Summary
Infections with Streptococcus pneumoniae, Group A Streptococcus (GAS), and Group B Streptococcus (GBS) are among the most frequent bacterial infections in humans and major causes of diseases. Immune responses against streptococci include the induction of type I interferons (IFNs), which are immunomodulatory cytokines with well-established antiviral functions, but ambiguous roles in bacterial infections. Studies of streptococcal infections further highlight the importance of type I IFN signaling in host defense against bacterial pathogens. We discuss the complexity of type I IFN induction by streptococcal ligands and their engagement in stimulating innate immune receptors. Furthermore, we elaborate on the broad physiological role and different impacts of type I IFN signaling on the outcome of streptococcal infections and influenza virus coinfections.
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Type I Interferons in Bacterial Infections
The function of type I interferons (IFNs) in viral infections is well established and can be almost uniformly described as protective. In contrast, their role in the context of bacterial infections is much less clear, as both beneficial and detrimental effects of type I IFN signaling have been reported in animal models [1, 2]. Examples where type I IFNs confer a protective role can be found in cases of infection with Salmonella typhimurium, Group B Streptococcus (GBS), Legionella pneumophila, and Streptococcus pneumoniae [3–6]. The molecular mechanisms underlying type I IFN function in the context of these infections range from the induction of cytokines and iNOS, to the enhanced differentiation of inflammatory macrophages, and may also include more complex processes, which orchestrate innate and adaptive immune responses. On the other hand, in cases of infection with Listeria monocytogenes and Francisella tularensis, type I IFNs exert unfavorable functions [7–11]. Various mechanisms can explain these harmful effects, such as type I IFN-mediated apoptosis of infected lymphocytes and macrophages, IFN-dependent reduction of IL-17 production by γδT cells, or diminished neutrophil activity. In summary, it is currently not possible to identify the denominator of either beneficial or detrimental effects of type I IFNs. Given the profound effects of these immunomodulatory cytokines on the outcome of bacterial infections, elucidating their incompletely understood induction by bacteria is of immense importance [12]. In the following, we will review the current understanding of the role of type I IFNs, as well as of the mechanisms of their induction in host defense against Streptococcus pyogenes (Group A Streptococcus, GAS), Streptococcus agalactiae (GBS), and S. pneumoniae. These streptococcal species are major human pathogens which, despite a long history of intense research, continue to pose a serious health threat worldwide. In this context, new therapies employing modulation of cytokine activities are attractive yet underexplored strategies.
Group A Streptococcus (S. pyogenes)
Pathogenicity
GAS, also called S. pyogenes, is a leading Gram-positive human pathogen. GAS causes a broad range of mostly self-limiting diseases including pharyngitis (strep throat), scarlet fever or impetigo [13, 14]. It may however establish invasive and life-threatening infections, such as necrotizing fasciitis and toxic shock, which result in mortality rates of more than 30 % [13]. GAS accounts for over 700 million mild and more than 650,000 severe invasive infections worldwide annually [15]. GAS and S. pneumoniae are the most frequently found coinfecting bacteria in specimens of the 1918 influenza pandemic and in patients of the recent H1N1 influenza outbreak [16, 17]. Analysis of patient samples and animal studies reveal that the exceptionally wide range of GAS-caused diseases along with the transition from contained to invasive infections is determined by the virulence factor armament of a particular bacterial strain and by the genetic inventory of the host immune system [13, 18–20]. The underlying host–pathogen interactions are not well understood. Virulence factors include T cell-activating superantigens, surface-localized proteins such as the serotype-determining M protein interfering with the complement system and phagocytosis, the internalization-inhibiting hyaluronic acid capsule, secreted proteases with cytokine/chemokine-inactivating properties, secreted DNases that help bacterial dissemination, and the cytolysins SLO and SLS [19, 21–23]. Horizontal bacteriophage-mediated genetic transfer and the counteracting CRISPR system contribute to the virulence diversity observed between GAS strains [24–26]. On the host side, animal studies demonstrated that innate immune cells, most notably macrophages, dendritic cells, and neutrophils, play an essential role in successful defense during subcutaneous infection, a model of invasive GAS infection [27–29]. In models of upper respiratory tract infections, mucosal Th17 cells have been found to exert protective effects although the specific effector function of these cells in GAS infections remain to be identified [30, 31]. IL-17-mediated activation of antibacterial innate immune mechanisms could be involved in the Th17-dependent defense. Interestingly, in mice the variability of individual innate immune responses contributes to differences in susceptibility to GAS infections more than the variability in T cell-mediated responses [32].
Despite the fact that GAS is a human-specific pathogen, and mice are resistant against GAS outside of laboratory conditions [33, 34], animal infection models are invaluable for understanding GAS diseases and improvements of current therapies. Consistently, much of what is known about host–pathogen interactions in GAS infections has been established from studies using gene-targeted mice. In future studies, the use of humanized mice [35] will be helpful for functional and mechanistic assessment of GAS virulence factors that target human but not murine defense systems.
Type I IFN Induction
GAS activates type I IFN production by both human and mouse innate immune cells [4, 36–38]. In addition, GAS infection of primary human macrophages triggers an IFN signaling signature resulting, among others, in the activation of the transcription factor STAT1 [37]. This signaling signature is prevented by antibodies neutralizing IFN-α and IFN-β; however, the precise nature of type I IFNs induced by GAS in human cells remains unclear. In mice, primary bone marrow-derived macrophages (BMDMs) and conventional dendritic cells (cDCs), but not plasmacytoid dendritic cells (pDCs), were shown to produce IFN-β upon GAS infection [36, 38]. In fact, GAS-derived DNA triggers IFN-β in macrophages, whereas GAS RNA stimulates IFN-β in cDCs [38] (Table 1). Generally, IFN-β is the primary type I IFN produced upon infection—it up-regulates the transcription factor IRF7 which then triggers IFN-α genes [39].
GAS-induced IFN-β activates the transcription factor STAT1 and STAT1 target genes in an IFNAR (type I IFN receptor)-dependent manner, confirming and clarifying a functional involvement of IFN signaling downstream of type I IFN production [36]. In contrast, the mechanism of type I IFN induction by GAS is incompletely understood (Fig. 1). Most importantly, the pattern recognition receptors (PRRs) triggering the IFN-β gene are not known [38]. In general, the identity of PRRs able to sense GAS remains one of the most challenging questions. The sole involvement of TLR2, the PRR recognizing cell wall components of Gram-positive bacteria, as well as of TLR1, TLR4, and TLR6 have been excluded [36, 38, 40]. Similarly, nucleic acid-recognizing TLR3, TLR7, and TLR9 are dispensable for production of inflammatory cytokines and type I IFNs by GAS-infected innate immune cells [36, 38, 40]. Further, type I IFNs are induced independently of the cytosolic PRRs NOD1 and NOD2 [38], which were shown to be required for IFN-β stimulation in several viral and bacterial infection models [41, 42]. Attempts at identifying the proximal GAS sensor have been performed employing cells derived from mice deficient in multiple TLRs. TLR2/TLR4 and TLR2/TLR6 double-deficient BMDMs and cDCs were not impaired in GAS recognition. It remains to be elucidated whether and how the newly characterized TLR13 is involved in GAS recognition and type I IFN induction. TLR13 is activated by a conserved sequence within the 23S rRNA of both Gram-negative and -positive bacteria [43, 44]. TLR13 stimulation causes production of inflammatory cytokines including TNF, IL-6, and IL-1β, but its role in type I IFN induction has not been clarified yet. Similarly, the role of TLR13 in host defense against bacterial pathogens remains to be investigated despite the ability of this PRR to recognize RNA of important pathogens such as S. aureus or GAS [43, 45]. The fact that TLR13 is expressed in mice but not humans raises the question whether humans possess an alternative route of bacterial RNA recognition. Yet another receptor that could potentially play a role in type I IFN induction by GAS is the recently characterized cyclic GMP-AMP synthase (cGAS) which acts as a cytosolic DNA sensor [46, 47]. cGAS is a danger recognition receptor which upon binding to DNA synthesizes the second messenger cyclic GMP-AMP (cGAMP). cGAMP binds and activates the ER protein STING to trigger IRF3 and IFN-β gene expression [48, 49]. While cGAS is involved in cellular defense against viruses [50–53], a role of cGAS in bacterial infections and/or in induction of type I IFNs by bacteria has not been demonstrated yet.
Signaling events downstream of the type I IFN-inducing GAS-specific PRRs are better understood (Fig. 1 and Table 1). Activation of Ifnb gene expression by GAS-derived DNA in macrophages is dependent on the TBK1 kinase and the transcription factor IRF3 [38]. In contrast, the IFN-β-inducing pathway triggered by GAS RNA in cDCs requires the adaptor MyD88 as well as the transcription factor IRF5, but not IRF3 [38]. Uptake of GAS is needed for triggering IFN-β production suggesting that phagolysosomal processing of internalized GAS liberates the bacterial IFN-β inducers. Whether both BMDMs and cDCs are involved in IFN-β production in vivo and whether these cell types play a redundant or distinct roles have yet to be examined.
Type I IFN Functions
Mice lacking the type I IFN receptor IFNAR1 are more susceptible to subcutaneous GAS infection [38], a standard model of severe invasive cellulitis [20]. The mortality rate of GAS-infected IFNAR1-deficient mice is 70 % whereas it is only 25 % in WT mice. IFNAR1 knockouts were shown to exhibit increased recruitment of neutrophils to the site of infection but the molecular and cellular basis of the beneficial effects of type I IFNs in GAS infection remain to be elucidated. The high neutrophil number observed in mice lacking type I IFN signaling is consistent with previous observations demonstrating inhibitory effects of type I IFNs on macrophage production of the chemokines CXCL1, CXCL2, and CCL2 during S. pneumoniae infections [54, 55]. These chemokines play a key role in attracting neutrophils to the site of infection. It is at present unclear how the increased neutrophil recruitment in GAS-infected IFNAR knockout mice could evoke more detrimental disease. One can speculate that an exaggerated inflammatory response elicited by recruited neutrophils causes severe tissue damage, thereby allowing better dissemination of the pathogen. Such scenario is conceivable as GAS expresses several DNases that help liberate it from neutrophil extracellular traps (NETs) [56, 57]. Consistently, the DNase Sda1 is a potent virulence factor which promotes GAS to acquire an invasive infection phenotype [58]. GAS exhibits a profound propensity to induce NETs, structures that contain large amounts of inflammation-promoting material such as neutrophil DNA, histones, and other chromatin-associated proteins [59, 60]. Interestingly, TLR9, a PRR able to sense self DNA [61], might be involved in sensing GAS-induced NETs as it is beneficial in an intraperitoneal model of GAS infection [62]. This indirect role of TLR9 in GAS infections is supported by the lack of effect of TLR9 knockout on direct GAS recognition by BMDMs and cDCs [4, 38, 40]. Thus, the enhanced neutrophil recruitment in IFNAR1-deficient mice might result in more intense, hence lethal inflammation. Effects of type I IFNs on other immune reactions such as recruitment of macrophages by GAS-induced TNF [63], or IL-1β production by the GAS-activated NLRP3 inflammasome [64], should be addressed in future studies to reveal the precise role of type I IFN signaling.
Group B Streptococcus (S. agalactiae)
Pathogenicity
GBS, also called S. agalactiae, is a Gram-positive human pathogen and leading infectious agent in neonatal sepsis worldwide [65]. Neonatal sepsis causes over two million deaths annually, with decreasing incidence largely due to improved prophylactic measures [66]. In early onset neonatal disease (within 6 days after birth), GBS is transmitted vertically from mothers vaginally colonized by the pathogen. In late onset disease (7–89 days after birth), GBS infection is usually a consequence of horizontal transfer in communities. GBS is also a significant cause of maternal morbidity (bacteremia, endometritis) [67]. GBS virulence factors include the polysaccharide capsule, membrane damaging exotoxins, and adherence molecules which enable evasion of the immune system and colonization of the host [68]. Innate immune system-derived TNF, IL-1β, and nitric oxide are key defense factors in host protection [69–71]. The vulnerability of neonates to GBS results in part from underdeveloped adaptive immunity but more importantly from deficiencies in innate immunity, including limited capacity of neutrophil production and increased risk of bone marrow exhaustion [67, 72]. The neonate immune insufficiency allows colonization and infection by GBS resulting mostly in meningitis or pneumonia. Prophylactic vaccination and immunomodulation appear the most promising approaches to eradicate GBS disease [67, 73].
Type I IFN Induction and Function
Type I IFN signaling has a protective function in GBS infections: mice deficient in either type I IFN receptor or IFN-β exhibit increased mortality in a neonatal infection model, both after intravenous or intraperitoneal GBS administration [74]. This lethal infection outcome is caused by uncontrolled bacteremia, suggesting that type I IFN signaling is required for launching a complete immune and antibacterial response. Both macrophages and cDCs, but not pDCs, were identified as the source of type I IFNs [4, 74, 75] (Table 1). A direct comparison of type I IFN amounts indicate that cDCs are the major producers in vitro [4] but the principle type I IFN-producing cell in vivo has yet to be confirmed. Type I IFN production is dependent on uptake and phagolysosomal processing of GBS [4, 75] (Fig. 2). In macrophages, GBS DNA was identified as type I IFN inducer that acts along the TBK1 and IRF3 axis [75] (Fig. 2 and Table 1). GBS DNA was proposed to escape phagosomes into the cytosol where it is detected by an unknown cytosolic DNA receptor, which is different from the double-stranded DNA sensor DAI [75, 76]. The inducer of type I IFNs in cDCs was shown to be GBS RNA, which was sensed in a MyD88-dependent manner in phagosomes of infected cells [4] (Fig. 2 and Table 1). The endosomal TLR7 was found to be involved in sensing of GBS RNA. Interestingly, GBS RNA was reported to induce TNF in macrophages independently of TLR3, TLR7, and TLR8, but it required MyD88 [77]. This RNA recognition occurs in endosomal compartments as it is dependent on Unc93b, a chaperon fundamentally involved in trafficking of endosomal TLRs. Together, these studies indicate that recognition of GBS is cell type-specific, and that GBS RNA induces type I IFNs in cDCs but not in macrophages. The molecular basis of the different outcome of GBS RNA sensing in macrophages and cDCs remains to be deciphered. As is the case with GAS, the analysis of the recently identified sensors TLR13 and cGAS might be helpful in resolving the open questions.
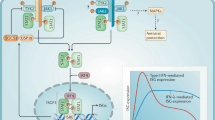
Type IFN signaling and induction by GBS. Induction of type I IFNs by GBS requires uptake and phagolysosomal processing of the pathogen. In macrophages, GBS-derived DNA triggers a cytosolic sensor which signals via TBK1 and IRF3 to induce IFN-β gene expression. In cDCs, GAS-derived RNA triggers the in Unc93b-dependent way the endosomal TLR7 which signals via MyD88 toward the IFN-β gene
Streptococcus pneumoniae
Pathogenesis
S. pneumoniae (pneumococcus) is a Gram-positive human pathogen regarded as the most frequent cause of community-acquired pneumonia [78, 79]. Pneumonia is the leading lethal infectious disease in developed countries [78, 79]. S. pneumoniae is one of the most prominent examples of a human-specific commensal microbe that frequently turns into an infectious agent. S. pneumoniae asymptomatically colonizes the nasopharynx in up to 60 % of all preschool children. Yet, S. pneumoniae represents the prime bacterial killer among children below the age of 5 with 1.2 million deaths annually worldwide. S. pneumoniae also poses a serious health risk to elderly people as a consequence of age-related immunosenescence. Of particular importance is a secondary S. pneumoniae infection of influenza patients, S. pneumoniae is one of the most frequent coinfecting pathogens in cases of influenza outbreaks [16, 17]. Both the genetic makeup of the pathogen and the condition of the host immune system play decisive roles in the transition from a commensal microbe into invasive pathogen. However, the exact parameters regulating this shift are not well understood. S. pneumoniae occurs in more than 90 serotypes which differ in their virulence. The serotypes are characterized by their polysaccharide capsule, which plays an important role in evasion of the immune system by inhibiting phagocytosis and complement binding [80]. An armament of other virulence factors including pneumolysin, hyaluronidase, neuraminidase, the serine protease PrtA, cholin-binding proteins, etc. contribute to various extents to pneumococcal diseases [81, 82]. The immune response against S. pneumoniae is initiated by its interactions with innate immune receptors. TLR2 is triggered by S. pneumoniae cell wall components (e.g., LTA), TLR4 can be activated by pneumolysin and TLR9 recognizes pneumococcal DNA [80, 83–86]. Furthermore, the cytosolic receptors NLRP3, NOD2, and AIM2 contribute to S. pneumoniae-induced inflammatory cytokine induction [80, 87–90].
Type I IFN Induction
S. pneumoniae induces type I IFNs in nasal-associated lymphoid and epithelial tissues, as well as in human and mouse alveolar macrophages and mouse BMDMs [6, 55, 91, 92]. The type I IFN inducer is S. pneumoniae DNA, which is recognized upon internalization of the pathogen and/or pneumolysin-dependent cytosolic delivery [6] (Table 1). The double-stranded DNA sensor DAI participates in the detection of S. pneumoniae DNA as DAI-deficient cells produce less IFN-β than control cells [6] (Fig. 3). Similar to GAS and GBS, the signaling pathway downstream of the proximal sensor includes TBK1, STING and IRF3, and is possibly indirectly dependent on NOD2 [6, 55] (Fig. 3 and Table 1). Signal transduction toward the IFN-β gene proceeds in the absence of TLR4, MyD88, NOD2, and TRIF. Thus, the IFN-β-inducing properties of S. pneumoniae-derived DNA resemble those of GAS and GBS. It remains to be investigated whether S. pneumoniae RNA also possesses immunostimulatory capabilities as described for GAS and GBS.
Type I IFN Function
Intravenous infection of type I IFN signaling-deficient mice with S. pneumoniae results in increased lethality [74]. Further evidence for a beneficial role of type I IFNs was provided by a study using a more natural route of infection, i.e., intranasal [6]. This particular study reported an impaired clearance of the pathogen from the site of infection, i.e., from the upper respiratory tract, in mice lacking IFNAR1, despite more potent recruitment of monocytes and dendritic cells. The exact mechanism of how type I IFNs elicit protective effects in pneumococcal infections remains to be characterized.
A distinct mode of pneumococcal infection is represented by coinfections with the influenza virus. These coinfections exhibit high morbidity and are life threatening in elderly patients. In animal models of coinfections, mice are first exposed to the influenza virus and a few days later S. pneumoniae is delivered intranasally. Both, S. pneumoniae and influenza virus are able to induce type I IFNs. Coinfections lead to synergistic induction of type I IFNs and, remarkably, this high level of type I IFN signaling is detrimental to the host [54, 55, 93]. The mechanisms of the harmful effects of type I IFNs on post-influenza bacterial infection include decreased production of the chemokines CCL2, CXCL1, and CXCL2, which act as chemoattractants for monocytes and neutrophils. As a result, less monocytes and neutrophils are recruited to infected tissues, although the precise nature of the most affected leukocytes is a matter of debate [54, 55]. Further studies are needed to clarify the molecular principles of coinfections. Such studies should particularly address the inability to tolerate tissue damage, which has recently been reported to play a critical role in influenza and L. pneumophila coinfections [94].
Type I Interferons in Streptococcal Infections: Unifying Themes and Divergences
Although they share several common features, GAS, GBS, and S. pneumoniae cause diverse diseases in humans. They are Gram-positive encapsulated pathogens exhibiting a largely extracellular life cycle. Their key virulence factors are cytolysins, which possess cytotoxic properties and promote intracellular survival and/or phagolysosomal damage. These pathogens’ ability to survive and grow within infected cells is very limited, although it has been reported that GAS is capable of acquiring a significant intracellular life span [13, 95]. Nonetheless, most internalized GAS are efficiently killed by the host phagolysosomal lytic and oxidative mechanisms. GAS that has escaped from the hostile phagosomal environment is rapidly recognized in the cytosol by the autophagy machinery and eradicated [96, 97]. The highly successful destruction of streptococci in the phagosomes results in the release of, among others, bacterial nucleic acids, which can act as type I IFN inducers. Consequently, endosomal recognition of GAS and GBS RNA induces type I IFNs [4, 38]. In this context, the role of S. pneumoniae RNA has yet to be investigated. In contrast, all three streptococcal species have been reported to induce type I IFNs by their DNA, which is sensed by cytosolic DNA receptors [6, 38, 75]. Cytolysins are likely to be involved in the passage of DNA through the phagosomal membrane, but the precise mechanisms of streptococcal DNA delivery into the host cell cytosol remain unclear. The issue of type I IFN-inducing receptors also requires further investigation. Whereas TLR7 was identified as the RNA-sensing type I IFN inducer in response to GBS but not GAS [4, 38], the DNA sensor DAI was found to induce type I IFNs in response to S. pneumoniae but not GBS [6, 75]. Future studies, now also include newly identified receptors, will show whether there are common type I IFN-inducing pathways in streptococcal infections.
Type I IFNs exhibit protective functions in infections against all three streptococcal species discussed here, yet the precise nature of these beneficial functions are not well explained. As the three streptococcal species cause different diseases and display in part different tissue tropism, the mode of action of type I IFNs will most likely involve multiple possibly non-overlapping mechanisms. Elucidation of type I IFN functions is essential for our better understanding of the surprisingly detrimental effects of these cytokines during viral coinfections [54, 55, 93]. Further, it has yet to be investigated whether the negative impact of type I IFNs during coinfections is restricted to respiratory pathogens.
Outlook
Despite significant advances in our understating of type I IFNs in bacterial infections, the key questions remain unresolved for most bacterial pathogens. These questions include the identity of type I IFN-inducing sensors and the specific effector functions of type I IFNs. Analyses of a broader range of innate immune receptors, ideally by employing unbiased approaches such as mass spectroscopy or genetic screens, will give us a more comprehensive picture of type I IFN induction. To elucidate the effector functions of type I IFNs, better infection models are needed. These will have to include animals allowing cell type-specific deletion of IFNAR1 [98], analysis of animals lacking different type I IFNs (particularly IFN-β), and in vivo and intravital imaging techniques. A so far unexplored aspect in streptococcal infections is the timing of type I IFN signaling. In the view of recent findings describing an unexpected harmful function of type I IFNs during persistent viral infections [99, 100], time-resolved analysis of type I IFN signaling in streptococcal infections and viral coinfections will need to be conducted in future studies. Another major challenge is the evaluation of the relevance of animal studies for the understanding of streptococcal diseases in humans. Clearly, the use of gene-targeted mice will remain fundamental for mechanistic and proof-of-principle studies. However, the increasingly better understood differences between the human and mouse immune systems, including their partially different repertoires of innate immune receptors, should be carefully considered when using animal models for human-specific pathogens.
Modulation of immune responses is recognized as a highly promising approach in the treatment of severe infectious diseases, and it may be the sole strategy for the treatment of acute life-threatening conditions such as streptococcal toxic shock syndrome. Type I IFNs are major immune modulators, possessing both immunostimulatory and immunosuppressive properties [101, 102]; as such, the elucidation of their mechanism of action in streptococcal infections could eventually establish type I IFN signaling as a target for novel therapies.
References
Trinchieri G (2010) Type I interferon: friend or foe? J Exp Med 207:2053–2063
Decker T, Muller M, Stockinger S (2005) The yin and yang of type I interferon activity in bacterial infection. Nat Rev Immunol 5:675–687
Freudenberg MA, Merlin T, Kalis C, Chvatchko Y, Stubig H et al (2002) Cutting edge: a murine, IL-12-independent pathway of IFN-gamma induction by gram-negative bacteria based on STAT4 activation by Type I IFN and IL-18 signaling. J Immunol 169:1665–1668
Mancuso G, Gambuzza M, Midiri A, Biondo C, Papasergi S et al (2009) Bacterial recognition by TLR7 in the lysosomes of conventional dendritic cells. Nat Immunol 10:587–594
Plumlee CR, Lee C, Beg AA, Decker T, Shuman HA et al (2009) Interferons direct an effective innate response to Legionella pneumophila infection. J Biol Chem 284:30058–30066
Parker D, Martin FJ, Soong G, Harfenist BS, Aguilar JL et al (2011) Streptococcus pneumoniae DNA initiates type I interferon signaling in the respiratory tract. MBio 2:e00016-00011
O’Connell RM, Saha SK, Vaidya SA, Bruhn KW, Miranda GA et al (2004) Type I interferon production enhances susceptibility to Listeria monocytogenes infection. J Exp Med 200:437–445
Auerbuch V, Brockstedt DG, Meyer-Morse N, O’Riordan M, Portnoy DA (2004) Mice lacking the type I interferon receptor are resistant to Listeria monocytogenes. J Exp Med 200:527–533
Carrero JA, Calderon B, Unanue ER (2004) Type I interferon sensitizes lymphocytes to apoptosis and reduces resistance to listeria infection. J Exp Med 200:535–540
Stockinger S, Kastner R, Kernbauer E, Pilz A, Westermayer S et al (2009) Characterization of the interferon-producing cell in mice infected with Listeria monocytogenes. PLoS Pathog 5:e1000355
Henry T, Kirimanjeswara GS, Ruby T, Jones JW, Peng K et al (2010) Type I IFN signaling constrains IL-17A/F secretion by gammadelta T cells during bacterial infections. J Immunol 184:3755–3767
Monroe KM, McWhirter SM, Vance RE (2010) Induction of type I interferons by bacteria. Cell Microbiol 12:881–890
Johansson L, Thulin P, Low DE, Norrby-Teglund A (2010) Getting under the skin: the immunopathogenesis of Streptococcus pyogenes deep tissue infections. Clin Infect Dis 51:58–65
Wessels MR (2011) Clinical practice. Streptococcal pharyngitis. N Engl J Med 364:648–655
Carapetis JR, Steer AC, Mulholland EK, Weber M (2005) The global burden of group A streptococcal diseases. Lancet Infect Dis 5:685–694
Morens DM, Fauci AS (2007) The 1918 influenza pandemic: insights for the 21st century. J Infect Dis 195:1018–1028
Zakikhany K, Degail MA, Lamagni T, Waight P, Guy R et al (2011) Increase in invasive Streptococcus pyogenes and Streptococcus pneumoniae infections in England, December 2010 to January 2011. Euro Surveill 16
Kotb M, Norrby-Teglund A, McGeer A, El-Sherbini H, Dorak MT et al (2002) An immunogenetic and molecular basis for differences in outcomes of invasive group A streptococcal infections. Nat Med 8:1398–1404
Olsen RJ, Shelburne SA, Musser JM (2009) Molecular mechanisms underlying group A streptococcal pathogenesis. Cell Microbiol 11:1–12
Medina E (2010) Murine model of cutaneous infection with Streptococcus pyogenes. Methods Mol Biol 602:395–403
Bisno AL, Brito MO, Collins CM (2003) Molecular basis of group A streptococcal virulence. Lancet Infect Dis 3:191–200
Lynskey NN, Lawrenson RA, Sriskandan S (2011) New understandings in Streptococcus pyogenes. Curr Opin Infect Dis 24:196–202
Molloy EM, Cotter PD, Hill C, Mitchell DA, Ross RP (2011) Streptolysin S-like virulence factors: the continuing sagA. Nat Rev Microbiol 9:670–681
Sitkiewicz I, Nagiec MJ, Sumby P, Butler SD, Cywes-Bentley C et al (2006) Emergence of a bacterial clone with enhanced virulence by acquisition of a phage encoding a secreted phospholipase A2. Proc Natl Acad Sci U S A 103:16009–16014
Deltcheva E, Chylinski K, Sharma CM, Gonzales K, Chao Y et al (2011) CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471:602–607
Nozawa T, Furukawa N, Aikawa C, Watanabe T, Haobam B et al (2011) CRISPR inhibition of prophage acquisition in Streptococcus pyogenes. PLoS One 6:e19543
Goldmann O, Rohde M, Chhatwal GS, Medina E (2004) Role of macrophages in host resistance to group A streptococci. Infect Immun 72:2956–2963
Loof TG, Rohde M, Chhatwal GS, Jung S, Medina E (2007) The contribution of dendritic cells to host defenses against Streptococcus pyogenes. J Infect Dis 196:1794–1803
Navarini AA, Lang KS, Verschoor A, Recher M, Zinkernagel AS et al (2009) Innate immune-induced depletion of bone marrow neutrophils aggravates systemic bacterial infections. Proc Natl Acad Sci U S A 106:7107–7112
Wang B, Dileepan T, Briscoe S, Hyland KA, Kang J et al (2010) Induction of TGF-beta1 and TGF-beta1-dependent predominant Th17 differentiation by group A streptococcal infection. Proc Natl Acad Sci U S A 107:5937–5942
Dileepan T, Linehan JL, Moon JJ, Pepper M, Jenkins MK et al (2011) Robust antigen specific Th17 T cell response to group A Streptococcus is dependent on IL-6 and intranasal route of infection. PLoS Pathog 7:e1002252
Goldmann O, Lengeling A, Bose J, Bloecker H, Geffers R et al (2005) The role of the MHC on resistance to group a streptococci in mice. J Immunol 175:3862–3872
Fox GF, Anderson LC, Loew FM, Quimby FW (2002) Laboratory animal medicine. Academic, Elsevier
Tart AH, Walker MJ, Musser JM (2007) New understanding of the group A Streptococcus pathogenesis cycle. Trends Microbiol 15:318–325
Brehm MA, Jouvet N, Greiner DL, Shultz LD (2013) Humanized mice for the study of infectious diseases. Curr Opin Immunol 25:428–435
Gratz N, Siller M, Schaljo B, Pirzada ZA, Gattermeier I et al (2008) Group A Streptococcus activates type I interferon production and MyD88-dependent signaling without involvement of TLR2, TLR4, and TLR9. J Biol Chem 283:19879–19887
Miettinen M, Lehtonen A, Julkunen I, Matikainen S (2000) Lactobacilli and Streptococci activate NF-kappa B and STAT signaling pathways in human macrophages. J Immunol 164:3733–3740
Gratz N, Hartweger H, Matt U, Kratochvill F, Janos M et al (2011) Type I interferon production induced by Streptococcus pyogenes-derived nucleic acids is required for host protection. PLoS Pathog 7:e1001345
Marie I, Durbin JE, Levy DE (1998) Differential viral induction of distinct interferon-alpha genes by positive feedback through interferon regulatory factor-7. EMBO J 17:6660–6669
Loof TG, Goldmann O, Medina E (2008) Immune recognition of Streptococcus pyogenes by dendritic cells. Infect Immun 76:2785–2792
Watanabe T, Asano N, Murray PJ, Ozato K, Tailor P et al (2008) Muramyl dipeptide activation of nucleotide-binding oligomerization domain 2 protects mice from experimental colitis. J Clin Invest 118:545–559
Sabbah A, Chang TH, Harnack R, Frohlich V, Tominaga K et al (2009) Activation of innate immune antiviral responses by Nod2. Nat Immunol 10:1073–1080
Oldenburg M, Kruger A, Ferstl R, Kaufmann A, Nees G et al (2012) TLR13 recognizes bacterial 23S rRNA devoid of erythromycin resistance-forming modification. Science 337:1111–1115
Li XD, Chen ZJ (2012) Sequence specific detection of bacterial 23S ribosomal RNA by TLR13. Elife 1:e00102
Hidmark A, von Saint Paul A, Dalpke AH (2012) Cutting edge: TLR13 is a receptor for bacterial RNA. J Immunol 189:2717–2721
Sun L, Wu J, Du F, Chen X, Chen ZJ (2012) Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339(6121):786–791
Wu J, Sun L, Chen X, Du F, Shi H et al (2012) Cyclic GMP-AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA. Science 339(6121):826–830
Ishikawa H, Barber GN (2008) STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455:674–678
Ishikawa H, Ma Z, Barber GN (2009) STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461:788–792
Li XD, Wu J, Gao D, Wang H, Sun L et al (2013) Pivotal roles of cGAS-cGAMP signaling in antiviral defense and immune adjuvant effects. Science 341:1390–1394
Gao D, Wu J, Wu YT, Du F, Aroh C et al (2013) Cyclic GMP-AMP synthase is an innate immune sensor of HIV and other retroviruses. Science 341:903–906
Schoggins JW, MacDuff DA, Imanaka N, Gainey MD, Shrestha B et al (2014) Pan-viral specificity of IFN-induced genes reveals new roles for cGAS in innate immunity. Nature 505:691–695
Ablasser A, Schmid-Burgk JL, Hemmerling I, Horvath GL, Schmidt T et al (2013) Cell intrinsic immunity spreads to bystander cells via the intercellular transfer of cGAMP. Nature 503:530–534
Shahangian A, Chow EK, Tian X, Kang JR, Ghaffari A et al (2009) Type I IFNs mediate development of postinfluenza bacterial pneumonia in mice. J Clin Invest 119:1910–1920
Nakamura S, Davis KM, Weiser JN (2011) Synergistic stimulation of type I interferons during influenza virus coinfection promotes Streptococcus pneumoniae colonization in mice. J Clin Invest 121:3657–3665
Buchanan JT, Simpson AJ, Aziz RK, Liu GY, Kristian SA et al (2006) DNase expression allows the pathogen group A Streptococcus to escape killing in neutrophil extracellular traps. Curr Biol 16:396–400
Chang A, Khemlani A, Kang H, Proft T (2011) Functional analysis of Streptococcus pyogenes nuclease A (SpnA), a novel group A streptococcal virulence factor. Mol Microbiol 79:1629–1642
Walker MJ, Hollands A, Sanderson-Smith ML, Cole JN, Kirk JK et al (2007) DNase Sda1 provides selection pressure for a switch to invasive group A streptococcal infection. Nat Med 13:981–985
Kolaczkowska E, Kubes P (2013) Neutrophil recruitment and function in health and inflammation. Nat Rev Immunol 13:159–175
Brinkmann V, Zychlinsky A (2007) Beneficial suicide: why neutrophils die to make NETs. Nat Rev Microbiol 5:577–582
Barrat FJ, Meeker T, Gregorio J, Chan JH, Uematsu S et al (2005) Nucleic acids of mammalian origin can act as endogenous ligands for Toll-like receptors and may promote systemic lupus erythematosus. J Exp Med 202:1131–1139
Zinkernagel AS, Hruz P, Uchiyama S, von Kockritz-Blickwede M, Schuepbach RA et al (2012) Importance of Toll-like receptor 9 in host defense against M1T1 group A Streptococcus infections. J Innate Immun 4:213–218
Mishalian I, Ordan M, Peled A, Maly A, Eichenbaum MB et al (2011) Recruited macrophages control dissemination of group A Streptococcus from infected soft tissues. J Immunol 187:6022–6031
Harder J, Franchi L, Munoz-Planillo R, Park JH, Reimer T et al (2009) Activation of the Nlrp3 inflammasome by Streptococcus pyogenes requires streptolysin O and NF-{kappa}B activation but proceeds independently of TLR signaling and P2X7 receptor. J Immunol 183:5823–5829
Dagnew AF, Cunnington MC, Dube Q, Edwards MS, French N et al (2012) Variation in reported neonatal group B streptococcal disease incidence in developing countries. Clin Infect Dis 55:91–102
Lozano R, Wang H, Foreman KJ, Rajaratnam JK, Naghavi M et al (2011) Progress towards millennium development goals 4 and 5 on maternal and child mortality: an updated systematic analysis. Lancet 378:1139–1165
Koenig JM, Keenan WJ (2009) Group B Streptococcus and early-onset sepsis in the era of maternal prophylaxis. Pediatr Clin North Am 56:689–708, Table of Contents
Henneke P, Berner R (2006) Interaction of neonatal phagocytes with group B Streptococcus: recognition and response. Infect Immun 74:3085–3095
Mancuso G, Midiri A, Beninati C, Biondo C, Galbo R et al (2004) Dual role of TLR2 and myeloid differentiation factor 88 in a mouse model of invasive group B streptococcal disease. J Immunol 172:6324–6329
Costa A, Gupta R, Signorino G, Malara A, Cardile F et al (2012) Activation of the NLRP3 inflammasome by group B streptococci. J Immunol 188:1953–1960
Deshmukh SD, Muller S, Hese K, Rauch KS, Wennekamp J et al (2012) NO is a macrophage autonomous modifier of the cytokine response to streptococcal single-stranded RNA. J Immunol 188:774–780
Teti G, Mancuso G, Tomasello F, Chiofalo MS (1992) Production of tumor necrosis factor-alpha and interleukin-6 in mice infected with group B streptococci. Circ Shock 38:138–144
Pannaraj PS, Edwards MS, Ewing KT, Lewis AL, Rench MA et al (2009) Group B streptococcal conjugate vaccines elicit functional antibodies independent of strain O-acetylation. Vaccine 27:4452–4456
Mancuso G, Midiri A, Biondo C, Beninati C, Zummo S et al (2007) Type I IFN signaling is crucial for host resistance against different species of pathogenic bacteria. J Immunol 178:3126–3133
Charrel-Dennis M, Latz E, Halmen KA, Trieu-Cuot P, Fitzgerald KA et al (2008) TLR-independent type I interferon induction in response to an extracellular bacterial pathogen via intracellular recognition of its DNA. Cell Host Microbe 4:543–554
Takaoka A, Wang Z, Choi MK, Yanai H, Negishi H et al (2007) DAI (DLM-1/ZBP1) is a cytosolic DNA sensor and an activator of innate immune response. Nature 448:501–505
Deshmukh SD, Kremer B, Freudenberg M, Bauer S, Golenbock DT et al (2011) Macrophages recognize streptococci through bacterial single-stranded RNA. EMBO Rep 12:71–76
Sinclair A, Xie X, Teltscher M, Dendukuri N (2013) Systematic review and meta-analysis of a urine-based pneumococcal antigen test for diagnosis of community-acquired pneumonia caused by Streptococcus pneumoniae. J Clin Microbiol 51:2303–2310
van der Poll T, Opal SM (2009) Pathogenesis, treatment, and prevention of pneumococcal pneumonia. Lancet 374:1543–1556
Koppe U, Suttorp N, Opitz B (2012) Recognition of Streptococcus pneumoniae by the innate immune system. Cell Microbiol 14:460–466
Patenge N, Fiedler T, Kreikemeyer B (2013) Common regulators of virulence in streptococci. Curr Top Microbiol Immunol 368:111–153
Mitchell AM, Mitchell TJ (2010) Streptococcus pneumoniae: virulence factors and variation. Clin Microbiol Infect 16:411–418
Knapp S, Wieland CW, van’t Veer C, Takeuchi O, Akira S et al (2004) Toll-like receptor 2 plays a role in the early inflammatory response to murine pneumococcal pneumonia but does not contribute to antibacterial defense. J Immunol 172:3132–3138
Schroder NW, Morath S, Alexander C, Hamann L, Hartung T et al (2003) Lipoteichoic acid (LTA) of Streptococcus pneumoniae and Staphylococcus aureus activates immune cells via Toll-like receptor (TLR)-2, lipopolysaccharide-binding protein (LBP), and CD14, whereas TLR-4 and MD-2 are not involved. J Biol Chem 278:15587–15594
Albiger B, Dahlberg S, Sandgren A, Wartha F, Beiter K et al (2007) Toll-like receptor 9 acts at an early stage in host defence against pneumococcal infection. Cell Microbiol 9:633–644
Malley R, Henneke P, Morse SC, Cieslewicz MJ, Lipsitch M et al (2003) Recognition of pneumolysin by Toll-like receptor 4 confers resistance to pneumococcal infection. Proc Natl Acad Sci U S A 100:1966–1971
Davis KM, Nakamura S, Weiser JN (2011) Nod2 sensing of lysozyme-digested peptidoglycan promotes macrophage recruitment and clearance of S. pneumoniae colonization in mice. J Clin Invest 121:3666–3676
Witzenrath M, Pache F, Lorenz D, Koppe U, Gutbier B et al (2011) The NLRP3 inflammasome is differentially activated by pneumolysin variants and contributes to host defense in pneumococcal pneumonia. J Immunol 187:434–440
McNeela EA, Burke A, Neill DR, Baxter C, Fernandes VE et al (2010) Pneumolysin activates the NLRP3 inflammasome and promotes proinflammatory cytokines independently of TLR4. PLoS Pathog 6:e1001191
Fang R, Tsuchiya K, Kawamura I, Shen Y, Hara H et al (2011) Critical roles of ASC inflammasomes in caspase-1 activation and host innate resistance to Streptococcus pneumoniae infection. J Immunol 187:4890–4899
Joyce EA, Popper SJ, Falkow S (2009) Streptococcus pneumoniae nasopharyngeal colonization induces type I interferons and interferon-induced gene expression. BMC Genomics 10:404
Koppe U, Hogner K, Doehn JM, Muller HC, Witzenrath M et al (2012) Streptococcus pneumoniae stimulates a STING- and IFN regulatory factor 3-dependent type I IFN production in macrophages, which regulates RANTES production in macrophages, cocultured alveolar epithelial cells, and mouse lungs. J Immunol 188:811–817
Li W, Moltedo B, Moran TM (2012) Type I interferon induction during influenza virus infection increases susceptibility to secondary Streptococcus pneumoniae infection by negative regulation of gammadelta T cells. J Virol 86:12304–12312
Jamieson AM, Pasman L, Yu S, Gamradt P, Homer RJ et al (2013) Role of tissue protection in lethal respiratory viral-bacterial coinfection. Science 340(6137):1230–1234
Hertzen E, Johansson L, Kansal R, Hecht A, Dahesh S et al (2012) Intracellular Streptococcus pyogenes in human macrophages display an altered gene expression profile. PLoS One 7:e35218
Nakagawa I, Amano A, Mizushima N, Yamamoto A, Yamaguchi H et al (2004) Autophagy defends cells against invading group A Streptococcus. Science 306:1037–1040
Yamaguchi H, Nakagawa I, Yamamoto A, Amano A, Noda T et al (2009) An initial step of GAS-containing autophagosome-like vacuoles formation requires Rab7. PLoS Pathog 5:e1000670
Prinz M, Schmidt H, Mildner A, Knobeloch KP, Hanisch UK et al (2008) Distinct and nonredundant in vivo functions of IFNAR on myeloid cells limit autoimmunity in the central nervous system. Immunity 28:675–686
Teijaro JR, Ng C, Lee AM, Sullivan BM, Sheehan KC et al (2013) Persistent LCMV infection is controlled by blockade of type I interferon signaling. Science 340:207–211
Wilson EB, Yamada DH, Elsaesser H, Herskovitz J, Deng J et al (2013) Blockade of chronic type I interferon signaling to control persistent LCMV infection. Science 340:202–207
Gonzalez-Navajas JM, Lee J, David M, Raz E (2012) Immunomodulatory functions of type I interferons. Nat Rev Immunol 12:125–135
Kovarik P, Sauer I, Schaljo B (2007) Molecular mechanisms of the anti-inflammatory functions of interferons. Immunobiology 212:895–901
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Kovarik, P., Castiglia, V., Janos, M. (2014). Type I Interferons in Immune Defense Against Streptococci. In: Parker, D. (eds) Bacterial Activation of Type I Interferons. Springer, Cham. https://doi.org/10.1007/978-3-319-09498-4_4
Download citation
DOI: https://doi.org/10.1007/978-3-319-09498-4_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-09497-7
Online ISBN: 978-3-319-09498-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)