [8]. FGFR4 is distinct in that it has only one isoform, homologous to the IIIc variant of FGFR1–3 [9].
FGFR structure, control of ligand specificity and receptor autoinhibition via alternative splicing. Each receptor monomer is comprised of an extracellular domain including three Ig loops, IgI, IgII and IgIII (also referred to as D1, D2 and D3, respectively), an acid box in the IgI–IgII linker region (represented by a white box), a transmembrane domain and an intracellular split kinase domain. Disulphide bonds are present in each Ig loop. IgI and the acid box are involved in autoinhibition of the receptor, while IgII and IgIII are involved in ligand binding. The HS-binding site lies in IgII, indicated in green. Ligand binding specificity is generated by alternative splicing of the IgIII domain. The first half of IgIII is encoded by an invariant exon (IIIa), which is spliced to either exon IIIb or IIIc (represented in blue and red, respectively), both of which splice to the exon that encodes the transmembrane domain (TM) region. Epithelial tissues predominantly express the IIIb isoform and mesenchymal tissues express IIIc. FGFR4 is expressed as a single isoform that is paralogous to FGFR-IIIc. An additional alternative splicing event can occur leading to the deletion of exons coding for IgI and/or the acid box/linker region. This leads to loss of receptor autoinhibition [7]
Receptor expression is generally cell type specific, for example IIIb and IIIc isoforms of FGFR1 and 2 are expressed in epithelial and mesenchymal cells, respectively [10, 11]. However, as shall be discussed later, this cell type specificity can change when FGFRs are associated with diseases such as cancer.
6.2 FGF:FGFR:HS Complex
Heparin, used in vitro as the model heparan sulphate (HS), is a member of the HS family of proteoglycans (HSPGs) and has been used to establish the necessity of HS binding in FGF:FGFR:HS complex formation [12]. This acidic molecule resembles the highly sulphated saccharide chains of HS [13]. Upon binding to FGFs/FGFRs, HS saccharide chains induce a conformational change. The length of the saccharide chain is important in FGF–FGFR interactions. Ornitz and colleagues reported interaction of a dodecasaccharide with both high- and low-affinity heparin-binding sites of ligands and showed that octasaccharides, thought to be the smallest saccharides with biological activity in FGF–FGFR interactions, could only engage the low-affinity binding sites of the ligand [14]. However, others have postulated that smaller chains, including hexasaccharides and disaccharides, may have biological activity [13, 15]. The heparin-binding residues found in the IgII loop of FGFRs (Fig. 6.2) are highly conserved [16], while heparin-binding residues of FGFs are diverse. Because of this, different FGFs require various HS sulphation patterns and/or length of chains for their optimum activity. Variability of HS sulphation patterns and length across cell types has an effect on FGF–FGFR interactions and may be a mediator of the biological activity of FGFRs [13–15, 17].
Another highly sulphated glycosaminoglycan (GAG), chondroitin sulphate (CS), is also able to interact with FGFs and FGFRs to promote complex formation. Studies have shown that insufficient synthesis of GAGs, which are assembled in the Golgi, impairs FGF/FGFR signalling capabilities [18]. The sulphation pattern and chain length of GAGs is so variable that there may be tissue- and even cell-specific GAG chains with varying specificities for ligands and receptors [19]. The difference in these chains may be of particular importance in the regulation of FGF/FGFR signalling. Work by a number of groups has also shown that variations in GAG sequences capable of interacting with FGFs and FGFRs can both inhibit or facilitate FGF signalling [17, 20–23].
A widely accepted model of FGFR interactions [16] proposed a complex of FGF:FGFR:HS in a 2:2:2 ratio (Fig. 6.4a). Two independent FGF:FGFR:HS ternary complexes are formed in a 1:1:1 ratio via HS binding to both receptor and ligand. They bind via receptor interactions, as well as interactions between the ligand in one complex and the receptor in another, thus forming a stable, symmetrical dimer. Direct ligand–ligand interactions are not observed. This FGF-FGFR complex can only be formed in the presence of HS. In summary, stabilisation of the dimer is through the following interactions: receptor–ligand, receptor–HS, ligand–HS and receptor–receptor.
Alternative FGF:FGFR:HS models. The basic structure of the FGF:FGFR complex comprises two receptor molecules and two ligands. Two models are presented which differ in the number of heparan sulphate proteoglycans (HSPGs) they contain. Dimerisation occurs upon ligand binding, leading to autophosphorylation of the kinase domain and subsequent downstream signalling. (a) FGF:FGFR:HS 2:2:2 model. First presented by Schlessinger et al. [16], and taken to be the most biologically relevant, this model proposes a symmetrical dimer utilising two HSPGs which bind ligands, bringing them into close proximity with the receptor, facilitating dimerisation. The HS chains also bind to the heparan-binding site of the IgII loop of the receptor to form a complete, active molecule capable of autophosphorylation and subsequent phosphorylation of signalling molecules. (b) FGF:FGFR:HS 2:2:1 model. Proposed by Pellegrini et al. [24], this dimer is formed using only one HSPG which binds both ligands necessary for each receptor monomer. Dimerisation occurs with the HS chain binding to both ligands and receptors, leading to signalling cascade activation
A second model [24] proposed FGF:FGFR:HS complex formation in a 2:2:1 ratio (Fig. 6.4b). Crystal structure analysis of FGFR2–FGF1 interactions showed a central heparin molecule linking two ligands and two receptor molecules. In this model, each ligand binds to a receptor monomer with heparin interacting with both ligands but only one receptor molecule. Two 1:1 FGF:FGFR complexes are joined to form a dimer via interactions with one HS chain.
6.3 Signalling Pathways
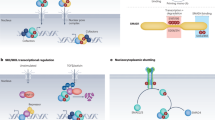
Upon dimerisation, reciprocal phosphorylation of the tyrosine kinase domains of the receptors occurs. These phosphorylated receptors are then able to act as docking sites for intracellular proteins, leading to activation of signalling cascades (Fig. 6.5) [25–27]. This autophosphorylation occurs in a specific order; ‘first-phase’ phosphorylation increases the catalytic activity of the kinase after ligand binding, while ‘second-phase’ phosphorylation creates phosphotyrosine-binding sites for docking molecules containing Src homology-2 (SH2) and phosphotyrosine-binding (PTB) domains [25, 28]. From this, four signalling pathways can be activated: MAP Kinase (MAPK), PI3K/AKT, PLCγ and STAT [25]. The key difference between FGFRs in signalling is the strength of their tyrosine kinase activity; their target proteins are the same [29].
FGF:FGFR-induced downstream signalling. Ligand-receptor binding induces four signalling cascades: MAPK, PI3K/AKT, PLCγ and STAT. These pathways comprise a series of phosphorylation events, culminating in the regulation of target genes, which dictate cellular processes, for example proliferation and migration
The lipid-anchored adapter protein FRS2 plays an integral role in the MAPK and PI3K/AKT pathways. FRS2α binds to the receptor via its PTB domain [30, 31] and undergoes phosphorylation. GRB2, another adapter molecule, is then recruited to FRS2α. From this point, two FGF-induced signalling pathways can be activated:
6.3.1 Phosphoinositide-3 Kinase
GRB2/FRS2α binds to and phosphorylates GAB1 via the SH3 domain of GRB2 [32]. This FRS2α/GRB2/GAB1 complex recruits PI3K via the SH2 domain of its p85 subunit. Activated PI3K produces phosphatidyl-inositol (3, 4, 5)-trisphosphate (PIP3), resulting in activation of the AKT pathway. Anti-apoptotic signalling, as well as cell growth and proliferation, is then initiated [32].
6.3.2 Mitogen-Activated Protein Kinase
Activation of the mitogen-activated protein kinase (MAPK) pathway results in mitogenic activity and cell survival [33]. The MAPK pathway is initiated by RAS binding to the FGFR/FRS2α/GRB2/SOS complex. RAS then recruits and phosphorylates RAF, leading to phosphorylation of MEK (MAPK/ERK kinase) and subsequent phosphorylation and activation of MAPK [33]. MAPK, also known as Extracellular Signal-Regulated Kinase (ERK), is then able to activate transcription factors in the nucleus, for example c-MYC, and influence the cell cycle.
The PLCγ and STAT pathways are mediated through other mechanisms.
6.3.3 Phospholipase C γ
Autophosphorylation of FGFR residue Tyr766 in FGFR1 creates a specific binding site for the SH2 domain of phospholipase Cγ (PLCγ), leading to tyrosine phosphorylation of PLCγ [34]. Recruitment of PLCγ is aided by PIP3, generated in response to PI3K stimulation [35]. Activation of PLCγ leads to cleavage of phosphotidyl-inositol-4, 5-bisphosphate (PIP2) into the second messengers inositol trisphosphate (IP3) and diacylglycerol (DAG) [36]. IP3 then releases calcium stores from the endoplasmic reticulum (ER) [37]. Calcium ions, along with DAG, then activate protein kinase C (PKC). PKC is then able to phosphorylate RAF and activate the MAPK pathway.
6.3.4 Signal Transducer and Activator of Transcription
The STAT family of cytoplasmic transcription factors can be activated by non-receptor tyrosine kinases, the Janus Kinases (JAK), leading to cell proliferation, differentiation or apoptosis [38]. Upon FGFR dimerisation and autophosphorylation, JAKs are phosphorylated by the receptor, forming a FGFR/JAK complex. This acts as a docking site for STATs, which are in turn tyrosine phosphorylated in their SH2 domain [39]. STAT dimers form and translocate to the nucleus, where they bind to gamma-activated site (GAS) enhancers to activate or repress gene transcription [39].
6.4 Regulation of FGF Signalling
Regulation of FGF signalling is critical to ensure a balanced response to receptor stimulation. This occurs largely through four mechanisms:
6.4.1 Receptor Internalisation
CBL, a multidomain protein that possesses an intrinsic ubiquitin ligase activity [40], binds to the FRS2α/GRB2 complex via the SH3 domain of GRB2 and the proline-rich region of CBL. Recruitment of CBL to FRS2α leads to ubiquitination of both FGFR and FRS2α and therefore attenuation of FGFR-mediated signalling.
6.4.2 Receptor Cleavage
Numerous growth factor receptors undergo ectodomain shedding, a process known to downregulate signalling. Ectodomain shedding, or S1 cleavage, is a process of proteolytic cleavage either within or near the membrane by members of the metalloprotease (MMP) and A Disintegrin And Metalloproteinase (ADAM) family [41]. Induction of this cleavage occurs in response to receptor activation [42]. Cleavage within the transmembrane domain by γ-secretase, known as S2 cleavage, often follows. Together, these cleavage events are known as Regulated Intramembrane Proteolysis [43].
FGFR1 is cleaved by MMP2 [44], and FGFR2 can be targeted by ADAM9 or 15 [45, 46]. Cleavage of both receptors leads to attenuation of signalling via two main mechanisms: downregulation of the number of active receptors at the cell surface and generation of a soluble extracellular domain able to compete with membrane-bound receptors for ligand binding [47]. Interestingly, FGFR1 also can be cleaved intracellularly by the serine protease Granzyme B. Although this was reported initially as a means of cytotoxic T lymphocytes inducing target cell apoptosis [48], the process is hijacked in cancer cells to allow nuclear trafficking of the C-terminus of the receptor, which acts to regulate transcription of a pro-migratory gene signature [49].
FGFR3 is unique in that the S1 cleavage occurs in an endosomal compartment, where it is cleaved by an as yet unknown protease, rather than involving a member of the ADAM family [41]. S2 cleavage via γ-secretase then occurs, generating a soluble intracellular domain capable of trafficking to the nucleus. Here, the nuclear FGFR3 fragment may be responsible for novel interactions in addition to the well-established downstream signalling pathways of receptor activation.
6.4.3 Induction of Negative Regulators
The first identified negative regulator of FGFRs was sprouty (SPRY) [50], one of a family of four proteins. SPRYs are thought to act through one of two mechanisms. Firstly, they may interact with GRB2, interrupting the FRS2α/GRB2 complex and therefore decreasing signal transduction [51]. Alternatively, SPRY–RAF interactions may occur, preventing RAF phosphorylation and therefore inhibiting MAPK signalling [52].
MAPK signalling can also be inhibited by Sprouty-related Enabled/vasodilator-stimulated phosphoprotein Homology 1 Domain-containing proteins (SPRED1 and 2) [53]. SPRED proteins prevent RAF activation of MEK by forming a complex between RAS and RAF. Co-localisation of SPRED2 with the protein Neighbor of BRCA1 (NBR1) results in sequestration of FGFR and lysosomal degradation [54].
Similar Expression to FGF (SEF) proteins also negatively regulate FGF signalling via a number of mechanisms: targeted inhibition at or downstream of MEK [55]; inhibition of RAS activation, which also inhibits the PI3K pathway [56]; direct interaction with FGFR and subsequent inhibition of FGFR and FRS2α phosphorylation [56–58]; and blockage of ERK/MEK dissociation, where SEF acts as a spatial regulator of phospho-ERK migration to the nucleus [59].
Another mechanism of negative regulation is via direct phosphorylation of MAPK pathway proteins. For example, SOS and RAF are substrates of MAPK. Phosphorylation of SOS by MAPK disrupts interactions between SOS and GRB2. This decreases recruitment of SOS to the membrane and results in diminished RAS activation [60]. MAPK also phosphorylates RAF, reducing RAF kinase activity and therefore decreasing MEK and MAPK phosphorylation [61]. Induction of the MAPK pathway can also lead to attenuation of the PI3K/AKT pathway. Activation of MAPK leads to GAB1 phosphorylation. This decreases PI3K recruitment to GAB1, in turn reducing AKT pathway activation [62].
Alternative internal control mechanisms of FGF signalling exist, including autoinhibition of the receptor [16, 63, 64]. The FGFRs exist in ‘closed’ and ‘open’ conformation equilibrium [7]. The first Ig loop (IgI) and the IgI/IgII linker region containing the acid box, a glutamate, aspartate and serine-rich sequence [6], are responsible for formation of the ‘closed’, autoinhibited state. Spectroscopic investigations have shown the acid box engages in electrostatic interactions with the HS-binding site of the IgII loop, inhibiting receptor–HS interactions and, therefore, receptor activation. This then encourages intramolecular interactions between IgI and the ligand-binding sites of the IgII and IgIII loops, further aiding the acquisition of a closed conformation [65]. Alternative splicing of exons encoding the IgI and/or acid box region leads to enhanced affinity of the receptor for its ligand and HS, increasing downstream signalling [65]. Loss of this region has been implicated in cancer [66, 67]. This mechanism of autoinhibition supports FGF binding specificity of receptors as only specific ligands with high affinity for the receptors will overcome the inhibition and bind to the receptor.
6.4.4 Klotho Interactions
FGFRs can also interact with klotho family proteins. These senescence-related, single-pass transmembrane proteins function as FGF19 subfamily signalling cofactors. FGFs are split into seven subfamilies, with the FGF19 subfamily comprising FGF19, 21 and 23. These endocrine factors regulate metabolic processes [68]. The HS-binding sites of this subfamily differ greatly from other FGFs, reducing their affinity for HS [69, 70]. Because of this, they require Klotho as a cofactor, to signal through FGFRs. Klotho expression is confined to a limited number of tissues [71]. It is able to bind FGFR1c independent of HS binding and convert it into a FGF23 receptor in the kidney [72]. Mutations in klotho proteins or the FGF19 subfamily are associated with diseases including autosomal dominant hypophosphatemic rickets, premature ageing disorders and diabetes [73]. Klotho is able to actively compete with FGF2 for FGFR1c binding, therefore attenuating FGF2 signalling [72].
6.5 FGFRs in Development
The critical role played by FGFR signalling during embryogenesis is highlighted by its conservation throughout evolution, from invertebrates through to higher mammals. There are a number of reviews that provide exquisite detail on FGFR signalling in a wide range of model organisms, including Caenorhabditis elegans [74], Drosophila melanogaster [75] and vertebrates [76–78]. However, we highlight below some key findings in the major model organisms.
6.5.1 Caenorhabditis elegans
C. elegans has just one FGFR, EGL-15, which was identified in mutant screens as a result of its importance in the migration of hermaphrodite sex myoblasts [79]. The EGL-15 receptor is essential for sensing the chemoattractant FGF ligand EGL-17, expressed in the target gonad and vulva [80]. Another FGF ortholog, LET-756, which shows structural homology to the FGF-9 subfamily [81], is essential for larval viability [82]. Further elements of the downstream signalling pathway were elucidated with the identification of a receptor tyrosine phosphatase, CLR-1 [83], and components of the MAPK cascade [84], which are key to FGF signalling in the worm, as they are in other model organisms. Interestingly, the FGF co-receptor Klotho has two functional orthologs in the worm, and these are essential in mediating the longevity and stress resistance effects of EGL-15/EGL-17 signalling [85]. Beyond the scope of this chapter, there are a number of non-canonical FGFR interactions in the worm that are the subject of an elegant review elsewhere [74].
6.5.2 Drosophila melanogaster
The tracheal system in Drosophila has been a key system for the identification of aspects of the FGF signalling pathway over the past 30 years. Breathless—one of two FGFRs in the fly—regulates tracheal branching [86], acting in concert with its cognate FGF ligand, branchless [87], to activate downstream MAPK signalling [75]. Further genetic dissection of the branching process identified Sprouty as a negative regulator of FGFR signalling [50] and described how the Notch pathway interacts with FGFR signalling in controlling cell fate [88], although orthologs of other negative regulators of the FGFR pathway, Sef and XFLRT3, are not present in invertebrates [89].
A further FGFR ortholog, Heartless, was identified by virtue of its pivotal role in mesoderm migration and subsequent specification [90, 91], although its ligands, the FGF-8 orthologs Pyramus and Thisbe, were not identified until much later [92, 93]. Downstream of FGF signalling, a novel adapter, Dof, is critical for activating intracellular signalling [94], in much the same fashion that FRS2 acts in the vertebrate pathway.
6.5.3 Zebrafish
FGF signalling is an important factor in patterning the zebrafish embryo, interacting with signalling by TGF-β superfamily members to regulate mesoderm induction [95]. FGF signalling acts as a posteriorising factor driving trunk and tail development during anterior–posterior patterning [96], regulating downstream T-box family transcription factors Notail and Spadetail [97], and also regulates dorso-ventral patterning [98].
One of the main regulators of FGF signalling, SEF, was found first in zebrafish [57, 99], and fundamental understanding of the roles for FGF-8 signalling in neural development has been identified through the study of mutant zebrafish strains [100–102].
6.5.4 Xenopus
The first studies of FGF signalling in early development focused on its role as a competence factor, using Xenopus as a model system and showing that cell fate in the developing embryo was regulated by FGFs [103, 104]. FGF signalling was shown to be essential for cells to respond to mesoderm inducing TGF-β superfamily members [105, 106], and components of the entire pathway, from FGFRs to HSPGs to signal transduction proteins, have all been studied in detail in the frog [76]. Defects caused when FGFR signalling is inhibited, by small molecule inhibition [107], morpholino knockdown [108, 109] or expression of dominant negative receptor [105, 110, 111], confirm its fundamental importance in mesoderm induction, morphogenetic movements, neural induction, neuronal determination and anterior–posterior patterning.
6.5.5 Chick
The ability to manipulate and culture chick embryos has helped reveal several key roles for FGFR signalling, including elegant grafting studies showing the importance of FGFR activity in specifying and driving limb development [112, 113] and dynamic studies of presomitic mesoderm determination and subsequent somitogenesis [114–116]. Furthermore, FGFR signalling has been shown to act as a competence factor for neural induction [117, 118].
6.5.6 Mouse
All of the FGFs and FGFRs have been targeted using genetically modified mouse models, with approaches including germline deletion, conditional knockout and constitutive/inducible expression of either dominant negative or activating mutation constructs. The phenotypes of FGF ligand knockout mice are summarised elsewhere [119]. Extensive studies have revealed key roles for FGFRs in development, homeostasis and disease, and these are detailed in Table ‘FGFR1–4 at a glance’.
6.6 FGFR1
FGFR1 is used as the model receptor in the majority of studies and many of the findings are relevant to all FGFRs.
Syndecan 4 (S4), a transmembrane proteoglycan with extracellular HS chains, can regulate FGFR1 signalling, as well as signal independently as a growth factor receptor, to initiate cell adhesion and migration [120]. Recent work has hypothesised S4 could also be involved in FGFR1 trafficking [121]. S4 has a PDZ-binding domain, which is capable of activating the small GTPase, RhoG. RhoG is kept in a complex with S4 in its inactive form. Upon FGF binding to FGFR1, aided by the HS chains of S4, a ligand–receptor–S4 complex is formed. Signalling pathways of the individual receptors are initiated, for example MAPK from FGFR1. Upon this complex formation, RhoG is released from S4 and is activated by guanine exchange factors (GEFs). This induces membrane ruffling, leading to macropinocytosis of the complete FGF–FGFR–S4 complex. Trafficking of the internalised complex is dependent on another small GTPase, Rab5. When Rab5 function is absent, the vesicles cannot mature and become functional signalling endosomes. In this scenario, the MAPK pathway is not activated. However, when Rab5 activity is restored and localises to the macropinosome containing the FGF-FGFR-S4 complex, maturation of the vesicle is facilitated and MAPK signalling is activated. When S4 is absent, RhoG activity is high, leading to increased macropinocytosis and therefore receptor internalisation. S4 controls the rate of FGF–FGFR–S4 complex macropinocytosis; overactive S4 and Rab5 can lead to inadequate attenuation of the MAPK signal leading to continuous downstream signalling effects, for example cell migration. Hence, a novel method of FGFR1 MAPK signalling regulation via S4-medated trafficking is proposed.
Nuclear localisation of both FGFs and FGFRs has been reported in a number of cell lines and tissues [122]. The mechanism of nuclear translocation of FGFR1 has recently been elucidated by Chioni & Grose.
Studies have also shown Importin β is involved in FGFR1 nuclear translocation [123]. It is proposed that this occurs via the Integrative Nuclear FGFR1 Signalling (INFS) pathway [124]. FGFR1 is released from the cytoplasmic membrane into the cytosol. As it does not contain a nuclear localisation signal (NLS), FGFR1 associates with Importin β, a carrier protein that does. FGFR1 can then be transported into the nucleus where it is able to influence expression of, for example, C-JUN.
6.7 FGFR2
Developmental disorders are commonly associated with FGFR mutations, including Kallmann and Lacrimo-Auriculo-Dento-Digital (LADD) syndromes [125]. In skeletal disorders, for example, Crouzon, Pfiffer and Jackson–Weiss syndromes, receptor mutations tend to cluster in the linker region, connecting IgII and IgIII, and in both IgIII and the IgIII-transmembrane domain linker, functioning by either promoting receptor dimerisation or altering ligand–receptor specificity. Mutations in two conserved cysteine residues in IgIII of FGFR2 are commonly found in these skeletal disorders [33]. These cysteine residues usually function by linking to another cysteine in IgIII of the receptor via intramolecular bonds. Substitution of this amino acid with another creates an unpaired cysteine residue able to form an intermolecular disulphide bridge, leading to receptor dimerisation and therefore activation.
The craniosynostosis syndrome, Apert syndrome, depends on FGFR2 mutations. Gain-of-function changes in the highly conserved residues S252 and P253 of the IgII and IgIII linker of FGFR2 result in a change in ligand binding specificity [126, 127]. These are the cause of the majority of Apert syndrome cases [128, 129]. This has been further shown in mouse models; S252W FGFR2c mutants showed activation of the c isoform of the receptor by mesenchymally expressed FGF7, while FGFR2b was activated by FGFs associated with epithelial expression [130]. It is also possible that S252W and P253R mutations lead to the modified receptor remaining on the cell membrane for an extended period of time, rather than undergoing rapid endocytosis into the lysosomes like its wild-type counterpart. Downstream signalling pathways are affected, leading to increased ERK phosphorylation and therefore increased cell proliferation and migration capabilities, as well as premature differentiation [131].
Mouse modelling of Apert syndrome has shown that a soluble, truncated FGFR2 isoform is upregulated and influences FGF1-FGFR2 binding. This glycosylated IIIa-TM isoform is generated by direct splicing of exon 7 (IIIa) to exon 10 (TM), generating a premature stop codon three amino acids into the TM exon [8]. This loss-of-function mutation can thus negatively regulate FGF signalling.
A number of cancers have been found to contain somatic mutations identical to germ line mutations in FGFRs associated with developmental disorders. For example, FGFR2 mutations commonly seen in Apert syndrome and Pfeiffer are frequently identified in endometrial cancer [132], for example S252W and N550K, both of which result in receptor activation. The S252W mutation resides in the linker region between IgII and IgIII, the area responsible for providing key contacts with the ligand. This increases the binding affinity of the receptor for a range of FGFs while also leading to violation of ligand specificity of the receptor isoforms [133].
Other FGFR2 mutations in endometrial cancer include S373C and Y376C, which result in gain of a cysteine residue, allowing formation of intermolecular disulphide bonds [134]. This leads to constitutive receptor dimerisation and therefore downstream signalling. Although these findings were established using FGFR2c functional studies, it is known FGFR3 contains paralogous mutations.
6.8 FGFR3
FGFR3 is mutated in a range of developmental and skeletal disorders and is the most frequently mutated FGFR in cancer, as noted by the extensive list in the Catalogue of Somatic Mutations in Cancer (COSMIC) database. Gain-of-function FGFR3 mutations are involved in the most severe form of dwarfism in humans, thanatrophoric dysplasia types I and II [135, 136] and achondroplasia [137, 138]. Similar somatic mutations have been found in bladder and cervical cancer, amongst others, where they are believed to have a positive effect on proliferation and inhibit apoptosis [139].
The FGFR3 germline mutation, A391E, is known to cause abnormal cranium growth and is responsible for Crouzon syndrome [140]. This mutation is also found in bladder cancer [141]. A391E leads to stabilisation of the transmembrane domain of the dimerised receptor independent of ligand binding [142] and is therefore responsible for ligand-independent receptor activation [143].
6.9 FGFR4
FGFR4 has a diverse range of roles, from involvement in the vascular system to regulation of hepatic bile acid and lipid metabolism [144, 145]. Recently, a single nucleotide polymorphism (SNP) in FGFR4 has been identified which is thought to have both positive and negative prognostic value in different diseases. This SNP (rs351855) results in a glycine-arginine change (G388R) in the transmembrane domain, leading to increased receptor stability and sustained receptor activation [146].
FGFR4 is expressed at high levels in coronary artery disease (CAD). Investigation of the SNP status of CAD patients in a Chinese population study showed this SNP is low in CAD patients [145]. It is therefore thought that having this SNP may be beneficial, acting as a protective factor against CAD development in Asian populations. This SNP is also associated with poor prognosis in prostate and breast cancer [146, 147].
6.10 FGFRL1: The Fifth FGFR
A fifth member of the FGFR family has been discovered, Fibroblast Growth Factor Receptor Like 1 (FGFRL1). This protein, which exists as a homodimer consisting of the three characteristic extracellular Ig-like domains, acid box between IgI and IgII and a transmembrane helix, differs from the classic receptors in that it has no intracellular tyrosine kinase domain [148–150]. Instead, the intracellular portion of FGFRL1 consists of only 100 residues including a histidine-rich sequence and a tandem tyrosine-based motif [148, 151, 152]. These two sequences function as signals for FGFRL1 trafficking from the plasma membrane to endosomes and lysosomes. Deletion of these sequences resulted in inefficient FGFRL1 internalisation and prolonged time at the plasma membrane [151].
Interactions with both FGFs and heparin have also been confirmed through dissociation studies [148]. FGFRL1 binds strongly to FGF3, 4, 8, 10 and 22 [153] and the affinity of FGF3 for FGFRL1 is at least one order of magnitude higher than the majority of FGFs for their receptors [151]. Affinity of this magnitude between FGFs and their receptors is only seen in mutant receptors in, for example, Pfeiffer, Apert and Muenke craniosynostosis syndromes [154]. The gain-of-function P253R mutation in Apert syndrome exactly matches an arginine residue at position 243 in FGFRL1; this residue could be responsible for the high affinity of FGF3 for FGFRL1 [155]. Its interaction with HS is also stronger than that of classic FGFRs and heparin [156, 157]. FGFs bind FGFRL1 between IgII and IgIII domains while heparin binds at the basic region at the beginning of the IgII loop [149, 158]. Autoinhibition of FGFRL1 via the IgI loop also occurs and the protein can be post-transcriptionally modified on one of its four glycosylation sites.
As FGFRL1 does not contain a tyrosine kinase domain it is not able to signal in the classical FGFR fashion. Its signalling function is yet to be fully determined, but a number of theories have been postulated. Firstly, the receptor could have an inhibitory effect on FGF signalling by sequestering ligands and therefore preventing them binding to FGFR1–4 [148, 149, 153]. Secondly, FGFRL1 could aid in internalisation and degradation of the classic receptors by binding to the same HS chain as the signalling receptor and effectively dragging it into endosomes/lysosomes. Thirdly, the tandem tyrosine-based motif and histidine-rich sequence could act as a docking site for tyrosine phosphatases, which could act on the signalling receptors and therefore attenuate signalling.
Although signalling mechanisms are yet to be elucidated, FGFRL1 can affect multiple cellular behaviours, inhibiting cell proliferation, increasing cell differentiation, regulating cell–cell contact and inducing cell–cell and cell–matrix adhesion. FGFRL1 is often found at the site of cell–cell contact and it is thought it may mediate cell adhesion by interacting with HS expressed on other cells [157]. FGFRL1 is also thought to play a role in craniosynostosis diseases and mutations in the protein have been found in ovarian cancer [151, 159].
Although not itself a receptor tyrosine kinase, it is clear that FGFRL1 plays an important role in FGF/FGFR signalling. Though full understanding of this role is yet to be determined, it is important to consider this fifth member of the FGFR family when understanding the complexity of FGF signalling.
6.11 Disease
As discussed for each receptor individually, both germ line and somatic FGFR mutations are known to play a role in a range of diseases, most notably craniosynostosis dysplasia and cancer (Table 6.1). Given the ability of the FGF signalling pathway to initiate cell survival and proliferation, amongst other cellular responses, it is not surprising this pathway is hijacked in cancer cells. Mutations in FGFRs in cancer are generally indicative of a more malignant phenotype. The majority of these mutations are activating, resulting in increased proliferation, migration and angiogenesis. However, recent data suggest that loss-of-function FGFR mutations may play a role in the development of some cancers [160, 161].
In cancer, chromosomal translocations lead to expression of constitutively active fusion proteins in which the FGFR tyrosine kinase domain is fused downstream of a constitutive dimerisation domain from a fusion protein. This has been seen in myeloproliferative syndromes (MPS), amongst other malignancies [162]. FGFR1 fusion proteins are known to cause 8p11 myeloproliferative syndrome (EMS), a form of MPS [163]. These fusion proteins are known to cause constitutive tyrosine kinase activation of FGFR1 while also leading to signalling independent of FRS2. Fusion proteins containing the FGFR3 kinase domain are also associated with multiple myeloma and peripheral T-cell lymphoma [33].
Other cancers associated with FGFR signalling deregulation include breast cancer, where FGFR1 and FGFR2 are amplified in approximately 10 and 2 % of breast cancers, respectively [164, 165]. Approximately 10 % of melanoma cases have FGFR2 mutations [160]. Interestingly, functional analysis has shown these mutations in melanomas result in loss of function of the receptor. The mutation spectrum, characteristic of those induced by UV radiation, includes 20 missense mutations occurring at conserved residues in FGFR2. Receptor loss of function due to this mutation is caused by loss of ligand binding affinity, impaired receptor dimerisation and decreased kinase activity.
FGFRs can also be involved in cellular transformation by interacting with other proteins. For example, in epithelial ovarian cancer (EOC), Neural Cell Adhesion Molecule (NCAM) is unregulated and promotes malignancy via interaction with FGFR [166]. However, as NCAM is known to inhibit FGF2–FGFR binding [167], NCAM/FGFR interactions increase malignancy via inhibition of normal FGF–FGFR interactions. FGF2/FGFR and NCAM/FGFR interactions therefore stimulate different receptor-mediated responses in EOC; NCAM/FGFR leads to increased cell migration, while FGF2/FGFR leads to increased proliferation. It is also possible the varying receptor interactions cause differential regulation of receptor trafficking to the endosome, resulting in different cellular responses and signalling kinetics [168]. Mouse models have shown that targeted abolition of NCAM/FGFR interaction with a monoclonal antibody results in elimination of metastatic dissemination of EOC. This has been shown via NCAM/FGFR1 interaction studies. However, as NCAM binds FGFR2 and FGFR4 [169], interaction of NCAM with multiple FGFRs may increase malignancy of EOC [166].
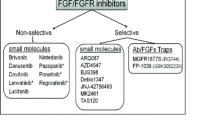
The high rate of FGFR mutation in a range of diseases makes this family of proteins a potential therapeutic target. Numerous studies have shown the benefits of FGFR knockdown and inhibition in cancer cell lines where the result is, for example, a decrease in cell proliferation [170]. However, translating this into a therapy for patients has proven difficult. Even specific FGFR inhibitors have off-target effects.
The most clinically advanced FGFR inhibitors to date are mixed kinase inhibitors, targeting the kinase domain of receptors to prevent downstream signalling. These include Dovitinib [171] and SU6668 [172]. However, their anti-FGFR activity is often weak, leading to investigation of more potent FGFR inhibitors. One such inhibitor currently in phase I clinical trials is AZD4547 [173]. This pyrazoloamide derivative targets FGFR1, 2 and 3 and resulted in cell growth inhibition versus cancer cell lines with known FGFR mutations and induces apoptosis. However, even this inhibitor has off-target effects, e.g. selectivity against VEGFR2 (also known as Kinase insert Domain Receptor, KDR), Insulin-like Growth Factor (IGF), PI3K and AKT, although this off-target inhibition is much lower than that of FGFRs. Such inhibitors still need more investigation, but the possibilities of potential FGFR inhibition are an exciting field of cancer therapeutics.
Abbreviations
- ADAM:
-
A Disintegrin And Metalloprotease
- CAD:
-
Coronary Artery Disease
- CBL:
-
Casitas B-lineage Lymphoma Protein
- CLR-1:
-
Cryptic Loci Regulator
- c-MYC:
-
Cellular-Myelocytomatosis Oncogene
- COSMIC:
-
Catalogue of Somatic Mutations in Cancer
- CS:
-
Chondroitin Sulphate
- DAG:
-
Diacylglycerol
- DOF:
-
Downstream of FGFR
- EGL:
-
Egg Laying Abnormal
- EOC:
-
Epithelial Ovarian Cancer
- ER:
-
Endoplasmic Reticulum
- ERK:
-
Extracellular Signal-Regulated Kinase
- FGF:
-
Fibroblast Growth Factor
- FGFR:
-
Fibroblast Growth Factor Receptor
- FGFRL1:
-
FGFR-Like 1
- FRS2:
-
Fibroblast Growth Factor Receptor Substrate 2
- GAB1:
-
GRB2-Associated Binding protein 1
- GAG:
-
Glycosaminoglycan
- GAS:
-
Gamma-Activated Site
- GEF:
-
Guanine Exchange Factor
- GRB2:
-
Growth Factor Receptor-Bound Protein 2
- HS:
-
Heparan Sulphate
- HSPG:
-
Heparan Sulphate Proteoglycan
- IGF:
-
Insulin-like Growth Factor
- INFS:
-
Integrative Nuclear FGFR1 Signalling
- IP3 :
-
Inositol trisphosphate
- JAK:
-
Janus Kinase
- KDR:
-
Kinase Insert Domain Receptor
- LADD:
-
Lacrimo-Auriculo-Dento-Digital
- LET-756:
-
Lethal Protein 756
- MAPK:
-
Mitogen-Activated Protein Kinase
- MEK:
-
ERK Kinase
- MMP:
-
Metalloprotease
- MPS:
-
Myeloproliferative Syndrome
- NBR1:
-
Neighbor of BRCA1
- NCAM:
-
Neural Cell Adhesion Molecule
- NLS:
-
Nuclear Localisation Signal
- PI3K:
-
Phosphoinositide-3 Kinase
- PIP2 :
-
Phosphatidyl-inositol-4, 5-bisphosphate
- PIP3 :
-
Phosphatidyl-inositol (3, 4, 5)-trisphosphate
- PKC:
-
Protein Kinase C
- PLCγ:
-
Phospholipase C γ
- PTB:
-
Phosphotyrosine Binding
- Rab5:
-
Ras-Related Proteins in Brain 5
- RAF:
-
Rapidly Accelerated Fibrosarcoma
- RAS:
-
Rat Sarcoma
- RhoG:
-
Ras Homology Growth-Related
- RTK:
-
Receptor Tyrosine Kinase
- S4:
-
Syndecan 4
- SEF:
-
Similar Expression to FGF
- SH2:
-
Src Homology 2
- SH3:
-
SRC Homology 3
- SNP:
-
Single Nucelotide Polymorphism
- SOS:
-
Son of Sevenless
- SPRED:
-
Sprouty-Related Enabled/Vasodilator-stimulated Phosphoprotein Homology 1 Domain-Containing Protein
- SPRY:
-
Sprouty
- STAT:
-
Signal Transducer and Activator of Transcription
- TGFβ:
-
Transforming Growth Factor β
- TM:
-
Transmembrane
- VEGFR:
-
Vascular Endothelial Growth Factor Receptor
- XFLRT3:
-
Xenopus Fibronectin Leucine-Rich Transmembrane Protein 3
References
Yayon A, Klagsbrun M, Esko JD, Leder P, Ornitz DM. Cell surface, heparin-like molecules are required for binding of basic fibroblast growth factor to its high affinity receptor. Cell. 1991;64:841–8.
Coughlin SR, Barr PJ, Cousens LS, Fretto LJ, Williams LT. Acidic and basic fibroblast growth factors stimulate tyrosine kinase activity in vivo. J Biol Chem. 1988;263:988–93.
Ornitz DM, Itoh N. Fibroblast growth factors. Genome Biol. 2001; 2: REVIEWS3005.
Ornitz DM, Xu J, Colvin JS, McEwen DG, MacArthur CA, Coulier F, Gao G, Goldfarb M. Receptor specificity of the fibroblast growth factor family. J Biol Chem. 1996;271:15292–7.
Zhang X, Ibrahimi OA, Olsen SK, Umemori H, Mohammadi M, Ornitz DM. Receptor specificity of the fibroblast growth factor family. The complete mammalian FGF family. J Biol Chem. 2006;281:15694–700.
Johnson DE, Williams LT. Structural and functional diversity in the FGF receptor multigene family. Adv Cancer Res. 1993;60:1–41.
Kalinina J, Dutta K, Ilghari D, Beenken A, Goetz R, Eliseenkova AV, Cowburn D, Mohammadi M. The alternatively spliced acid box region plays a key role in FGF receptor autoinhibition. Structure. 2012;20:77–88.
Wheldon LM, Khodabukus N, Patey SJ, Smith TG, Heath JK, Hajihosseini MK. Identification and characterization of an inhibitory fibroblast growth factor receptor 2 (FGFR2) molecule, up-regulated in an Apert Syndrome mouse model. Biochem J. 2011;436:71–81.
Vainikka S, Partanen J, Bellosta P, Coulier F, Birnbaum D, Basilico C, Jaye M, Alitalo K. Fibroblast growth factor receptor-4 shows novel features in genomic structure, ligand binding and signal transduction. EMBO J. 1992;11:4273–80.
Orr-Urtreger A, Bedford M, Burakova T, Arman E, Zimmer Y, Yayon A, Givol D, Lonai P. Developmental localization of the splicing alternatives of fibroblast growth-factor receptor-2 (FGFR2). Dev Biol. 1993;158:475–86.
Yan G, Fukabori Y, McBride G, Nikolaropolous S, McKeehan WL. Exon switching and activation of stromal and embryonic fibroblast growth factor (FGF)-FGF receptor genes in prostate epithelial cells accompany stromal independence and malignancy. Mol Cell Biol. 1993;13:4513–22.
Lindahl U, Hook M. Glycosaminoglycans and their binding to biological macromolecules. Annu Rev Biochem. 1978;47:385–417.
Gambarini AG, Miyamoto CA, Lima GA, Nader HB, Dietrich CP. Mitogenic activity of acidic fibroblast growth factor is enhanced by highly sulfated oligosaccharides derived from heparin and heparan sulfate. Mol Cell Biochem. 1993;124:121–9.
Ornitz DM, Yayon A, Flanagan JG, Svahn CM, Levi E, Leder P. Heparin is required for cell-free binding of basic fibroblast growth factor to a soluble receptor and for mitogenesis in whole cells. Mol Cell Biol. 1992;12:240–7.
Ornitz DM, Herr AB, Nilsson M, Westman J, Svahn CM, Waksman G. FGF binding and FGF receptor activation by synthetic heparan-derived di- and trisaccharides. Science. 1995;268:432–6.
Schlessinger J, Plotnikov AN, Ibrahimi OA, Eliseenkova AV, Yeh BK, Yayon A, Linhardt RJ, Mohammadi M. Crystal structure of a ternary FGF-FGFR-heparin complex reveals a dual role for heparin in FGFR binding and dimerization. Mol Cell. 2000;6:743–50.
Guimond SE, Turnbull JE. Fibroblast growth factor receptor signalling is dictated by specific heparan sulphate saccharides. Curr Biol. 1999;9:1343–6.
Ornitz DM. FGFs, heparan sulfate and FGFRs: complex interactions essential for development. Bioessays. 2000;22:108–12.
McDowell LM, Frazier BA, Studelska DR, Giljum K, Chen J, Liu J, Yu K, Ornitz DM, Zhang L. Inhibition or activation of Apert syndrome FGFR2 (S252W) signaling by specific glycosaminoglycans. J Biol Chem. 2006;281:6924–30.
Guimond S, Maccarana M, Olwin BB, Lindahl U, Rapraeger AC. Activating and inhibitory heparin sequences for FGF-2 (basic FGF). Distinct requirements for FGF-1, FGF-2, and FGF-4. J Biol Chem. 1993;268:23906–14.
Pye DA, Vives RR, Turnbull JE, Hyde P, Gallagher JT. Heparan sulfate oligosaccharides require 6-O-sulfation for promotion of basic fibroblast growth factor mitogenic activity. J Biol Chem. 1998;273:22936–42.
Taylor KR, Rudisill JA, Gallo RL. Structural and sequence motifs in dermatan sulfate for promoting fibroblast growth factor-2 (FGF-2) and FGF-7 activity. J Biol Chem. 2005;280:5300–6.
Trowbridge JM, Rudisill JA, Ron D, Gallo RL. Dermatan sulfate binds and potentiates activity of keratinocyte growth factor (FGF-7). J Biol Chem. 2002;277:42815–20.
Pellegrini L, Burke DF, von Delft F, Mulloy B, Blundell TL. Crystal structure of fibroblast growth factor receptor ectodomain bound to ligand and heparin. Nature. 2000;407:1029–34.
Furdui CM, Lew ED, Schlessinger J, Anderson KS. Autophosphorylation of FGFR1 kinase is mediated by a sequential and precisely ordered reaction. Mol Cell. 2006;21:711–7.
Mohammadi M, Dionne CA, Li W, Li N, Spivak T, Honegger AM, Jaye M, Schlessinger J. Point mutation in FGF receptor eliminates phosphatidylinositol hydrolysis without affecting mitogenesis. Nature. 1992;358:681–4.
Mohammadi M, Schlessinger J, Hubbard SR. Structure of the FGF receptor tyrosine kinase domain reveals a novel autoinhibitory mechanism. Cell. 1996;86:577–87.
Mohammadi M, Dikic I, Sorokin A, Burgess WH, Jaye M, Schlessinger J. Identification of six novel autophosphorylation sites on fibroblast growth factor receptor 1 and elucidation of their importance in receptor activation and signal transduction. Mol Cell Biol. 1996; 16:977–89.
Raffioni S, Thomas D, Foehr ED, Thompson LM, Bradshaw RA. Comparison of the intracellular signaling responses by three chimeric fibroblast growth factor receptors in PC12 cells. Proc Natl Acad Sci USA. 1999;96:7178–83.
Dhalluin C, Yan KS, Plotnikova O, Lee KW, Zeng L, Kuti M, Mujtaba S, Goldfarb MP, Zhou MM. Structural basis of SNT PTB domain interactions with distinct neurotrophic receptors. Mol Cell. 2000;6:921–9.
Ong SH, Guy GR, Hadari YR, Laks S, Gotoh N, Schlessinger J, Lax I. FRS2 proteins recruit intracellular signaling pathways by binding to diverse targets on fibroblast growth factor and nerve growth factor receptors. Mol Cell Biol. 2000;20:979–89.
Gotoh N. Regulation of growth factor signaling by FRS2 family docking/scaffold adaptor proteins. Cancer Sci. 2008;99:1319–25.
Eswarakumar VP, Lax I, Schlessinger J. Cellular signaling by fibroblast growth factor receptors. Cytokine Growth Factor Rev. 2005;16:139–49.
Mohammadi M, Honegger AM, Rotin D, Fischer R, Bellot F, Li W, Dionne CA, Jaye M, Rubinstein M, Schlessinger J. A tyrosine-phosphorylated carboxy-terminal peptide of the fibroblast growth factor receptor (Flg) is a binding site for the SH2 domain of phospholipase C-gamma 1. Mol Cell Biol. 1991;11:5068–78.
Falasca M, Iurisci C, Carvelli A, Sacchetti A, Corda D. Release of the mitogen lysophosphatidylinositol from H-Ras-transformed fibroblasts; a possible mechanism of autocrine control of cell proliferation. Oncogene. 1998;16:2357–65.
Klint P, Claesson-Welsh L. Signal transduction by fibroblast growth factor receptors. Front Biosci. 1999;4:165–77.
Rameh LE, Rhee SG, Spokes K, Kazlauskas A, Cantley LC, Cantley LG. Phosphoinositide 3-kinase regulates phospholipase Cgamma-mediated calcium signaling. J Biol Chem. 1998;273:23750–7.
Ebong S, Yu CR, Carper DA, Chepelinsky AB, Egwuagu CE. Activation of STAT signaling pathways and induction of suppressors of cytokine signaling (SOCS) proteins in mammalian lens by growth factors. Invest Ophthalmol Vis Sci. 2004;45:872–8.
Darnell Jr JE. STATs and gene regulation. Science. 1997;277:1630–5.
Wang L, Rudert WA, Loutaev I, Roginskaya V, Corey SJ. Repression of c-Cbl leads to enhanced G-CSF Jak-STAT signaling without increased cell proliferation. Oncogene. 2002;21:5346–55.
Degnin CR, Laederich MB, Horton WA. Ligand activation leads to regulated intramembrane proteolysis of fibroblast growth factor receptor 3. Mol Biol Cell. 2011;22:3861–73.
Blobel CP, Carpenter G, Freeman M. The role of protease activity in ErbB biology. Exp Cell Res. 2009;315:671–82.
Brown MS, Ye J, Rawson RB, Goldstein JL. Regulated intramembrane proteolysis: a control mechanism conserved from bacteria to humans. Cell. 2000;100:391–8.
Levi E, Fridman R, Miao HQ, Ma YS, Yayon A, Vlodavsky I. Matrix metalloproteinase 2 releases active soluble ectodomain of fibroblast growth factor receptor 1. Proc Natl Acad Sci USA. 1996;93:7069–74.
Maretzky T, Le Gall SM, Worpenberg-Pietruk S, Eder J, Overall CM, Huang XY, Poghosyan Z, Edwards DR, Blobel CP. Src stimulates fibroblast growth factor receptor-2 shedding by an ADAM15 splice variant linked to breast cancer. Cancer Res. 2009;69:4573–6.
Peduto L, Reuter VE, Shaffer DR, Scher HI, Blobel CP. Critical function for ADAM9 in mouse prostate cancer. Cancer Res. 2005;65:9312–9.
Ancot F, Foveau B, Lefebvre J, Leroy C, Tulasne D. Proteolytic cleavages give receptor tyrosine kinases the gift of ubiquity. Oncogene. 2009;28:2185–95.
Loeb CR, Harris JL, Craik CS. Granzyme B proteolyzes receptors important to proliferation and survival, tipping the balance toward apoptosis. J Biol Chem. 2006;281:28326–35.
Chioni AM, Grose R. FGFR1 cleavage and nuclear translocation regulates breast cancer cell behavior. J Cell Biol. 2012;197:801–17.
Hacohen N, Kramer S, Sutherland D, Hiromi Y, Krasnow MA. Sprouty encodes a novel antagonist of FGF signaling that patterns apical branching of the Drosophila airways. Cell. 1998;92:253–63.
Thisse B, Thisse C. Functions and regulations of fibroblast growth factor signaling during embryonic development. Dev Biol. 2005;287:390–402.
Sasaki A, Taketomi T, Kato R, Saeki K, Nonami A, Sasaki M, Kuriyama M, Saito N, Shibuya M, Yoshimura A. Mammalian Sprouty4 suppresses Ras-independent ERK activation by binding to Raf1. Cell Cycle. 2003;2:281–2.
Wakioka T, Sasaki A, Kato R, Shouda T, Matsumoto A, Miyoshi K, Tsuneoka M, Komiya S, Baron R, Yoshimura A. Spred is a Sprouty-related suppressor of Ras signalling. Nature. 2001;412:647–51.
Mardakheh FK, Yekezare M, Machesky LM, Heath JK. Spred2 interaction with the late endosomal protein NBR1 down-regulates fibroblast growth factor receptor signaling. J Cell Biol. 2009;187:265–77.
Yang RB, Ng CK, Wasserman SM, Komuves LG, Gerritsen ME, Topper JN. A novel interleukin-17 receptor-like protein identified in human umbilical vein endothelial cells antagonizes basic fibroblast growth factor-induced signaling. J Biol Chem. 2003;278:33232–8.
Kovalenko D, Yang X, Nadeau RJ, Harkins LK, Friesel R. Sef inhibits fibroblast growth factor signaling by inhibiting FGFR1 tyrosine phosphorylation and subsequent ERK activation. J Biol Chem. 2003;278:14087–91.
Tsang M, Friesel R, Kudoh T, Dawid IB. Identification of Sef, a novel modulator of FGF signalling. Nat Cell Biol. 2002;4:165–9.
Xiong S, Zhao Q, Rong Z, Huang G, Huang Y, Chen P, Zhang S, Liu L, Chang Z. hSef inhibits PC-12 cell differentiation by interfering with Ras-mitogen-activated protein kinase MAPK signaling. J Biol Chem. 2003;278:50273–82.
Torii S, Kusakabe M, Yamamoto T, Maekawa M, Nishida E. Sef is a spatial regulator for Ras/MAP kinase signaling. Dev Cell. 2004;7:33–44.
Buday L, Warne PH, Downward J. Downregulation of the Ras activation pathway by MAP kinase phosphorylation of Sos. Oncogene. 1995;11:1327–31.
Ueki K, Matsuda S, Tobe K, Gotoh Y, Tamemoto H, Yachi M, Akanuma Y, Yazaki Y, Nishida E, Kadowaki T. Feedback regulation of mitogen-activated protein kinase kinase kinase activity of c-Raf-1 by insulin and phorbol ester stimulation. J Biol Chem. 1994;269:15756–61.
Gual P, Giordano S, Anguissola S, Parker PJ, Comoglio PM. Gab1 phosphorylation: a novel mechanism for negative regulation of HGF receptor signaling. Oncogene. 2001;20:156–66.
Plotnikov AN, Schlessinger J, Hubbard SR, Mohammadi M. Structural basis for FGF receptor dimerization and activation. Cell. 1999;98:641–50.
Stauber DJ, DiGabriele AD, Hendrickson WA. Structural interactions of fibroblast growth factor receptor with its ligands. Proc Natl Acad Sci USA. 2000;97:49–54.
Olsen SK, Ibrahimi OA, Raucci A, Zhang F, Eliseenkova AV, Yayon A, Basilico C, Linhardt RJ, Schlessinger J, Mohammadi M. Insights into the molecular basis for fibroblast growth factor receptor autoinhibition and ligand-binding promiscuity. Proc Natl Acad Sci USA. 2004;101:935–40.
Kobrin MS, Yamanaka Y, Friess H, Lopez ME, Korc M. Aberrant expression of type I fibroblast growth factor receptor in human pancreatic adenocarcinomas. Cancer Res. 1993;53:4741–4.
Mansson PE, Adams P, Kan M, McKeehan WL. Heparin-binding growth factor gene expression and receptor characteristics in normal rat prostate and two transplantable rat prostate tumors. Cancer Res. 1989;49:2485–94.
Itoh N, Ornitz DM. Evolution of the Fgf and Fgfr gene families. Trends Genet. 2004;20:563–9.
Goetz R, Beenken A, Ibrahimi OA, Kalinina J, Olsen SK, Eliseenkova AV, Xu C, Neubert TA, Zhang F, Linhardt RJ, Yu X, White KE, Inagaki T, Kliewer SA, Yamamoto M, Kurosu H, Ogawa Y, Kuro-o M, Lanske B, Razzaque MS, Mohammadi M. Molecular insights into the klotho-dependent, endocrine mode of action of fibroblast growth factor 19 subfamily members. Mol Cell Biol. 2007;27:3417–28.
Harmer NJ, Pellegrini L, Chirgadze D, Fernandez-Recio J, Blundell TL. The crystal structure of fibroblast growth factor (FGF) 19 reveals novel features of the FGF family and offers a structural basis for its unusual receptor affinity. Biochemistry. 2004;43:629–40.
Kurosu H, Kuro OM. The Klotho gene family as a regulator of endocrine fibroblast growth factors. Mol Cell Endocrinol. 2009;299:72–8.
Urakawa I, Yamazaki Y, Shimada T, Iijima K, Hasegawa H, Okawa K, Fujita T, Fukumoto S, Yamashita T. Klotho converts canonical FGF receptor into a specific receptor for FGF23. Nature. 2006;444:770–4.
Polanska UM, Edwards E, Fernig DG, Kinnunen TK. The cooperation of FGF receptor and Klotho is involved in excretory canal development and regulation of metabolic homeostasis in Caenorhabditis elegans. J Biol Chem. 2011;286:5657–66.
Polanska UM, Fernig DG, Kinnunen T. Extracellular interactome of the FGF receptor-ligand system: complexities and the relative simplicity of the worm. Dev Dyn. 2009;238:277–93.
Ghabrial A, Luschnig S, Metzstein MM, Krasnow MA. Branching morphogenesis of the Drosophila tracheal system. Annu Rev Cell Dev Biol. 2003;19:623–47.
Bottcher RT, Niehrs C. Fibroblast growth factor signaling during early vertebrate development. Endocr Rev. 2005;26:63–77.
Dorey K, Amaya E. FGF signalling: diverse roles during early vertebrate embryogenesis. Development. 2010;137:3731–42.
Huang P, Stern MJ. FGF signaling in flies and worms: more and more relevant to vertebrate biology. Cytokine Growth Factor Rev. 2005;16:151–8.
DeVore DL, Horvitz HR, Stern MJ. An FGF receptor signaling pathway is required for the normal-cell migrations of the sex myoblasts in C-elegans hermaphrodites. Cell. 1995;83:611–20.
Burdine RD, Branda CS, Stern MJ. EGL-17(FGF) expression coordinates the attraction of the migrating sex myoblasts with vulval induction in C. elegans. Development. 1998;125:1083–93.
Popovici C, Conchonaud F, Birnbaum D, Roubin R. Functional phylogeny relates LET-756 to fibroblast growth factor 9. J Biol Chem. 2004;279:40146–52.
Roubin R, Naert K, Popovici C, Vatcher G, Coulier F, Thierry-Mieg J, Pontarotti P, Birnbaum D, Baillie D, Thierry-Mieg D. let-756, a C. elegans fgf essential for worm development. Oncogene. 1999;18:6741–7.
Kokel M, Borland CZ, DeLong L, Horvitz HR, Stern MJ. clr-1 encodes a receptor tyrosine phosphatase that negatively regulates an FGF receptor signaling pathway in Caenorhabditis elegans. Genes Dev. 1998;12:1425–37.
Borland CZ, Schutzman JL, Stern MJ. Fibroblast growth factor signaling in Caenorhabditis elegans. Bioessays. 2001;23:1120–30.
Chateau MT, Araiz C, Descamps S, Galas S. Klotho interferes with a novel FGF-signalling pathway and insulin/Igf-like signalling to improve longevity and stress resistance in Caenorhabditis elegans. Aging. 2010;2:567–81.
Glazer L, Shilo B-Z. The Drosophila FGF-R homolog is expressed in the embryonic tracheal system and appears to be required for directed tracheal cell extension. Genes Dev. 1991;5:697–705.
Sutherland D, Samakovlis C, Krasnow MA. branchless encodes a Drosophila FGF homolog that controls tracheal cell migration and the pattern of branching. Cell. 1996;87:1091–101.
Ghabrial AS, Krasnow MA. Social interactions among epithelial cells during tracheal branching morphogenesis. Nature. 2006;441:746–9.
Tsang M, Dawid IB. Promotion and attenuation of FGF signaling through the Ras-MAPK pathway. Sci STKE. 2004:pe17.
Beiman M, Shilo BZ, Volk T. Heartless, a Drosophila FGF receptor homolog, is essential for cell migration and establishment of several mesodermal lineages. Genes Dev. 1996;10:2993–3002.
Gisselbrecht S, Skeath JB, Doe CQ, Michelson AM. Heartless encodes a fibroblast growth factor receptor (DFR1/DFGF-R2) involved in the directional migration of early mesodermal cells in the Drosophila embryo. Genes Dev. 1996;10:3003–17.
Gryzik T, Muller HA. FGF8-like1 and FGF8-like2 encode putative ligands of the FGF receptor Htl and are required for mesoderm migration in the Drosophila gastrula. Curr Biol. 2004;14:659–67.
Stathopoulos A, Tam B, Ronshaugen M, Frasch M, Levine M. pyramus and thisbe: FGF genes that pattern the mesoderm of Drosophila embryos. Genes Dev. 2004;18:687–99.
Vincent S, Wilson R, Coelho C, Affolter M, Leptin M. The Drosophila protein Dof is specifically required for FGF signaling. Mol Cell. 1998;2:515–25.
Rodaway A, Takeda H, Koshida S, Broadbent J, Price B, Smith JC, Patient R, Holder N. Induction of the mesendoderm in the zebrafish germ ring by yolk cell-derived TGF-beta family signals and discrimination of mesoderm and endoderm by FGF. Development. 1999;126:3067–78.
Griffin K, Patient R, Holder N. Analysis of FGF function in normal and no tail zebrafish embryos reveals separate mechanisms for formation of the trunk and the tail. Development. 1995;121:2983–94.
Griffin KJ, Amacher SL, Kimmel CB, Kimelman D. Molecular identification of spadetail: regulation of zebrafish trunk and tail mesoderm formation by T-box genes. Development. 1998;125:3379–88.
Furthauer M, Thisse C, Thisse B. A role for FGF-8 in the dorsoventral patterning of the zebrafish gastrula. Development. 1997;124:4253–64.
Furthauer M, Lin W, Ang SL, Thisse B, Thisse C. Sef is a feedback-induced antagonist of Ras/MAPK-mediated FGF signalling. Nat Cell Biol. 2002;4:170–4.
Brand M, Heisenberg CP, Jiang YJ, Beuchle D, Lun K, Furutani-Seiki M, Granato M, Haffter P, Hammerschmidt M, Kane DA, Kelsh RN, Mullins MC, Odenthal J, van Eeden FJ, Nusslein-Volhard C. Mutations in zebrafish genes affecting the formation of the boundary between midbrain and hindbrain. Development. 1996;123:179–90.
Reifers F, Bohli H, Walsh EC, Crossley PH, Stainier DY, Brand M. Fgf8 is mutated in zebrafish acerebellar (ace) mutants and is required for maintenance of midbrain-hindbrain boundary development and somitogenesis. Development. 1998;125:2381–95.
Shanmugalingam S, Houart C, Picker A, Reifers F, Macdonald R, Barth A, Griffin K, Brand M, Wilson SW. Ace/Fgf8 is required for forebrain commissure formation and patterning of the telencephalon. Development. 2000;127:2549–61.
Kimelman D, Kirschner M. Synergistic induction of mesoderm by FGF and TGF-beta and the identification of an mRNA coding for FGF in the early Xenopus embryo. Cell. 1987;51:869–77.
Slack JM, Darlington BG, Heath JK, Godsave SF. Mesoderm induction in early Xenopus embryos by heparin-binding growth factors. Nature. 1987;326:197–200.
Amaya E, Musci T, Kirschner M. Expression of a dominant negative mutant of the FGF receptor disrupts mesoderm formation in xenopus embryos. Cell. 1991;66:257–70.
Smith JC, Price BM, Green JB, Weigel D, Herrmann BG. Expression of a Xenopus homolog of Brachyury (T) is an immediate-early response to mesoderm induction. Cell. 1991;67:79–87.
Sivak JM, Petersen LF, Amaya E. FGF signal interpretation is directed by Sprouty and Spred proteins during mesoderm formation. Dev Cell. 2005;8:689–701.
Fisher ME, Isaacs HV, Pownall ME. eFGF is required for activation of XmyoD expression in the myogenic cell lineage of Xenopus laevis. Development. 2002;129:1307–15.
Fletcher RB, Baker JC, Harland RM. FGF8 spliceforms mediate early mesoderm and posterior neural tissue formation in Xenopus. Development. 2006;133:1703–14.
Amaya E, Stein PA, Musci TJ, Kirschner MW. FGF signalling in the early specification of mesoderm in Xenopus. Development. 1993;118:477–87.
Hongo I, Kengaku M, Okamoto H. FGF signaling and the anterior neural induction in Xenopus. Dev Biol. 1999;216:561–81.
Cohn MJ, Izpisua-Belmonte JC, Abud H, Heath JK, Tickle C. Fibroblast growth factors induce additional limb development from the flank of chick embryos. Cell. 1995;80:739–46.
Laufer E, Nelson CE, Johnson RL, Morgan BA, Tabin C. Sonic hedgehog and Fgf-4 act through a signaling cascade and feedback loop to integrate growth and patterning of the developing limb bud. Cell. 1994;79:993–1003.
Benazeraf B, Francois P, Baker RE, Denans N, Little CD, Pourquie O. A random cell motility gradient downstream of FGF controls elongation of an amniote embryo. Nature. 2010;466:248–52.
Dubrulle J, McGrew MJ, Pourquie O. FGF signaling controls somite boundary position and regulates segmentation clock control of spatiotemporal Hox gene activation. Cell. 2001;106:219–32.
Dubrulle J, Pourquie O. fgf8 mRNA decay establishes a gradient that couples axial elongation to patterning in the vertebrate embryo. Nature. 2004;427:419–22.
Streit A, Berliner AJ, Papanayotou C, Sirulnik A, Stern CD. Initiation of neural induction by FGF signalling before gastrulation. Nature. 2000;406:74–8.
Wilson SI, Graziano E, Harland R, Jessell TM, Edlund T. An early requirement for FGF signalling in the acquisition of neural cell fate in the chick embryo. Curr Biol. 2000;10:421–9.
Itoh N, Ornitz DM. Fibroblast growth factors: from molecular evolution to roles in development, metabolism and disease. J Biochem. 2010;149:121–30.
Elfenbein A, Simons M. Auxiliary and autonomous proteoglycan signaling networks. Methods Enzymol. 2010;480:3–31.
Elfenbein A, Lanahan A, Zhou TX, Yamasaki A, Tkachenko E, Matsuda M, Simons M. Syndecan 4 regulates FGFR1 signaling in endothelial cells by directing macropinocytosis. Sci Signal. 2012;5:ra36.
Bryant DM, Stow JL. Nuclear translocation of cell-surface receptors: lessons from fibroblast growth factor. Traffic. 2005;6:947–54.
Reilly JF, Maher PA. Importin beta-mediated nuclear import of fibroblast growth factor receptor: role in cell proliferation. J Cell Biol. 2001;152:1307–12.
Stachowiak MK, Maher PA, Stachowiak EK. Integrative nuclear signaling in cell development–a role for FGF receptor-1. DNA Cell Biol. 2007;26:811–26.
Wilkie AO. Bad bones, absent smell, selfish testes: the pleiotropic consequences of human FGF receptor mutations. Cytokine Growth Factor Rev. 2005;16:187–203.
Ibrahimi OA, Eliseenkova AV, Plotnikov AN, Yu K, Ornitz DM, Mohammadi M. Structural basis for fibroblast growth factor receptor 2 activation in Apert syndrome. Proc Natl Acad Sci USA. 2001;98:7182–7.
Yu K, Herr AB, Waksman G, Ornitz DM. Loss of fibroblast growth factor receptor 2 ligand-binding specificity in Apert syndrome. Proc Natl Acad Sci USA. 2000;97:14536–41.
Oldridge M, Lunt PW, Zackai EH, McDonald-McGinn DM, Muenke M, Moloney DM, Twigg SR, Heath JK, Howard TD, Hoganson G, Gagnon DM, Jabs EW, Wilkie AO. Genotype-phenotype correlation for nucleotide substitutions in the IgII-IgIII linker of FGFR2. Hum Mol Genet. 1997;6:137–43.
Webster MK, Donoghue DJ. FGFR activation in skeletal disorders: too much of a good thing. Trends Genet. 1997;13:178–82.
Wang Y, Xiao R, Yang F, Karim BO, Iacovelli AJ, Cai J, Lerner CP, Richtsmeier JT, Leszl JM, Hill CA, Yu K, Ornitz DM, Elisseeff J, Huso DL, Jabs EW. Abnormalities in cartilage and bone development in the Apert syndrome FGFR2(+/S252W) mouse. Development. 2005;132:3537–48.
Ahmed Z, Schuller AC, Suhling K, Tregidgo C, Ladbury JE. Extracellular point mutations in FGFR2 elicit unexpected changes in intracellular signalling. Biochem J. 2008;413:37–49.
Pollock PM, Gartside MG, Dejeza LC, Powell MA, Mallon MA, Davies H, Mohammadi M, Futreal PA, Stratton MR, Trent JM, Goodfellow PJ. Frequent activating FGFR2 mutations in endometrial carcinomas parallel germline mutations associated with craniosynostosis and skeletal dysplasia syndromes. Oncogene. 2007;26:7158–62.
Greulich H, Pollock PM. Targeting mutant fibroblast growth factor receptors in cancer. Trends Mol Med. 2011;17:283–92.
Wilkie AO, Patey SJ, Kan SH, van den Ouweland AM, Hamel BC. FGFs, their receptors, and human limb malformations: clinical and molecular correlations. Am J Med Genet. 2002;112:266–78.
Rousseau F, el Ghouzzi V, Delezoide AL, Legeai-Mallet L, Le Merrer M, Munnich A, Bonaventure J. Missense FGFR3 mutations create cysteine residues in thanatophoric dwarfism type I (TD1). Hum Mol Genet. 1996;5:509–12.
Tavormina PL, Shiang R, Thompson LM, Zhu YZ, Wilkin DJ, Lachman RS, Wilcox WR, Rimoin DL, Cohn DH, Wasmuth JJ. Thanatophoric dysplasia (types I and II) caused by distinct mutations in fibroblast growth factor receptor 3. Nat Genet. 1995;9:321–8.
Rousseau F, Bonaventure J, Legeai-Mallet L, Pelet A, Rozet JM, Maroteaux P, Le Merrer M, Munnich A. Mutations of the fibroblast growth factor receptor-3 gene in achondroplasia. Horm Res. 1996;45:108–10.
Shiang R, Thompson LM, Zhu YZ, Church DM, Fielder TJ, Bocian M, Winokur ST, Wasmuth JJ. Mutations in the transmembrane domain of FGFR3 cause the most common genetic form of dwarfism, achondroplasia. Cell. 1994;78:335–42.
Knowles MA. Novel therapeutic targets in bladder cancer: mutation and expression of FGF receptors. Future Oncol. 2008;4:71–83.
Meyers GA, Orlow SJ, Munro IR, Przylepa KA, Jabs EW. Fibroblast growth factor receptor 3 (FGFR3) transmembrane mutation in Crouzon syndrome with acanthosis nigricans. Nat Genet. 1995;11:462–4.
van Rhijn BW, van Tilborg AA, Lurkin I, Bonaventure J, de Vries A, Thiery JP, van der Kwast TH, Zwarthoff EC, Radvanyi F. Novel fibroblast growth factor receptor 3 (FGFR3) mutations in bladder cancer previously identified in non-lethal skeletal disorders. Eur J Hum Genet. 2002;10:819–24.
Li E, You M, Hristova K. FGFR3 dimer stabilization due to a single amino acid pathogenic mutation. J Mol Biol. 2006;356:600–12.
Chen F, Degnin C, Laederich M, Horton WA, Hristova K. The A391E mutation enhances FGFR3 activation in the absence of ligand. Biochim Biophys Acta. 2011;1808:2045–50.
Chen Q, Jiang Y, An Y, Zhao N, Zhao Y, Yu C. Soluble FGFR4 extracellular domain inhibits FGF19-induced activation of FGFR4 signaling and prevents nonalcoholic fatty liver disease. Biochem Biophys Res Commun. 2011;409:651–6.
Zhu Q, Liu T. Fibroblast growth factor receptor 4 polymorphisms and coronary artery disease: a case control study. Mol Biol Rep. 2012;39:8679–85.
Wang J, Yu W, Cai Y, Ren C, Ittmann MM. Altered fibroblast growth factor receptor 4 stability promotes prostate cancer progression. Neoplasia. 2008;10:847–56.
Frullanti E, Berking C, Harbeck N, Jezequel P, Haugen A, Mawrin C, Parise Jr O, Sasaki H, Tsuchiya N, Dragani TA. Meta and pooled analyses of FGFR4 Gly388Arg polymorphism as a cancer prognostic factor. Eur J Cancer Prev. 2011;20:340–7.
Sleeman M, Fraser J, McDonald M, Yuan S, White D, Grandison P, Kumble K, Watson JD, Murison JG. Identification of a new fibroblast growth factor receptor, FGFR5. Gene. 2001;271:171–82.
Trueb B, Zhuang L, Taeschler S, Wiedemann M. Characterization of FGFRL1, a novel fibroblast growth factor (FGF) receptor preferentially expressed in skeletal tissues. J Biol Chem. 2003;278:33857–65.
Wiedemann M, Trueb B. Characterization of a novel protein (FGFRL1) from human cartilage related to FGF receptors. Genomics. 2000;69:275–9.
Rieckmann T, Zhuang L, Fluck CE, Trueb B. Characterization of the first FGFRL1 mutation identified in a craniosynostosis patient. Biochim Biophys Acta. 2009;1792:112–21.
Zhuang L, Karotki AV, Bruecker P, Trueb B. Comparison of the receptor FGFRL1 from sea urchins and humans illustrates evolution of a zinc binding motif in the intracellular domain. BMC Biochem. 2009;10:33.
Steinberg F, Zhuang L, Beyeler M, Kalin RE, Mullis PE, Brandli AW, Trueb B. The FGFRL1 receptor is shed from cell membranes, binds fibroblast growth factors (FGFs), and antagonizes FGF signaling in Xenopus embryos. J Biol Chem. 2010;285:2193–202.
Ibrahimi OA, Zhang F, Eliseenkova AV, Itoh N, Linhardt RJ, Mohammadi M. Biochemical analysis of pathogenic ligand-dependent FGFR2 mutations suggests distinct pathophysiological mechanisms for craniofacial and limb abnormalities. Hum Mol Genet. 2004;13:2313–24.
Trueb B. Biology of FGFRL1, the fifth fibroblast growth factor receptor. Cell Mol Life Sci. 2011;68:951–64.
Powell AK, Fernig DG, Turnbull JE. Fibroblast growth factor receptors 1 and 2 interact differently with heparin/heparan sulfate. Implications for dynamic assembly of a ternary signaling complex. J Biol Chem. 2002;277:28554–63.
Rieckmann T, Kotevic I, Trueb B. The cell surface receptor FGFRL1 forms constitutive dimers that promote cell adhesion. Exp Cell Res. 2008;314:1071–81.
Beenken A, Mohammadi M. The FGF family: biology, pathophysiology and therapy. Nat Rev Drug Discov. 2009;8:235–53.
Schild C, Trueb B. Aberrant expression of FGFRL1, a novel FGF receptor, in ovarian tumors. Int J Mol Med. 2005;16:1169–73.
Gartside MG, Chen H, Ibrahimi OA, Byron SA, Curtis AV, Wellens CL, Bengston A, Yudt LM, Eliseenkova AV, Ma J, Curtin JA, Hyder P, Harper UL, Riedesel E, Mann GJ, Trent JM, Bastian BC, Meltzer PS, Mohammadi M, Pollock PM. Loss-of-function fibroblast growth factor receptor-2 mutations in melanoma. Mol Cancer Res. 2009;7:41–54.
Turner N, Grose R. Fibroblast growth factor signalling: from development to cancer. Nat Rev Cancer. 2010;10:116–29.
Grose R, Dickson C. Fibroblast growth factor signaling in tumorigenesis. Cytokine Growth Factor Rev. 2005;16:179–86.
Fioretos T, Panagopoulos I, Lassen C, Swedin A, Billstrom R, Isaksson M, Strombeck B, Olofsson T, Mitelman F, Johansson B. Fusion of the BCR and the fibroblast growth factor receptor-1 (FGFR1) genes as a result of t(8;22)(p11;q11) in a myeloproliferative disorder: the first fusion gene involving BCR but not ABL. Genes Chromosomes Cancer. 2001;32:302–10.
Courjal F, Cuny M, Simony-Lafontaine J, Louason G, Speiser P, Zeillinger R, Rodriguez C, Theillet C. Mapping of DNA amplifications at 15 chromosomal localizations in 1875 breast tumors: definition of phenotypic groups. Cancer Res. 1997;57:4360–7.
Turner N, Pearson A, Sharpe R, Lambros M, Geyer F, Lopez-Garcia MA, Natrajan R, Marchio C, Iorns E, Mackay A, Gillett C, Grigoriadis A, Tutt A, Reis-Filho JS, Ashworth A. FGFR1 amplification drives endocrine therapy resistance and is a therapeutic target in breast cancer. Cancer Res. 2010;70:2085–94.
Zecchini S, Bombardelli L, Decio A, Bianchi M, Mazzarol G, Sanguineti F, Aletti G, Maddaluno L, Berezin V, Bock E, Casadio C, Viale G, Colombo N, Giavazzi R, Cavallaro U. The adhesion molecule NCAM promotes ovarian cancer progression via FGFR signalling. EMBO Mol Med. 2011;3:480–94.
Francavilla C, Loeffler S, Piccini D, Kren A, Christofori G, Cavallaro U. Neural cell adhesion molecule regulates the cellular response to fibroblast growth factor. J Cell Sci. 2007;120:4388–94.
Francavilla C, Cattaneo P, Berezin V, Bock E, Ami D, de Marco A, Christofori G, Cavallaro U. The binding of NCAM to FGFR1 induces a specific cellular response mediated by receptor trafficking. J Cell Biol. 2009;187:1101–16.
Christensen C, Lauridsen JB, Berezin V, Bock E, Kiselyov VV. The neural cell adhesion molecule binds to fibroblast growth factor receptor 2. FEBS Lett. 2006;580:3386–90.
Byron SA, Gartside MG, Wellens CL, Mallon MA, Keenan JB, Powell MA, Goodfellow PJ, Pollock PM. Inhibition of activated fibroblast growth factor receptor 2 in endometrial cancer cells induces cell death despite PTEN abrogation. Cancer Res. 2008;68:6902–7.
Trudel S, Li ZH, Wei E, Wiesmann M, Chang H, Chen C, Reece D, Heise C, Stewart AK. CHIR-258, a novel, multitargeted tyrosine kinase inhibitor for the potential treatment of t(4;14) multiple myeloma. Blood. 2005;105:2941–8.
Fabbro D, Manley PW. Su-6668. SUGEN. Curr Opin Investig Drugs. 2001;2:1142–8.
Gavine PR, Mooney L, Kilgour E, Thomas AP, Al-Kadhimi K, Beck S, Rooney C, Coleman T, Baker D, Mellor MJ, Brooks AN, Klinowska T. AZD4547: an orally bioavailable, potent, and selective inhibitor of the fibroblast growth factor receptor tyrosine kinase family. Cancer Res. 2012;72:2045–56.
Celli G, Larochelle WJ, MacKem S, Sharp R, Merlino G. Soluble dominant-negative receptor uncovers essential roles for fibroblast growth-factors in multiorgan induction and patterning. EMBO J. 1998;17:1642–55.
Ciruna B, Rossant J. FGF signaling regulates mesoderm cell fate specification and morphogenetic movement at the primitive streak. Dev Cell. 2001;1:37–49.
Ciruna BG, Schwartz L, Harpal K, Yamaguchi TP, Rossant J. Chimeric analysis of fibroblast growth-factor receptor-1 (FGFR1) function—a role for FGFR1 in morphogenetic movement through the primitive streak. Development. 1997;124:2829–41.
Deng CX, Wynshawboris A, Shen MM, Daugherty C, Ornitz DM, Leder P. Murine FGFR-1 is required for early postimplantation growth and axial organization. Genes Dev. 1994;8:3045–57.
Freeman KW, Gangula RD, Welm BE, Ozen M, Foster BA, Rosen JM, Ittmann M, Greenberg NM, Spencer DM. Conditional activation of fibroblast growth factor receptor (FGFR) 1, but not FGFR2, in prostate cancer cells leads to increased osteopontin induction, extracellular signal-regulated kinase activation, and in vivo proliferation. Cancer Res. 2003;63:6237–43.
Freeman KW, Welm BE, Gangula RD, Rosen JM, Ittmann M, Greenberg NM, Spencer DM. Inducible prostate intraepithelial neoplasia with reversible hyperplasia in conditional FGFR1-expressing mice. Cancer Res. 2003;63:8256–63.
Partanen J, Schwartz L, Rossant J. Opposite phenotypes of hypomorphic and Y766 phosphorylation site mutations reveal a function for Fgfr1 in anteroposterior patterning of mouse embryos. Genes Dev. 1998;12:2332–44.
Robinson ML, MacMillancrow LA, Thompson JA, Overbeek PA. Expression of a truncated FGF receptor results in defective lens development in transgenic mice. Development. 1995;121:3959–67.
Verheyden JM, Lewandoski M, Deng C, Harfe BD, Sun X. Conditional inactivation of Fgfr1 in mouse defines its role in limb bud establishment, outgrowth and digit patterning. Development. 2005;132:4235–45.
Welm BE, Freeman KW, Chen M, Contreras A, Spencer DM, Rosen JM. Inducible dimerization of FGFR1: development of a mouse model to analyze progressive transformation of the mammary gland. J Cell Biol. 2002;157:703–14.
Winter SF, Acevedo VD, Gangula RD, Freeman KW, Spencer DM, Greenberg NM. Conditional activation of FGFR1 in the prostate epithelium induces angiogenesis with concomitant differential regulation of Ang-1 and Ang-2. Oncogene. 2007;26:4897–907.
Xu X, Li C, Takahashi K, Slavkin HC, Shum L, Deng CX. Murine fibroblast growth factor receptor 1alpha isoforms mediate node regression and are essential for posterior mesoderm development. Dev Biol. 1999;208:293–306.
Yamaguchi TP, Harpal K, Henkemeyer M, Rossant J. FGFR-1 is required for embryonic growth and mesodermal patterning during mouse gastrulation. Genes Dev. 1994;8:3032–44.
Yang J, Meyer M, Muller AK, Bohm F, Grose R, Dauwalder T, Verrey F, Kopf M, Partanen J, Bloch W, Ornitz DM, Werner S. Fibroblast growth factor receptors 1 and 2 in keratinocytes control the epidermal barrier and cutaneous homeostasis. J Cell Biol. 2010;188:935–52.
Zhang H, Dessimoz J, Beyer TA, Krampert M, Williams LT, Werner S, Grose R. Fibroblast growth factor receptor 1-IIIb is dispensable for skin morphogenesis and wound healing. Eur J Cell Biol. 2004;83:3–11.
Zhao H, Kegg H, Grady S, Truong HT, Robinson ML, Baum M, Bates CM. Role of fibroblast growth factor receptors 1 and 2 in the ureteric bud. Dev Biol. 2004;276:403–15.
De Moerlooze L, Spencer-Dene B, Revest J, Hajihosseini M, Rosewell I, Dickson C. An important role for the IIIb isoform of fibroblast growth factor receptor 2 (FGFR2) in mesenchymal-epithelial signalling during mouse organogenesis. Development. 2000;127:483–92.
Eswarakumar VP, Monsonego-Ornan E, Pines M, Antonopoulou I, Morriss-Kay GM, Lonai P. The IIIc alternative of Fgfr2 is a positive regulator of bone formation. Development. 2002;129:3783–93.
Jackson D, Bresnick J, Rosewell I, Crafton T, Poulsom R, Stamp G, Dickson C. Fibroblast growth factor receptor signalling has a role in lobuloalveolar development of the mammary gland. J Cell Sci. 1997;110(Pt 11):1261–8.
Lin Y, Liu G, Zhang Y, Hu YP, Yu K, Lin C, McKeehan K, Xuan JW, Ornitz DM, Shen MM, Greenberg N, McKeehan WL, Wang F. Fibroblast growth factor receptor 2 tyrosine kinase is required for prostatic morphogenesis and the acquisition of strict androgen dependency for adult tissue homeostasis. Development. 2007;134:723–34.
Mailleux AA, Spencer-Dene B, Dillon C, Ndiaye D, Savona-Baron C, Itoh N, Kato S, Dickson C, Thiery JP, Bellusci S. Role of FGF10/FGFR2b signaling during mammary gland development in the mouse embryo. Development. 2002;129:53–60.
Parsa S, Kuremoto K, Seidel K, Tabatabai R, Mackenzie B, Yamaza T, Akiyama K, Branch J, Koh CJ, Al Alam D, Klein OD, Bellusci S. Signaling by FGFR2b controls the regenerative capacity of adult mouse incisors. Development. 2010;137:3743–52.
Peters K, Werner S, Liao X, Wert S, Whitsett J, Williams L. Targeted expression of a dominant negative FGF receptor blocks branching morphogenesis and epithelial differentiation of the mouse lung. EMBO J. 1994;13:3296–301.
Revest JM, Spencer-Dene B, Kerr K, De Moerlooze L, Rosewell I, Dickson C. Fibroblast growth factor receptor 2-IIIb acts upstream of Shh and Fgf4 and is required for limb bud maintenance but not for the induction of Fgf8, Fgf10, Msx1, or Bmp4. Dev Biol. 2001;231:47–62.
Werner S, Smola H, Liao X, Longaker M, Krieg T, Hofschneider P, Williams L. The function of KGF in morphogenesis of epithelium and reepithelialization of wounds. Science. 1994;266:819–22.
Werner S, Weinberg W, Liao X, Peters K, Blessing M, Yuspa S, Weiner R, Williams L. Targeted expression of a dominant-negative FGF receptor mutant in the epidermis of transgenic mice reveals a role of FGF in keratinocyte organization and differentiation. EMBO J. 1993;12:2635–43.
Xu XL, Weinstein M, Li CL, Naski M, Cohen RI, Ornitz DM, Leder P, Deng CX. Fibroblast-growth-factor-receptor-2 (FGFR2)-mediated reciprocal regulation loop between FGF8 and FGF10 is essential for limb induction. Development. 1998;125:753–65.
Yu K, Xu J, Liu Z, Sosic D, Shao J, Olson EN, Towler DA, Ornitz DM. Conditional inactivation of FGF receptor 2 reveals an essential role for FGF signaling in the regulation of osteoblast function and bone growth. Development. 2003;130:3063–74.
Colvin JS, Bohne BA, Harding GW, McEwen DG, Ornitz DM. Skeletal overgrowth and deafness in mice lacking fibroblast growth factor receptor 3. Nat Genet. 1996;12:390–7.
Deng C, Wynshaw BA, Zhou F, Kuo A, Leder P. Fibroblast growth factor receptor 3 is a negative regulator of bone growth. Cell. 1996;84:911–21.
L’Hote CG, Knowles MA. Cell responses to FGFR3 signalling: growth, differentiation and apoptosis. Exp Cell Res. 2005;304:417–31.
Su N, Xu X, Li C, He Q, Zhao L, Chen S, Luo F, Yi L, Du X, Huang H, Deng C, Chen L. Generation of Fgfr3 conditional knockout mice. Int J Biol Sci. 2010;6:327–32.
Wang JM, Du XL, Li CL, Yin LJ, Chen B, Sun J, Su N, Zhao L, Song RH, Song WW, Chen L, Deng CX. Gly374Arg mutation in Fgfr3 causes achondroplasia in mice. Zhonghua Yi Xue Yi Chuan Xue Za Zhi. 2004;21:537–41.
Wang Y, Spatz MK, Kannan K, Hayk H, Avivi A, Gorivodsky M, Pines M, Yayon A, Lonai P, Givol D. A mouse model for achondroplasia produced by targeting fibroblast growth factor receptor 3. Proc Natl Acad Sci USA. 1999;96:4455–60.
Weinstein M, Xu XL, Ohyama K, Deng CX. FGFR-3 and FGFR-4 function cooperatively to direct alveogenesis in the murine lung. Development. 1998;125:3615–23.
Yu C, Wang F, Jin C, Wu X, Chan WK, McKeehan WL. Increased carbon tetrachloride-induced liver injury and fibrosis in FGFR4-deficient mice. Am J Pathol. 2002;161:2003–10.
Yu C, Wang F, Kan M, Jin C, Jones RB, Weinstein M, Deng CX, McKeehan WL. Elevated cholesterol metabolism and bile acid synthesis in mice lacking membrane tyrosine kinase receptor FGFR4. J Biol Chem. 2000;275:15482–9.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
FGFR1–4 at a glance
FGFR1–4 at a glance
FGFR1 | FGFR2 | FGFR3 | FGFR4 | |
|---|---|---|---|---|
Alternative names | BFGFR, FLT2, FGFBR, FLG, CEK, CD331, H2, H3, H4, KAL2, N-SAM, c-Fgr, HBGFR, FLJ14326 | BEK, KGFR, KSAM, CD332, BFR-1, CEK3, CFD1, ECT1, TK14, TK25 | CD333, JTK4, ACH, CEK2, HBGFR | CD334, JTK2, TKF |
Chromosome location | 8p12 | 10q26 | 4p16.3 | 5q35.1-qter |
Gene size (bp) | 57,696 | 120,128 | 15,560 | 11,206 |
Intron/exon number | 18 exons | 18 exons | 17 exons | 18 exons |
mRNA size (5′, ORF, 3′) | Up to ~5900 bp | Up to ~4250 bp | Up to ~4150 bp | Up to ~3120 bp |
Amino acid number | Up to 853 | Up to 830 | Up to 808 | Up to 802 |
Protein size | Up to 95 kDa | Up to 93 kDa | Up to 88 kDa | 88 kDa |
Post-translational modifications | Autophosphorylated, ubiquitinated, N-glycosylated | Autophosphorylated, ubiquitinated, N-glycosylated | Autophosphorylated, ubiquitinated, N-glycosylated | Autophosphorylated, ubiquitinated, N-glycosylated |
Domains | Up to three Ig domains, transmembrane domain, tyrosine kinase domain | Up to three Ig domains, transmembrane domain, tyrosine kinase domain | Up to three Ig domains, transmembrane domain, tyrosine kinase domain | Up to three Ig domains, transmembrane domain, tyrosine kinase domain |
Pathways activated | MAPK, PI3K/AKT, PLCγ, STAT | MAPK, PI3K/AKT, PLCγ, STAT | MAPK, PI3K/AKT, PLCγ, STAT | MAPK, PI3K/AKT, PLCγ, STAT |
Knockout mouse phenotype | FGFR1−/− embryonic lethal around gastrulation | FGFR2−/− embryonic lethal around gastrulation | FGFR3 mutant alleles show skeletal phenotypes and hearing defects | FGFR4 null mice show no overt phenotype barring a reduction in body weight |
FGFR1-IIIb−/− no phenotype | FGFR2-IIIb−/− multiple defects in organogenesis | Conditional knockout available | ||
Conditional knockouts show multiple phenotypes in brain, limb and bone | Isoform specific conditional knockouts exist for IIIb, showing multiple phenotypes, and for IIIc, mimicking IIIb activity in Apert Syndrome | |||
Conditional knockout of entire gene gives multiple phenotypes | ||||
References |
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Fearon, A.E., Chioni, AM., Grose, R.P. (2015). The FGFR Receptor Family. In: Wheeler, D., Yarden, Y. (eds) Receptor Tyrosine Kinases: Family and Subfamilies. Springer, Cham. https://doi.org/10.1007/978-3-319-11888-8_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-11888-8_6
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-11887-1
Online ISBN: 978-3-319-11888-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)