Abstract
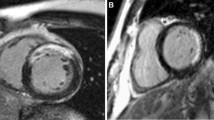
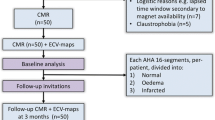
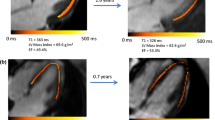
Many uncertainties remain about the relation between post-infarct scars and ventricular arrhythmia. Most post-infarct patients suffer scar-related arrhythmia several years after the infarct event suggesting that scar remodeling is a process that might require years until the affected tissue becomes arrhythmogenic. In clinical practice, a simple time-based rule is often used to assess risk and stratify patients. In other cases, left ventricular ejection fraction (LVEF) impairment is also taken into account but it is known to be suboptimal. More information is needed to better stratify patients and prescribe appropriate individualized treatments. In this paper we propose to use probabilistic disease progression modeling to obtain an image-based data-driven description of the infarct maturation process. Our approach includes monotonic constraints in order to impose a regular behaviour on the biomarkers’ trajectories. 49 post-MI patients underwent Computed Tomography (CT) and Late Gadolinium Enhanced Cardiac Magnetic Resonance (LGE-CMR) scans. Image-derived biomarkers were computed such as LVEF, LGE-CMR scar volume, fat volume, and size of areas with a different degree of left ventricular wall narrowing, from moderate to severe. We show that the model is able to estimate a plausible progression of post-infarct scar maturation. According to our results there is a progressive thinning process observable only with CT imaging; intramural fat appears in a late stage; LGE-CMR scar volume almost does not change and LVEF slightly changes during the scar maturation process.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Al-Khatib, S., et al.: AHA/ACC/HRS guideline for management of patients with ventricular arrhythmias and the prevention of sudden cardiac death: executive summary: a report of the American College of Cardiology/American Heart Association task force on clinical practice guidelines and the heart rhythm society. Heart Rhythm 35, e-91–e-220 (2017)
Anversa, P., Olivetti, G., Capasso, J.M.: Cellular basis of ventricular remodeling after myocardial infarction. Am. J. Cardiol 68(14), 7–16 (1991)
Cedilnik, N., Duchateau, J., Sacher, F., Jaïs, P., Cochet, H., Sermesant, M.: Fully automated electrophysiological model personalisation framework from CT imaging. In: Coudière, Y., Ozenne, V., Vigmond, E., Zemzemi, N. (eds.) FIMH 2019. LNCS, vol. 11504, pp. 325–333. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-21949-9_35
Goldberger, J.J., Cain, M.E., Hohnloser, S.H., et al.: American Heart Association/American College of Cardiology Foundation/Heart Rhythm Society scientific statement on noninvasive risk stratification techniques for identifying patients at risk for sudden cardiac death: a scientific statement from the American Heart Association Council on clinical cardiology committee on electrocardiography and arrhythmias and council on epidemiology and prevention. J. Am. Coll. Cardiol. 52(14), 1179–1199 (2008)
Jáuregui, B., Soto-Iglesias, D., Penela, D., Acosta, J., et al.: Follow-up after myocardial infarction to explore the stability of arrhythmogenic substrate: the FOOTPRINT study. JACC Clin. Electrophysiol. 6(2), 207–218 (2020)
Jia, S., et al.: Automatically segmenting the left atrium from cardiac images using successive 3D U-nets and a contour loss. In: International Workshop on Statistical Atlases and Computational Models of the Heart, pp. 221–229 (2018)
Lorenzi, M., Filippone, M., Frisoni, G.B., Alexander, D.C., et al.: Probabilistic disease progression modeling to characterize diagnostic uncertainty: application to staging and prediction in Alzheimer’s disease. NeuroImage 190, 56–68 (2019)
Martin, C.A., Gajendragadkar, P.R.: scar tissue: never too old to remodel. JACC Clini. Electrophysiol. 6(2), 219–220 (2020)
Martin, R., et al.: Characteristics of scar-related ventricular tachycardia circuits using ultra-high-density mapping: a multi-center study. Circ. Arrhythm. Electrophysiol 11(10), e006569 (2018)
Rasmussen, C.E.: Gaussian processes in machine learning. In: Bousquet, O., von Luxburg, U., Rätsch, G. (eds.) ML -2003. LNCS (LNAI), vol. 3176, pp. 63–71. Springer, Heidelberg (2004). https://doi.org/10.1007/978-3-540-28650-9_4
Ronneberger, O., Fischer, P., Brox, T.: U-Net: convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015, Part III. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Yezzi, A.J., Prince, J.L.: An Eulerian PDE approach for computing tissue thickness. IEEE Trans. Med. Imaging 22(10), 1332–1339 (2003)
Acknowledgements
Part of this work was funded by the ERC starting grant EC-STATIC (715093), the IHU LIRYC (ANR-10-IAHU-04), the Equipex MUSIC (ANR-11-EQPX-0030) and the ANR ERACoSysMed SysAFib projects. This work was also supported by the French government, through the 3IA Côte d’Azur Investments in the Future project managed by the National Research Agency (ANR) with the reference number ANR-19-P3IA-0002, and by the ANR JCJC project Fed-BioMed 19-CE45-0006-01. We would like to thank all patients who agreed to make available their clinical data for research.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Nuñez-Garcia, M., Cedilnik, N., Jia, S., Cochet, H., Lorenzi, M., Sermesant, M. (2021). Estimation of Imaging Biomarker’s Progression in Post-infarct Patients Using Cross-sectional Data. In: Puyol Anton, E., et al. Statistical Atlases and Computational Models of the Heart. M&Ms and EMIDEC Challenges. STACOM 2020. Lecture Notes in Computer Science(), vol 12592. Springer, Cham. https://doi.org/10.1007/978-3-030-68107-4_11
Download citation
DOI: https://doi.org/10.1007/978-3-030-68107-4_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-68106-7
Online ISBN: 978-3-030-68107-4
eBook Packages: Computer ScienceComputer Science (R0)