Abstract
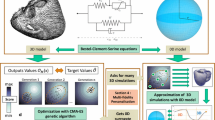
We propose a strategy to perform population-based personalization of a model, to overcome the limits of case-based personalization for generating virtual populations from models that include randomness. We formulate the problem as matching the synthetic and real populations by minimizing the Kullback-Leibler divergence between their distributions. As an analytical formulation of the models is complex or even impossible, the personalization is addressed by a gradient-free method: the CMA-ES algorithm, whose relevance was demonstrated for the case-based personalization of complex biomechanical cardiac models. The algorithm iteratively adapts the covariance matrix which in our problem encodes the distribution of the synthetic data.
We demonstrate the feasibility of this approach on two simple geometrical models of myocardial infarction, in 2D, to better focus on the relevance of the personalization process. Our strategy is able to reproduce the distribution of 2D myocardial infarcts from the segmented late Gadolinium images of 123 subjects with acute myocardial infarction.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Connolly, A.J., Bishop, M.J.: Computational representations of myocardial infarct scars and implications for arrhythmogenesis. Clin. Med. Insights Cardiol. 10, 27–40 (2016)
Kingma, D.P., Welling, M.: Auto-encoding variational Bayes. arXiv (2014)
Kerckhoffs, R.C., McCulloch, A.D., Omens, J.H., et al.: Effects of biventricular pacing and scar size in a computational model of the failing heart with left bundle branch block. Med. Image Anal. 13, 362–369 (2009). https://doi.org/10.1016/j.media.2008.06.013
Duchateau, N., De Craene, M., Allain, P., et al.: Infarct localization from myocardial deformation: prediction and uncertainty quantification by regression from a low-dimensional space. IEEE Trans. Med. Imaging 35, 2340–2352 (2016). https://doi.org/10.1109/TMI.2016.2562181
Leong, C.N., Lim, E., Andriyana, A., et al.: The role of infarct transmural extent in infarct extension: a computational study. Int. J. Numer. Method Biomed. Eng. 33, e02794 (2017). https://doi.org/10.1002/cnm.2794
Reimer, K.A., Lowe, J.E., Rasmussen, M.M., et al.: The wavefront phenomenon of ischemic cell death. 1. Myocardial infarct size vs duration of coronary occlusion in dogs. Circulation 56, 786–794 (1977). https://doi.org/10.1161/01.cir.56.5.786
Pashaei, A., Hoogendoorn, C., Sebastián, R., Romero, D., Cámara, O., Frangi, A.F.: Effect of scar development on fast electrophysiological models of the human heart: in-silico study on atlas-based virtual populations. In: Metaxas, D.N., Axel, L. (eds.) FIMH 2011. LNCS, vol. 6666, pp. 427–436. Springer, Heidelberg (2011). https://doi.org/10.1007/978-3-642-21028-0_54
Rumindo, G.K., Duchateau, N., Croisille, P., Ohayon, J., Clarysse, P.: Strain-based parameters for infarct localization: evaluation via a learning algorithm on a synthetic database of pathological hearts. In: Pop, M., Wright, G.A. (eds.) FIMH 2017. LNCS, vol. 10263, pp. 106–114. Springer, Cham (2017). https://doi.org/10.1007/978-3-319-59448-4_11
Belle, L., Motreff, P., Mangin, L., et al.: Comparison of immediate with delayed stenting using the minimalist immediate mechanical intervention approach in acute ST-segment-elevation myocardial infarction: the MIMI study. Circ. Cardiovasc. Interv. 9, e003388 (2016). https://doi.org/10.1161/CIRCINTERVENTIONS.115.003388
Hansen, N., Ostermeier, A.: Adapting arbitrary normal mutation distributions in evolution strategies: the covariance matrix adaptation. In: Proceedings ICEC, pp. 312–317 (1996). https://doi.org/10.1109/ICEC.1996.542381
Molléro, R., Pennec, X., Delingette, H., Garny, A., Ayache, N., Sermesant, M.: Multifidelity-CMA: a multifidelity approach for efficient personalisation of 3D cardiac electromechanical models. Biomech. Model. Mechanobiol 17(1), 285–300 (2017). https://doi.org/10.1007/s10237-017-0960-0
Duchateau, N., De Craene, M., Sitges, M., Caselles, V.: Adaptation of multiscale function extension to inexact matching: application to the mapping of individuals to a learnt manifold. In: Nielsen, F., Barbaresco, F. (eds.) GSI 2013. LNCS, vol. 8085, pp. 578–586. Springer, Heidelberg (2013). https://doi.org/10.1007/978-3-642-40020-9_64
Gretton, A., Borgwardt, K.M., Rasch, M.J., et al.: A kernel two-sample test. J. Mach. Learn. Res. 13, 723–73 (2012)
Acknowledgements
The authors acknowledge the support from the French ANR (LABEX PRIMES of Univ. Lyon [ANR-11-LABX-0063] within the program “Investissements d’Avenir” [ANR-11-IDEX-0007], the JCJC project “MIC-MAC” [ANR-19-CE45-0005]), and the Fédération Francaise de Cardiologie (“MI-MIX” project, Allocation René Foudon). They are also grateful to P Croisille and M Viallon (CREATIS, CHU Saint Etienne) for providing the imaging data for the MIMI population, and M Di Folco (CREATIS) for the preliminary exploration of the data alignment and simulation tools.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Mom, K., Clarysse, P., Duchateau, N. (2021). Population-Based Personalization of Geometric Models of Myocardial Infarction. In: Ennis, D.B., Perotti, L.E., Wang, V.Y. (eds) Functional Imaging and Modeling of the Heart. FIMH 2021. Lecture Notes in Computer Science(), vol 12738. Springer, Cham. https://doi.org/10.1007/978-3-030-78710-3_1
Download citation
DOI: https://doi.org/10.1007/978-3-030-78710-3_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-78709-7
Online ISBN: 978-3-030-78710-3
eBook Packages: Computer ScienceComputer Science (R0)