Abstract
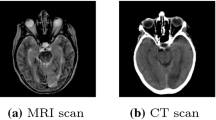
In medical image synthesis, model training could be challenging due to the inconsistencies between images of different modalities even with the same patient, typically caused by internal status/tissue changes as different modalities are usually obtained at a different time. This paper proposes a novel deep learning method, Structure-aware Generative Adversarial Network (SA-GAN), that preserves the shapes and locations of in-consistent structures when generating medical images. SA-GAN is employed to generate synthetic computed tomography (synCT) images from magnetic resonance imaging (MRI) with two parallel streams: the global stream translates the input from the MRI to the CT domain while the local stream automatically segments the inconsistent organs, maintains their locations and shapes in MRI, and translates the organ intensities to CT. Through extensive experiments on a pelvic dataset, we demonstrate that SA-GAN provides clinically acceptable accuracy on both synCTs and organ segmentation and supports MR-only treatment planning in disease sites with internal organ status changes.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Armanious, K., et al.: MedGAN: medical image translation using GANs. Computerized Med. Imag. Graph. 79, 101684 (2020)
Chen, S., Qin, A., Zhou, D., Yan, D.: U-net-generated synthetic CT images for magnetic resonance imaging-only prostate intensity-modulated radiation therapy treatment planning. Med. Phys. 45(12), 5659–5665 (2018)
Emami, H., Dong, M., Glide-Hurst, C.K.: Attention-guided generative adversarial network to address atypical anatomy in synthetic CT generation. In: 2020 IEEE 21st International Conference on Information Reuse and Integration for Data Science (IRI), pp. 188–193. IEEE (2020)
Emami, H., Dong, M., Nejad-Davarani, S.P., Glide-Hurst, C.K.: Generating synthetic CTs from magnetic resonance images using generative adversarial networks. Med. Phys. 45(8), 3627–3636 (2018)
Emami, H., Liu, Q., Dong, M.: FREA-UNet: frequency-aware U-Net for modality transfer. arXiv preprint arXiv:2012.15397 (2020)
Fu, J., et al.: Deep learning approaches using 2D and 3D convolutional neural networks for generating male pelvic synthetic computed tomography from magnetic resonance imaging. Med. Phys. 46(9), 3788–3798 (2019)
Fu, J., Yang, Y., Singhrao, K., Ruan, D., Low, D.A., Lewis, J.H.: Male pelvic synthetic CT generation from t1-weighted MRI using 2D and 3D convolutional neural networks. arXiv preprint arXiv:1803.00131 (2018)
Gatys, L.A., Ecker, A.S., Bethge, M.: Image style transfer using convolutional neural networks. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 2414–2423 (2016)
Ge, Y., et al.: Unpaired MR to CT synthesis with explicit structural constrained adversarial learning. In: 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019), pp. 1096–1099. IEEE (2019)
Ghiasi, G., Lee, H., Kudlur, M., Dumoulin, V., Shlens, J.: Exploring the structure of a real-time, arbitrary neural artistic stylization network. arXiv preprint arXiv:1705.06830 (2017)
Goodfellow, I., et al.: Generative adversarial nets. In: Advances in Neural Information Processing Systems, pp. 2672–2680 (2014)
Hamghalam, M., Lei, B., Wang, T.: High tissue contrast MRI synthesis using multi-stage attention-GAN for glioma segmentation. arXiv preprint arXiv:2006.05030 (2020)
Han, X.: MR-based synthetic CT generation using a deep convolutional neural network method. Med. Phys. 44(4), 1408–1419 (2017)
Huang, X., Belongie, S.: Arbitrary style transfer in real-time with adaptive instance normalization. In: Proceedings of the IEEE International Conference on Computer Vision, pp. 1501–1510 (2017)
Huynh, T., et al.: Estimating CT image from MRI data using structured random forest and auto-context model. IEEE Trans. Med. Imag. 35(1), 174–183 (2015)
Isola, P., Zhu, J.Y., Zhou, T., Efros, A.A.: Image-to-image translation with conditional adversarial networks. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 1125–1134 (2017)
Kim, J., et al.: Dosimetric evaluation of synthetic CT relative to bulk density assignment-based magnetic resonance-only approaches for prostate radiotherapy. Radiat. Oncol. 10(1), 239 (2015)
Lei, Y., et al.: MRI-only based synthetic CT generation using dense cycle consistent generative adversarial networks. Med. Phys. 46(8), 3565–3581 (2019)
Lin, J., Xia, Y., Qin, T., Chen, Z., Liu, T.Y.: Conditional image-to-image translation. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 5524–5532 (2018)
Mao, X., Li, Q., Xie, H., Lau, R.Y., Wang, Z., Paul Smolley, S.: Least squares generative adversarial networks. In: Proceedings of the IEEE International Conference on Computer Vision, pp. 2794–2802 (2017)
Maspero, M., et al.: Dose evaluation of fast synthetic-CT generation using a generative adversarial network for general pelvis MR-only radiotherapy. Phys. Med. Biol. 63(18), 185001 (2018)
Nakamura, N., et al.: Variability in bladder volumes of full bladders in definitive radiotherapy for cases of localized prostate cancer. Strahlentherapie und Onkologie 186(11), 637–642 (2010)
Nie, D., Cao, X., Gao, Y., Wang, L., Shen, D.: Estimating CT image from MRI data using 3D fully convolutional networks. In: Carneiro, G., et al. (eds.) LABELS/DLMIA -2016. LNCS, vol. 10008, pp. 170–178. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-46976-8_18
Nie, D., et al.: Medical image synthesis with deep convolutional adversarial networks. IEEE Trans. Biomed. Eng. 65(12), 2720–2730 (2018)
Nie, D., et al.: Medical image synthesis with context-aware generative adversarial networks. In: Descoteaux, M., Maier-Hein, L., Franz, A., Jannin, P., Collins, D.L., Duchesne, S. (eds.) MICCAI 2017. LNCS, vol. 10435, pp. 417–425. Springer, Cham (2017). https://doi.org/10.1007/978-3-319-66179-7_48
Pan, Y., Liu, M., Lian, C., Zhou, T., Xia, Y., Shen, D.: Synthesizing missing PET from MRI with cycle-consistent generative adversarial networks for Alzheimer’s disease diagnosis. In: Frangi, A.F., Schnabel, J.A., Davatzikos, C., Alberola-López, C., Fichtinger, G. (eds.) MICCAI 2018. LNCS, vol. 11072, pp. 455–463. Springer, Cham (2018). https://doi.org/10.1007/978-3-030-00931-1_52
Tie, X., Lam, S.K., Zhang, Y., Lee, K.H., Au, K.H., Cai, J.: Pseudo-CT generation from multi-parametric MRI using a novel multi-channel multi-path conditional generative adversarial network for nasopharyngeal carcinoma patients. Med. Phys. 47(4), 1750–1762 (2020)
Wolterink, J.M., Dinkla, A.M., Savenije, M.H.F., Seevinck, P.R., van den Berg, C.A.T., Išgum, I.: Deep MR to CT synthesis using unpaired data. In: Tsaftaris, S.A., Gooya, A., Frangi, A.F., Prince, J.L. (eds.) SASHIMI 2017. LNCS, vol. 10557, pp. 14–23. Springer, Cham (2017). https://doi.org/10.1007/978-3-319-68127-6_2
Zhong, H., Wen, N., Gordon, J.J., Elshaikh, M.A., Movsas, B., Chetty, I.J.: An adaptive MR-CT registration method for MRI-guided prostate cancer radiotherapy. Phys. Med. Biol. 60(7), 2837 (2015)
Zhu, J.Y., Park, T., Isola, P., Efros, A.A.: Unpaired image-to-image translation using cycle-consistent adversarial networks. In: Proceedings of the IEEE International Conference on Computer Vision, pp. 2223–2232 (2017)
Acknowledgments
This work was partially supported by the National Cancer Institute of the National Institutes of Health under Award Number R01CA204189.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Emami, H., Dong, M., Nejad-Davarani, S.P., Glide-Hurst, C.K. (2021). SA-GAN: Structure-Aware GAN for Organ-Preserving Synthetic CT Generation. In: de Bruijne, M., et al. Medical Image Computing and Computer Assisted Intervention – MICCAI 2021. MICCAI 2021. Lecture Notes in Computer Science(), vol 12906. Springer, Cham. https://doi.org/10.1007/978-3-030-87231-1_46
Download citation
DOI: https://doi.org/10.1007/978-3-030-87231-1_46
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-87230-4
Online ISBN: 978-3-030-87231-1
eBook Packages: Computer ScienceComputer Science (R0)