Abstract
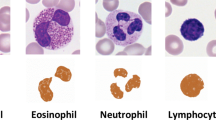
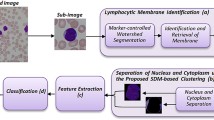
Features that are widely used in digital image analysis and pattern recognition tasks are from three main categories: shape, intensity, and texture invariant features. For computer-aided diagnosis in medical imaging for many specific types of medical problem, the most effective choice of a subset of these features through feature selection is still an open problem. In this work, we consider the problem of white blood cell (leukocyte) recognition into their five primary types: Neutrophils, Lymphocytes, Eosinophils, Monocytes and Basophils using a Support Vector Machine classifier. For features, we use four main intensity histogram calculations, set of 11 invariant moments, the relative area, co-occurrence and run-length matrices, dual tree complex wavelet transform, Haralick and Tamura features. Global sensitivity analysis using Sobol’s RS-HDMR which can deal with independent and dependent input variables is used to assess dominate discriminatory power and the reliability of feature models in presence of high dimensional input feature data to build an efficient feature selection. Both the numerical and empirical results of experiments are compared with forward sequential feature selection. Finally, the results obtained from the preliminary analysis of white blood cell classification are presented in confusion matrices and interpreted using Cohen’s kappa (κ) with the classification framework being validated with experiments conducted on poor quality white blood cell images.
Chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
References
Aliş, Ö., Rabitz, H.: Efficient implementation of high dimensional model representations. Journal of Mathematical Chemistry 29(2), 127–142 (2001)
Bacusmber, J.W., Gose, E.E.: Leukocyte pattern recognition. IEEE Transactions on Systems, Man and Cybernetics SMC-2(4), 513–526 (1972)
Ben-Hur, A., Weston, J.: A user’s guide to support vector machines. In: Data Mining Techniques for the Life Sciences, Methods in Molecular Biology
Bouatmane, S., Roula, M., Bouridane, A., Al-Maadeed, S.: Round-robin sequential forward selection algorithm for prostate cancer classification and diagnosis using multispectral imagery. Machine Vision and Applications 22(5), 865–878 (2011)
Buttarello, M., Plebani, M.: Automated blood cell counts -state of the art. American Journal of Clinical Pathology 130, 104–116 (2008)
Choi, K.S., Zeng, Y., Qin, J.: Using sequential floating forward selection algorithm to detect epileptic seizure in EEG signals. In: 11th International Conference on Signal Processing (ICSP), vol. 3, pp. 1637–1640 (2012)
Comaniciu, D., Meer, P.: Cell image segmentation for diagnostic pathology. In: Advanced Algorithmic Approaches to Medical Image Segmentation, pp. 541–558. Springer, New York (2002)
Dangott, B., Salama, M., Ramesh, N., Tasdizen, T.: Isolation and two-step classification of normal white blood cells in peripheral blood smears. Journal of Pathology Informatics 3(1), 13 (2012)
Dorini, L.B., Minetto, R., Leite, N.J.: Semi-automatic white blood cell segmentation based on multiscale analysis. IEEE Journal of Biomedical and Health Informatics 17(1), 250–256 (2013)
Fu, B., Zhou, J., Li, Y., Zhang, G., Wang, C.: Image analysis by modified legendre moments. Pattern Recognition 40(2), 691–704 (2007)
Grimaldi, E., Scopacasa, F.: Evaluation of the abbott CELL-DYN 4000 hematology analyzer. American Journal of Clinical Pathology 113(4), 497–505 (2000)
Habibzadeh, M., Krzyżak, A., Fevens, T.: Analysis of white blood cell differential counts using dual-tree complex wavelet transform and support vector machine classifier. In: Bolc, L., Tadeusiewicz, R., Chmielewski, L.J., Wojciechowski, K. (eds.) ICCVG 2012. LNCS, vol. 7594, pp. 414–422. Springer, Heidelberg (2012)
Habibzadeh, M., Krzyżak, A., Fevens, T.: Comparative study of shape, intensity and texture features and support vector machine for white blood cell classification. Journal of Theoretical and Applied Computer Science 7, 20–35 (2013)
Habibzadeh, M., Krzyżak, A., Fevens, T., Sadr, A.: Counting of RBCs and WBCs in noisy normal blood smear microscopic images. In: SPIE Medical Imaging: Computer-Aided Diagnosis, Orlando, FL, USA, vol. 7963, p. 79633I (February 2011)
Haralick, R.M., Shanmugam, K., Dinstein, I.: Textural features for image classification. IEEE Transactions on Systems, Man and Cybernetics SMC-3(6), 610–621 (1973)
Hosny, K.M.: Image representation using accurate orthogonal gegenbauer moments. Pattern Recognition Letters 32(6), 795–804 (2011)
Hu, M.K.: Visual pattern recognition by moment invariants. IEEE Transactions on Information Theory 8(2), 179–187 (1962)
Jain, A., Zongker, D.: Feature selection: evaluation, application, and small sample performance. IEEE Transactions on Pattern Analysis and Machine Intelligence 19(2), 153–158 (1997)
Jiang, K., Liao, Q.M., Dai, S.Y.: A novel white blood cell segmentation scheme using scale-space filtering and watershed clustering. In: IEEE International Conference on Machine Learning and Cybernetics, Xi’an, China, pp. 2820–2825 (November 2003)
Kaya, G.T., Kaya, H., Ersoy, O.K.: Feature selection by high dimensional model representation and its application to remote sensing. In: IEEE International Geoscience and Remote Sensing Symposium (IGARSS), pp. 4938–4941 (2012)
Kok-Swee, S., Faizy Salleh, A., Chee-way, C., Rosli, B., Hock-Ann, G.: Translation and scale invariants of Hahn moments. International Journal of Image and Graphics 09(02), 271–285 (2009)
Kumar, B.R., Joseph, D.K., Sreenivas, T.V.: Teager energy based blood cell segmentation. In: 14th International Conference on Digital Signal Processing, Santorini, Greece, pp. 619–622 (July 2002)
Landis, J.R., Koch, G.G.: The measurement of observer agreement for categorical data. Biometrics 33(1)
Lezoray, O., Elmoataz, A., Cardot, H., Gougeon, G., Lecluse, M., Elie, H., Revenu, M.: Segmentation of cytological images using color and mathematical morphology. Acta Stereologica 18(1), 1–14 (1999)
Li, S., Lee, M.C., Pun, C.M.: Complex Zernike moments features for shape-based image retrieval. IEEE Transactions on Systems, Man and Cybernetics, Part A: Systems and Humans 39(1), 227–237 (2009)
Mukherjee, D.P., Ray, N., Acton, S.T.: Level set analysis for leukocyte detection and tracking. IEEE Transactions on Image Processing 13(4), 562–572 (2004)
Mukundan, R., Ong, S.H., Lee, P.A.: Image analysis by Tchebichef moments. IEEE Transactions on Image Processing 10(9), 1357–1364 (2001)
Ongun, G., Halici, U., Leblebicioglu, K., Atalay, V., Beksac, M., Beksac, S.: Feature extraction and classification of blood cells for an automated differential blood count system. In: International Joint Conference on Neural Networks, Washington, DC, USA, pp. 2461–2466 (July 2001)
Ping, Z., Wu, R., Sheng, Y.: Image description with Chebyshev-Fourier moments. Journal of the Optical Society of America A 19(9), 1748–1754 (2002)
Rahman, S.: Extended polynomial dimensional decomposition for arbitrary probability distributions. Journal of Engineering Mechanics 135(12), 1439–1451 (2009)
Ramoser, H., Laurain, V., Bischof, H., Ecker, R.: Leukocyte segmentation and classification in blood-smear images. In: 27th IEEE Annual Conference Engineering in Medicine and Biology, Shanghai, China, September 1-4, pp. 3371–3374 (2005)
Ren, H., Ping, Z., Bo, W., Wu, W., Sheng, Y.: Multidistortion-invariant image recognition with radial harmonic fourier moments. Journal of the Optical Society of America A 20(4), 631–637 (2003)
Rodenacker, K., Bengtsson, E.: A feature set for cytometry on digitized microscopic images. Analytical Cellular Pathology 25(1), 1–36 (2001)
Rowan, R., England, J.M.: Automated examination of the peripheral blood smear. In: Automation and Quality Assurance in Hematology, ch. 5, pp. 129–177. Blackwell Scientific Oxford (1986)
Ruzicka, K., Veitl, M., Thalhammer-Scherrer, R., Schwarzinger, I.: New hematology analyzer Sysmex XE-2100: performance evaluation of a novel white blood cell dierential technology. Archives of Pathology and Laboratory Medicine 125(3), 391–396 (2001)
Sheng, Y., Shen, L.: Orthogonal fourier-mellin moments for invariant pattern recognition. Journal of the Optical Society of America A 11(6), 1748–1757 (1994)
Shitong, W., Min, W.: A new detection algorithm (NDA) based on fuzzy cellular neural networks for white blood cell detection. IEEE Transactions on Information Technology in Biomedicine 10(1), 5–10 (2006)
Sinha, N., Ramakrishnan, A.G.: Automation of differential blood count. In: IEEE International Conference on Convergent Technologies for Asia-Pacific Region, Bangalore, India, pp. 547–551 (October 2003)
Sobol, I.M.: Global sensitivity indices for nonlinear mathematical models and their Monte Carlo estimates. Mathematics and Computers in Simulation 55(1-3), 271–280 (2001)
Sobrevilla, P., Montseny, E., Keller, J.: White blood cell detection in bone marrow images. In: 18th International Conference of the North American Fuzzy Information Processing Societ (NAFIPS), pp. 403–407 (1999)
Tamura, H., Mori, S., Yamawaki, T.: Textural features corresponding to visual perception. IEEE Transactions on Systems, Man and Cybernetics 8(6), 460–473 (1978)
Tang, X.: Texture information in runlength matrices. IEEE Transactions on Image Processing 7(11), 1602–1609 (1998)
Theera-Umpon, N., Dhompongsa, S.: Morphological granulometric features of nucleus in automatic bone marrow white blood cell classification. IEEE Transactions on Information Technology in Biomedicine 11(3), 353–359 (2007)
Verso, M.L.: The evolution of blood-counting techniques. Journal of Medical History 8(2), 149–158 (1964)
Xia, T., Zhu, H., Shu, H., Haigron, P., Luo, L.: Image description with generalized pseudo-zernike moments. Journal of the Optical Society of America A 24(1), 50–59 (2007)
Xiao-min, Y., Li-min, L., Yu, W.: Automatic classification system for leukocytes in human blood. Journal of Computer Science and Technology 17(2), 130–136 (1994)
Yap, P.T., Paramesran, R., Ong, S.H.: Image analysis by Krawtchouk moments. IEEE Transactions on Image Processing 12(11), 1367–1377 (2003)
Kang, B., Ma, Z., Ma, J.: Translation and scale invariant of Legendre moments for images retrieval. Journal of Information & Computational Science 8(11), 2221–2229 (2011)
Ziehn, T., Tomlin, A.S.: GUI-HDMR - a software tool for global sensitivity analysis of complex models. Environmental Modelling & Software 24(7), 775–785 (2009)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer International Publishing Switzerland
About this paper
Cite this paper
Habibzadeh, M., Krzyżak, A., Fevens, T. (2014). Comparative Study of Feature Selection for White Blood Cell Differential Counts in Low Resolution Images. In: El Gayar, N., Schwenker, F., Suen, C. (eds) Artificial Neural Networks in Pattern Recognition. ANNPR 2014. Lecture Notes in Computer Science(), vol 8774. Springer, Cham. https://doi.org/10.1007/978-3-319-11656-3_20
Download citation
DOI: https://doi.org/10.1007/978-3-319-11656-3_20
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-11655-6
Online ISBN: 978-3-319-11656-3
eBook Packages: Computer ScienceComputer Science (R0)