Abstract
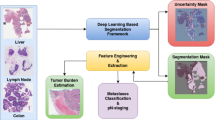
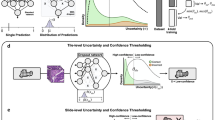
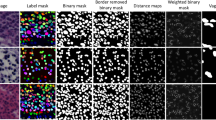
Deep learning assisted histopathology has the potential to extract reproducible and accurate measurements from digitised slides in a scalable fashion. A typical workflow of such analysis may involve instance segmentation of relevant tissues followed by feature measurements. Inherent segmentation uncertainties produced by these deep models, however, could propagate to the downstream measurements, causing biased distribution estimate of the whole slide. One challenging aspect when handling ambiguous tissues is that the number of instances could differ as the instance segmentation step may not generalise well to these tissues. As an attempt to address this problem, we propose to derive a confidence score from the segmentation uncertainties obtained from Bayesian Neural Networks (BNNs) and utilise these as weights to improve the distribution estimate. We generate a synthetic dataset that mimics the diverse and varying visual features of the original data to enable systematic experiments. With this dataset we demonstrate the robustness of the method by extracting several clinically relevant measurements with two different BNNs. Our results indicate that the distribution estimates are consistently improved when the instances are weighted by the entropy-derived confidence measure. In addition, we provide results on applying the method to the original data.
KHT is funded by the EPSRC and MRC grant number EP/L016052/1. JR and KS are supported by the Oxford NIHR Biomedical Research Centre and the PathLAKE consortium (Innovate UK App. Nr. 18181).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Blundell, C., Cornebise, J., Kavukcuoglu, K., Wierstra, D.: Weight uncertainty in neural networks. arXiv preprint arXiv:1505.05424 (2015)
Boykov, Y., Funka-Lea, G.: Graph cuts and efficient ND image segmentation. Int. J. Comput. Vis. 70(2), 109–131 (2006). https://doi.org/10.1007/s11263-006-7934-5
Chatrian, A., Sirinukunwattana, K., Verrill, C., Rittscher, J.: Towards the identification of histology based subtypes in prostate cancer. In: 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019), pp. 948–952. IEEE (2019)
DeVries, T., Taylor, G.W.: Leveraging uncertainty estimates for predicting segmentation quality. arXiv preprint arXiv:1807.00502 (2018)
Gal, Y., Ghahramani, Z.: Dropout as a Bayesian approximation: representing model uncertainty in deep learning. In: International Conference on Machine Learning, pp. 1050–1059 (2016)
Isola, P., Zhu, J.Y., Zhou, T., Efros, A.A.: Image-to-image translation with conditional adversarial networks. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp. 1125–1134 (2017)
Kendall, A., Badrinarayanan, V., Cipolla, R.: Bayesian SegNet: model uncertainty in deep convolutional encoder-decoder architectures for scene understanding. arXiv preprint arXiv:1511.02680 (2015)
Kendall, A., Gal, Y.: What uncertainties do we need in Bayesian deep learning for computer vision? In: Advances in Neural Information Processing Systems, pp. 5574–5584 (2017)
Kingma, D.P., Welling, M.: Auto-encoding variational bayes. arXiv preprint arXiv:1312.6114 (2013)
Kingma, D.P., Salimans, T., Welling, M.: Variational dropout and the local reparameterization trick. In: Advances in Neural Information Processing Systems, pp. 2575–2583 (2015)
Kohl, S., et al.: A probabilistic U-Net for segmentation of ambiguous images. In: Advances in Neural Information Processing Systems, pp. 6965–6975 (2018)
Kwon, Y., Won, J.H., Kim, B.J., Paik, M.C.: Uncertainty quantification using Bayesian neural networks in classification: application to biomedical image segmentation. Comput. Stat. Data Anal. 142, 106816 (2020)
Lakshminarayanan, B., Pritzel, A., Blundell, C.: Simple and scalable predictive uncertainty estimation using deep ensembles. In: Advances in Neural Information Processing Systems, pp. 6402–6413 (2017)
Larsen, A.B.L., Sønderby, S.K., Larochelle, H., Winther, O.: Autoencoding beyond pixels using a learned similarity metric. In: International Conference on Machine Learning, pp. 1558–1566 (2016)
Lin, T.Y., Goyal, P., Girshick, R., He, K., Dollár, P.: Focal loss for dense object detection. In: Proceedings of the IEEE International Conference on Computer Vision, pp. 2980–2988 (2017)
Maas, A.L., Hannun, A.Y., Ng, A.Y.: Rectifier nonlinearities improve neural network acoustic models. In: Proceedings of the ICML, vol. 30, no. 1, p. 3 (2013)
Miller, D., Dayoub, F., Milford, M., Sünderhauf, N.: Evaluating merging strategies for sampling-based uncertainty techniques in object detection. In: 2019 International Conference on Robotics and Automation (ICRA), pp. 2348–2354. IEEE (2019)
Miller, D., Nicholson, L., Dayoub, F., Sünderhauf, N.: Dropout sampling for robust object detection in open-set conditions. In: 2018 IEEE International Conference on Robotics and Automation (ICRA), pp. 1–7. IEEE (2018)
Morrison, D., Milan, A., Antonakos, E.: Uncertainty-aware instance segmentation using dropout sampling. Technical report, CVPR Robotic Vision Probabilistic Object Detection Challenge (2019)
Mukhoti, J., Gal, Y.: Evaluating Bayesian deep learning methods for semantic segmentation. arXiv preprint arXiv:1811.12709 (2018)
Ng, M., Guo, F., Biswas, L., Wright, G.A.: Estimating uncertainty in neuralnetworks for segmentation quality control. Technical report (2018). https://www.doc.ic.ac.uk/~bglocker/public/mednips2018/med-nips_2018_paper_105.pdf
Nketia, T.A., Noble, J.A., Rittscher, J.: Towards quantifying the impact of cell boundary estimation on morphometric analysis for phenotypic screening. In: 2015 IEEE 12th International Symposium on Biomedical Imaging (ISBI), pp. 781–784. IEEE (2015)
Orlando, J.I., et al.: U2-Net: a Bayesian U-Net model with epistemic uncertainty feedback for photoreceptor layer segmentation in pathological oct scans. In: 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019), pp. 1441–1445. IEEE (2019)
Ramdas, A., Trillos, N.G., Cuturi, M.: On wasserstein two-sample testing and related families of nonparametric tests. Entropy 19(2), 47 (2017)
Ronneberger, O., Fischer, P., Brox, T.: U-Net: convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Shridhar, K., Laumann, F., Liwicki, M.: A comprehensive guide to Bayesian convolutional neural network with variational inference. arXiv preprint arXiv:1901.02731 (2019)
Wang, Z., Bovik, A.C., Sheikh, H.R., Simoncelli, E.P.: Image quality assessment: from error visibility to structural similarity. IEEE Trans. Image Process. 13(4), 600–612 (2004)
Weissenbacher, A.: Normothermic kidney preservation. Ph.D. thesis, University of Oxford (2018)
Weissenbacher, A., Lo Faro, L., Boubriak, O., Soares, M.F., Roberts, I.S., Hunter, J.P., Voyce, D., Mikov, N., Cook, A., Ploeg, R.J., et al.: Twenty-four-hour normothermic perfusion of discarded human kidneys with urine recirculation. Am. J. Transplant. 19(1), 178–192 (2019)
Zhu, J.Y., Park, T., Isola, P., Efros, A.A.: Unpaired image-to-image translation using cycle-consistent adversarial networks. In: Proceedings of the IEEE International Conference on Computer Vision, pp. 2223–2232 (2017)
Zhu, J.Y., et al.: Toward multimodal image-to-image translation. In: Advances in neural information processing systems, pp. 465–476 (2017)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this paper
Cite this paper
Tam, K.H., Sirinukunwattana, K., Soares, M.F., Kaisar, M., Ploeg, R., Rittscher, J. (2020). Improving Pathological Distribution Measurements with Bayesian Uncertainty. In: Sudre, C.H., et al. Uncertainty for Safe Utilization of Machine Learning in Medical Imaging, and Graphs in Biomedical Image Analysis. UNSURE GRAIL 2020 2020. Lecture Notes in Computer Science(), vol 12443. Springer, Cham. https://doi.org/10.1007/978-3-030-60365-6_7
Download citation
DOI: https://doi.org/10.1007/978-3-030-60365-6_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-60364-9
Online ISBN: 978-3-030-60365-6
eBook Packages: Computer ScienceComputer Science (R0)