Abstract
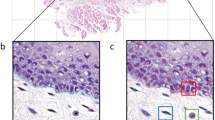
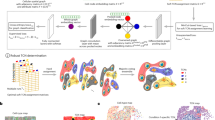
Graphs are useful in analysing histopathological images as they are able to represent neighbourhood interactions and spatial relationships. Typically graph nodes represent cells and the vertices are constructed by applying a nearest neighbor algorithm to cell’s locations. When passing these graphs through one graph neural network (GNN) message passing step, each node can only utilise features from nodes within its immediate neighbourhood to make a classification. To overcome this, we introduce two levels of hierarchically connected nodes that we term “supernodes”. These supernodes, used in conjunction with at least four GNN message passing steps, allow for cell node classifications to be influenced by a wider area, enabling the entire graph to learn tissue-level structures. The method is evaluated on a supervised task to classify individual cells as belonging to a specific tissue class. Results demonstrate that the inclusion of supernodes with multiple GNN message passing steps increases model accuracy.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Pati, P., et al.: Hierarchical graph representations in digital pathology. Med. Image Anal. 75, 102264 (2022)
Mi, H., et al.: Digital pathology analysis quantifies spatial heterogeneity of CD3, CD4, CD8, CD20, and FoxP3 immune markers in triple-negative breast cancer. Front. Physiol. 11, 583333 (2020)
Janowczyk, A., Madabhushi, A.: Deep learning for digital pathology image analysis: a comprehensive tutorial with selected use cases. J. Pathol. Inform. 7, 29 (2016)
Zhou, Y., et al.: CGC-net: cell graph convolutional network for grading of colorectal cancer histology images. In: 2019 IEEE/CVF International Conference on Computer Vision Workshop (ICCVW) (2019)
Lu, W., et al.: Capturing cellular topology in multi-gigapixel pathology images. In: 2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW) (2020)
Anklin, V., et al.: Learning whole-slide segmentation from inexact and incomplete labels using tissue graphs (2021). arXiv:2103.03129
Veličković, P., et al.: Graph attention networks (2017). arXiv:1710.10903
Gao, J.P., et al.: Tumor heterogeneity of gastric cancer: from the perspective of tumor-initiating cell. World J. Gastroenterol. 24(24), 2567–2581 (2018)
Anghel, A., et al.: A high-performance system for robust stain normalization of whole-slide images in histopathology. Front. Med. (Lausanne) 6, 193 (2019)
Chen, J., Chen, H.: Edge-featured graph attention network (2021). arXiv:2101.07671
Xu, K., et al.: Representation learning on graphs with jumping knowledge networks (2018). arXiv:1806.03536
Junttila, M.R., de Sauvage, F.J.: Influence of tumour micro-environment heterogeneity on therapeutic response. Nature 501(7467), 346–354 (2013)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Sims, J., Grabsch, H.I., Magee, D. (2022). Using Hierarchically Connected Nodes and Multiple GNN Message Passing Steps to Increase the Contextual Information in Cell-Graph Classification. In: Manfredi, L., et al. Imaging Systems for GI Endoscopy, and Graphs in Biomedical Image Analysis. ISGIE GRAIL 2022 2022. Lecture Notes in Computer Science, vol 13754. Springer, Cham. https://doi.org/10.1007/978-3-031-21083-9_10
Download citation
DOI: https://doi.org/10.1007/978-3-031-21083-9_10
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-21082-2
Online ISBN: 978-3-031-21083-9
eBook Packages: Computer ScienceComputer Science (R0)