Abstract
Interpolated climate surfaces have been widely used to predict species distributions and develop environmental niche models. However, the spatial coverage and density of meteorological sites used to develop these surfaces vary among countries and regions, such that the most biodiverse regions often have the most sparsely sampled climatic data. We explore the potential of satellite remote sensing (S-RS) products—which have consistently high spatial and temporal resolution and nearly global coverage—to quantify species-environment relationships that predict species distributions. We propose several new environmental metrics that take advantage of high temporal resolution in S-RS data and compare these approaches to classic climate-only approaches using the live oaks (Quercus section Virentes) as a case study. We show that models perform similarly but for some species, particularly in understudied regions, show less precision in predicting spatial distribution. These results provide evidence supporting efforts to enhance environmental niche models and species distribution models (ENMs/SDMs) with S-RS data and, when combined with other approaches for species detection, will likely enhance our ability to monitor biodiversity globally.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
- Species distribution models
- Environmental niche models
- Biotic niche
- Abiotic niche
- Remotely sensed environmental data
- Climate variables
- Phenology
9.1 Introduction
What drives species distributions? This is one of the most fundamental questions in ecology, evolution, and biogeography, and it drew the attention of early naturalists (Gaston 2009; Guisan et al. 2017). Although the question is classic and its answers sometime seem obvious—for example, Alfred Russel Wallace recognized the effect of geographical and environmental features on species distributional ranges (Wallace 1860)—the answers are highly complex as a consequence of historical evolutionary and biogeographic processes and the spatial and temporal dynamics of abiotic and biotic factors (Soberón and Peterson 2005; Soberón 2007; Colwell and Rangel 2009).
Here we explore the potential of satellite remote sensing (S-RS) products to quantify species-environment relationships that predict species distributions. We propose several new metrics that take advantage of the high temporal resolution in Moderate Resolution Imaging Spectroradiometer (MODIS) leaf area index (LAI) and MODIS normalized difference vegetation index (NDVI) data products. Evaluating the potential of remotely sensed data in environmental niche modeling (ENM) and species distribution modeling (SDM) is an important step toward the long-term goal of improving our ability to monitor and predict changes in biodiversity globally. To achieve this, we first modeled the environmental/ecological niches for the American live oak species (Quercus section Virentes) using environmental variables derived from (1) interpolated climate surfaces data (i.e., WorldClim) and (2) S-RS products. Live oaks are a small lineage, descended from a common ancestor (for a discussion on phylogenetics, see Meireles et al., Chap. 7) that includes seven species that vary in geographic range size and climatic breadth and are distributed in both temperate and tropical climates from the southeastern United States, Mesoamerica, and the Caribbean (Cavender-Bares et al. 2015). Their variation in range size and climatic distributions, their distributions in both highly studied and understudied regions of the globe, and the second author’s expert knowledge of their distributions make them an interesting case study for comparing SDM/ENMs that rely on classic data sources to those that use remotely sensed data sources, which have more consistent data accuracy and resolution. We used the live oaks as a test clade to evaluate the relationships among the modeled niches estimated from both sources of environmental data.
Given that the interpolated climate surfaces from WorldClim (Hijmans et al. 2005) are the most widely used data set for the study of species-environment relationships, we compare the performance of SDMs based on S-RS products to those based on WorldClim data. If there is a tight relationship between models from the two sources, this would indicate that the resultant models from S-RS products have similar performance to the resultant models from the WorldClim climatic predictors. Remotely sensed data products may provide an advantage in predicting species distributions in regions where climatic data is sparsely sampled. Although WorldClim provides interpolated climate surfaces for land areas across the world at multiple spatial resolutions, from 30 arc seconds (~1 km) to 10 arcmin (~18.5 km) (Hijmans et al. 2005), the spatial distribution of the base information (i.e., weather or climatic stations) used for interpolations is unevenly distributed across the world (Fig. 9.2c). This is not a small issue given the uncertainty associated with interpolated climatic variables when modeling species-environment relationships, especially in many tropical countries, where weather stations are frequently few and far apart (Soria-Auza et al. 2010). Given that tropical regions are precisely the regions where most species occur (Fig. 9.2c), finding alternative means to predict species is important for efforts to monitor and manage biodiversity globally. S-RS products, which provide quasi-global coverage of land and sea surfaces at high temporal and spatial resolution, represent promising alternatives that may be particularly important in the world’s most biologically diverse regions. Our aim here is to provide an understanding of the potential of S-RS products to quantify species ecological niches and estimate species distributions rather than to develop a definitive ecological and geographical profile for the live oaks themselves. If the consistent accuracy and high spatial resolution of S-RS products can actually improve estimates of species distributions, they will represent an advance in our ability to predict where species are likely persist under changing environments. Ultimately, such predictions can be combined with other remote sensing (RS) means of detecting species and biodiversity (Meireles et al., Chap. 7; Bolch et al., Chap. 12; Record et al., Chap. 10) to enable global-scale biodiversity change detection.
9.2 Theoretical Background
9.2.1 The BAM Diagram
One way to explore the ubiquitous relationship between the spatial and temporal dynamics of abiotic and biotic factors is through the BAM framework (Fig. 9.1, Soberón and Peterson 2005; Soberón 2007), which formally describes the individual and joint effects of biotic factors (B; e.g., species interactions), abiotic factors (A; e.g., environmental conditions or abiotically suitable area), and movement (M; e.g., species dispersal capacity) in determining species distributions in a geographical space (G; e.g., the study region). Notice that all factors in the BAM framework are placed within a spatial context. Within the geographical space (G), three overlapping circles are shown, each of which represents suitable conditions for a given species. The intersection between all factors “B∩A∩M” represents the occupied distributional area (G0) or the “realized” or occupied niche. The intersection between biotic and abiotic factors “B∩A” represents the invadable distributional area (G1) or areas that can be colonized because suitable biotic and abiotic conditions and both present. The intersection between abiotic and movement factors “A∩M” represents the area where the species cannot be found. Finally, the union between the occupied and invadable areas “G0∪G1” represents the geographic potential distribution area (GP) or biotically reduced niche (see Soberón 2007; Peterson et al. 2011 for detailed explanations).
The BAM diagram, where B biotic, A abiotically suitable area, and M movement or migration, illustrates the relationship among the three major determinants of species distributions. G1, the invadable distributional area, and G0, the occupied distributional area, represent the outcomes from the intersection between the major determinants
9.2.2 Where Are We Now?
Although the BAM framework was developed to understand and quantify species-environment relationships (Soberón 2007; see also Soberón and Peterson 2005), the concept and investigation of species-environment relationships are long-standing, dating back to Wallace (Wallace 1860) and early ecologists (Grinnell 1904, 1917; Elton 1927; Holdridge 1947; Hutchinson 1957). These early naturalists originally established the theoretical principles to analyze and describe biogeographical distributions in relation to environmental patterns (Colwell and Rangel 2009). Interestingly, despite the large body of theoretical advances and empirical applications, the quantification of ecological niches and estimation of species distributions is still a challenging task (but see Sanín and Anderson 2018; Smith et al. 2018) and one of the most active areas in macroecological and biodiversity research (Franklin 2010; Peterson et al. 2011; Anderson 2013; Guisan et al. 2017).
In fact, since the first algorithm for modeling species-environment relationship was presented (BIOCLIM, Nix 1986), the number of publications has increased dramatically (Lobo et al. 2010; Booth et al. 2013). A simple search in Google Scholar for the terms “ecological niche model” and “species distribution model” (last accessed on December 30, 2018) returned 2,950 and 6,400 citations, respectively, for 1990–2018 (Fig. 9.2a). Interestingly, the number of publications on these topics increased markedly in the past 10 years (Fig. 9.2, see also Lobo et al. 2010) and continues to grow, particularly in studies that emphasize the application of ENMs and SDMs to environmental assessment, forecasting, and hindcasting species distributions (Anderson 2013; Elith and Franklin 2013; Guisan et al. 2017). Interestingly, although the number of publications increased in the last 10 years, most of the studies were performed in United States and Europe (Fig. 9.2b) in countries with a high density of weather stations (Fig. 9.2c), with much less emphasis on the most diverse regions of the globe. The increasing access to species occurrence data (e.g., Global Biodiversity Information Facility, GBIF) and environmental data (climatic and satellite derived) has created the opportunity not only to model species-environment relationships but to expand the theoretical and practical applications of ENM and SDM to different research programs and fields, including conservation biology, wildlife and ecosystem management, evolutionary biology, and public health (Franklin 2010; Peterson et al. 2011; Guisan et al. 2017), and to do so in remote regions where access is limited and predictions of species distributions have disproportionate importance.
(a) Number of publications containing the term “species distribution model” or “ecological niche model” between 1990 and 2018 (Google Scholar search December 31, 2018). The solid line represents the combination of both SDM and ENM, while dashed and dotted lines indicate the individual terms. (b) Percentage of ENM/SDM studies performed in the United States and Europe in relation to the total number of publications from (a). (c) Distribution of weather stations (green dots) used to create the interpolated climate surfaces (i.e., WorldClim) and the number of species for the 30 most diverse countries. The numbers correspond to the estimated number of species—vertebrates and vascular plants—for each country. Notice that for most countries the weather stations are sparse and have low coverage. (Source: WorldClim: Global weather stations, 2014 (http://databasin.org/datasets/15a31dec689b4c958ee491ff30fcce75); biodiversity data: World Conservation Monitoring Centre of the United Nations Environment Programme (UNEP-WCMC), 2004)
Parallel to the development and evolution of ENM and SDM theory and applications, we have witnessed the growth of technological tools and S-RS products (Pettorelli et al. 2014a; Turner 2014). Many of these are particularly applicable for describing, quantifying, and mapping the spatial and temporal patterns of vegetation structure and function, the impacts of human activities, and environmental change (Turner et al. 2003; Pinto-Ledezma and Rivero 2014; Jetz et al. 2016; Cord et al. 2017) and more recently are used as predictors of broad patterns of biodiversity, including the associations between species co-occurrence patterns and ecosystem energy availability (Phillips et al. 2008; Pigot et al. 2016; Hobi et al. 2017). In addition, an unprecedented number of S-RS data and data products (S-RS) have been made freely available (Turner et al. 2003; Hobi et al. 2017) with the potential to track the spatial variation in the chemical composition of vegetation (Wang et al. 2019; Serbin and Townsend, Chap. 3), physiology, structure, and function (Lausch et al., Chap. 13; Serbin and Townsend, Chap. 3; Myneni et al. 2002; Saatchi et al. 2008; Jetz et al. 2016).
Despite the potential of S-RS products for measuring and modeling biodiversity (Gillespie et al. 2008; Pettorelli et al. 2014a, b; Turner 2014; Cord et al. 2013; Zimmermann et al. 2007), attention has only recently turned to using these data in studies of species-environment relationships (Cord et al. 2013; West et al. 2016), and most studies use bioclimatic data such as WorldClim (but see Paz et al., Chap. 11; Record et al., Chap. 10). Although early attempts indicated that S-RS products do not seem to improve the accuracy in estimating species distributions (Pearson et al. 2004; Thuiller 2004; Zimmermann et al. 2007), more recent publications (Kissling et al. 2012; Cord et al. 2013) suggest that despite these apparent limitations, S-RS products provide better spatial resolution that allow the discrimination of habitat characteristics not captured when bioclimatic data are used (Saatchi et al. 2008; Cord et al. 2013), and they can be used as surrogates of biotic and/or functional predictors such as LAI that increase the performance of individual species models (Kissling et al. 2012; Cord et al. 2013).
9.3 Modeling Ecological Niches and Predicting Geographic Distributions
Although the terms ENM and SDM are often used synonymously in the literature, the two are not equivalent (Anderson 2012; Soberón et al. 2017). A comprehensive discussion of this topic is beyond the scope of this chapter but is provided elsewhere (see Peterson et al. 2011; Anderson 2012; Soberón et al. 2017). A crucial step in differentiating the two terms is to establish a distinction between environmental space and geographical space (Hutchinson’s duality; Colwell and Rangel 2009). On the one hand, environmental space corresponds to a suite of environmental conditions at a given time (e.g., climate, topography); on the other hand, geographical space is the extent of a particular region or study area (Soberón and Nakamura 2009; Peterson et al. 2011) and includes important historical context. Thus, when modeling species ecological niches, we are modeling the existing abiotically suitable conditions for the species or the biotically reduced niche (Peterson et al. 2011; see also Fig. 9.1). However, when modeling species distributions, the intent is to project objects into geographical space (Fig. 9.1), and, depending on the factors considered, it is possible to estimate the occupied distributional area or the invadable distributional area (Soberón and Nakamura 2009; Peterson et al. 2011; Anderson 2012; Soberón et al. 2017).
9.3.1 Methods
9.3.1.1 Oak Species Data Sets
Occurrence data were downloaded from iDigBio between 20 and 24 July 2018, including localities collected by the authors, and cleaned for accuracy. Any botanical garden localities were discarded. All points were visually examined, and any localities that were outside the known range of the species, or in unrealistic locations (e.g., water bodies), were discarded.
9.3.1.2 Environmental Data Sets
For comparative purposes we obtained environmental data from two sources: (1) environmental variables derived from WorldClim and (2) S-RS data products (Fig. 9.3). Environmental variables derived from climatic data were obtained from the 10 to 2.5 arcmin WorldClim (Hijmans et al. 2005; spatial resolution of ~18.5 and 4.5 km at the equator, respectively) for annual mean temperature (BIO1), temperature seasonality (BIO4), minimum temperature of coldest month (BIO6), mean temperature of warmest quarter (BIO10), annual precipitation (BIO12), and precipitation seasonality (BIO15). These environmental variables were selected as critical for the distribution of oak species (Hipp et al. 2017) generally and were previously shown to be important in differentiating live oak (Virentes) species specifically (Cavender-Bares et al. 2011; Koehler et al. 2011; Cavender-Bares et al. 2015).
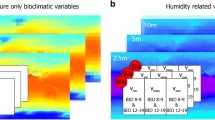
A selection of S-RS products and climatic variables used in this study. The panels show (a) Climate Hazards group Infrared Precipitation with Stations (CHIRPS); (b) MODIS mean LAI; (c) MODIS mean NDVI; (d) mean annual temperature; (e) mean annual precipitation; and (f) altitude or mean elevation from Shuttle Radar Topography Mission (SRTM)
Environmental variables from S-RS products were obtained from MODIS over a 15-year period (2001–2015) from NASA using the interface EOSDIS Earthdata (https://earthdata.nasa.gov). Data include two MODIS Collection 5 land products: LAI (8-day temporal resolution) and NDVI (16-day temporal resolution). LAI and NDVI products (Fig. 9.3b, c) are derived from Terra/Aqua MOD15A2 and Terra MOD13A2, respectively (see Myneni et al. 2002 for a detailed explanation of MODIS products). We also obtained precipitation data from Climate Hazards group Infrared Precipitation with Stations (CHIRPS , Fig. 9.3a), an S-RS product designed for monitoring drought and global environmental land change (Funk et al. 2015). Notice that the original MODIS products present a spatial resolution of 1 km and CHIRPS, a spatial resolution of 3 arcmin or ~5.5 km at the equator. To standardize the spatial resolution of both MODIS products and CHIRPS, we upscaled the spatial resolution of MODIS products to that of CHIRPS.
Prior to following the ENM/SDM procedures (outlined below), we calculated five new metrics taking advantage of the high temporal resolution LAI and NDVI data by doing simple arithmetic calculations: LAI/NDVI cumulative, LAI/NDVI mean, LAI/NDVI max, LAI/NDVI min, and LAI/NDVI seasonality or the coefficient of variation (see Saatchi et al. 2008; Hobi et al. 2017 for details). These metrics represent the spatial variation in vegetation productivity over a year (Berry et al. 2007; Hobi et al. 2017) and allow detection of biodiversity changes, description of habitats of different species, and tracking of phenology within species geographical ranges (Fig. 9.4). We used these two S-RS products given LAI and NDVI provide information on net primary productivity, dynamics of the growing season, and vegetation seasonality, all potentially important variables for characterizing plant species ranges (Myneni et al. 2002; Saatchi et al. 2008). All data processing and metric calculations were performed in R v3.5 (R Core Team 2018) using customized scripts and core functions from the packages raster (Hijmans 2018), gdalUtils (Greenberg and Mattiuzzi 2018), and rgdal (Bivand et al. 2018). R scripts for data processing and metric calculations can be found at https://github.com/jesusNPL/RS-SDM_ENM.
Satellite remotely sensed vegetation phenology based on MODIS LAI product. The time periods (t) represent the 46 time intervals every 8 days within a year starting from 1 January. The curves for vegetation phenology represent the variation in LAI over a 1-year interval calculated as the mean LAI within a species geographical distribution at 8-day intervals averaged over 15 years (see Hobi et al. 2017 for details). Shown is seasonal variation in the temperate forest vegetation where Q. virginiana occurs in North America compared with seasonal variation in the tropical dry forest vegetation where Q. oleoides occurs in Mexico and Central America
9.3.1.3 Modeling Procedure
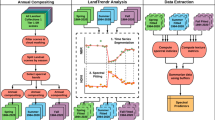
To model the ecological niche and distribution for oak species, we used an ensemble framework—prediction of a niche or a distributional area made by combining results of different modeling algorithms (Araújo and New 2007; Diniz-Filho et al. 2009). Within this framework we fit six species models and projected potential distributions for current environmental conditions for both environmental data sets (Table 9.1). The modeling algorithms included three statistical models (generalized linear models [GLM], generalized additive models [GAM], and adaptive regression splines [MARS]) and three machine learning models (MAXENT, support vector machines [SVM], and Random Forest [RF]). A description for each algorithm is detailed in Franklin (2010) (see also Peterson et al. 2011). All algorithms were fit in R and used the packages dismo (Hijmans et al. 2017), kernlab (Karatzoglou et al. 2004), randomForest (Liaw and Wiener 2002), mgcv (Wood 2006), and earth (Milborrow 2016).
Within our ensemble framework, species’ ecological niches are modeled using the six algorithms by fitting the occurrences of a single species and the predictors. The resulting six species models (one for each algorithm) are stacked into a single species model by averaging all models (Araújo and New 2007). We chose this approach because a major source of uncertainty in ENM/SDM arises from the algorithm used for modeling (Diniz-Filho et al. 2009; Qiao et al. 2015) and because the choice of the “best” modeling algorithm depends on the aims of the modeling applications (Peterson et al. 2011). Finally, using the stacked species models, we estimated macroecological patterns of species richness and the uncertainty associated with model parametrization. These patterns are less interesting in their own right for a small clade with only seven species, but they demonstrate an effective approach that can be applied to much larger groups of species.
We estimated live oak species richness by summing the projected potential species distributions; uncertainty was estimated as the variance attributable to the source of uncertainty (i.e., algorithms and their interactions) by performing a one-way analysis of variance (ANOVA) without replicates (Sokal and Rohlf 1995). The resulting uncertainty map shows regions with low and high uncertainty associated with the source of uncertainty (i.e., algorithm).
9.3.1.3.1 Statistical Analyses
To explore the performance of environmental data derived from RS for ENM/SDM compared to traditionally used environmental data from climatic variables (e.g., WorldClim), we evaluated the relationship between the modeled ecological niches from: (1) S-RS products; and (2) environmental variables from WorldClim. In doing so, we used correlation analyses corrected according Clifford’s method to obtain the effective degrees of freedom for Pearson’s coefficients while controlling for spatial autocorrelation (Clifford et al. 1989). Statistical analyses were performed in R using the package SpatialPack (Vallejos and Osorio 2014).
9.3.2 Results
Live oak models calibrated using different sources and combinations of environmental predictors (Table 9.1) within the ensemble framework generally provided similar suitability distributions (Fig. 9.5). Interestingly, increasing the number of predictors or increasing model complexity (S-SR2 in Table 9.1) affected model performance as measured by the Cohen’s Kappa coefficient and AUC (area under the receiver operating characteristic curve) indices (Table 9.2), and thus affected the geographic predictions: Complex models tended to have higher statistical performance but to underestimate the distributions of live oak species when compared with simpler models (Fig. 9.5). Individual live oak species models made from S-RS products and WorldClim differed somewhat in their performances (see Table 9.2 and Fig. 9.5). Models from WorldClim tended to have slightly better statistical performance in inferring species distributions based on the AUC and Kappa criteria. However, these metrics do not capture differences in the precision and spatial resolution of the approaches. In several species, the WorldClim models predicted low precision locations compared to the S-RS data. In particular, the IUCN (International Union for Conservation of Nature) red-listed narrow endemic Brandegee Oak (Quercus brandegeei) in southern Baja California is very imprecisely predicted compared with the S-RS data. Using high-resolution interpolated climatic predictors did not improve the performance of individual models (WC25 in Table 9.2) and returned similar suitability predictions to those estimated under lower spatial resolution climatic predictors (Fig. 9.5). Although WorldClim models seems to have better statistical performance as shown in Table 9.2, we can at most discriminate the accuracy of interpolating continuous surface-derived models, only when we are inferring habitat suitability models (ENM) and not the projected species geographical distribution (SDM). Using S-RS data as predictors not only helps to identify the species habitat suitability but also incorporates local ecological conditions necessary to predict local species distributions and co-occurrence (Radeloff et al. 2019). This is because S-RS data have the potential to get at biological mechanisms, for example, through the detection of species phenological variation over space and time (Figs. 9.3c and 9.4).
Maps of predicted distributions for seven live oak species under three combinations of environmental variables. Legend colors represent values of suitability, where 1 and 0 represent maximum and minimum suitability, respectively. (a–d) Quercus brandegeei; (e–h) Quercus fusiformis; (i–l) Quercus geminata; (LL–Ñ) Quercus minima; (o–r) Quercus oleoides; (s–v) Quercus sagraena; and (w–z) Quercus virginiana. Notice that each species model was estimated using an ensemble framework such that each species model represents the average of six algorithms weighted by their AUC. For representation purposes we cropped the predicted distributions using the species geographical ranges obtained from BIEN database (Enquist et al. 2016)
When macroecological patterns of species richness and uncertainty maps were constructed, we observed similar patterns of species richness between maps made from the simpler combination of S-RS and WorldClim models (Fig. 9.6a, c, and d; see Table 9.1 for a description of the environmental combinations of S-RS and WorldClim). Notably, species richness estimation from the complex S-RS tends to restrict live oak assemblages to southeastern North America (Fig. 9.6b), which is the only region where three live oak species co-occur. Uncertainty maps (Fig. 9.6e–h) show that uncertainty values for the simpler S-RS models (Fig. 9.6e) are lower when compared with the other models and WorldClim models tend to show high uncertainty values across south-central North America (e.g., Texas).
Macroecological patterns of species richness (top panel) and uncertainty (bottom panel) for live oak species quantified under four combinations of environmental variables. WorldClim data were used at two spatial resolutions, 10 and 2.5 arcmin. See Table 9.1 for a description of the environmental combinations. Numbers on legends for the top panel represent the number of species within each pixel, where 4 means that four live oak are co-occurring in those pixels. Numbers on legends on the bottom panel represent the percentage of uncertainty or variance between algorithms, where higher values represent higher uncertainty
The geographical relationship between the individual species models made for the four combinations of environmental predictors varied depending on the species evaluated (Table 9.3), although they show positive relationships in all cases. In general, spatial relationships between species models from S-RS products (i.e., S-RS vs S-RS2) and WorldClim (i.e., WC10 vs WC25) were strongly correlated (see also Paz et al., Chap. 11), while the relationship between models made from S-RS products and WorldClim data varied slightly; the simpler models made from the S-RS data showed stronger spatial relationship to the WorldClim models (Table 9.3). Interestingly, the spatial relationships increased as a function of increasing the species potential distributions. For example, weaker spatial relationships (r = 0.5384 for S-RS/WC10 and r = 0.6360 for S-RS/WC25) were found for Quercus brandegeei, and stronger spatial relationships were found for dwarf live oak (Quercus minima; r = 0.8150 for S-RS/WC10 and r = 0.8332) and southern live oak (Quercus virginiana; r = 0.8872 for S-RS/WC10 and r = 0.8523 for S-RS/WC25) that are distributed across the southeastern United States (Fig. 9.5LL–Ñ and Fig. 9.5W–Z, respectively).
Finally, we found that macroecological patterns of species richness derived from the four sets of environmental predictors were strongly correlated (Table 9.4). Although the algorithms used for modeling have been emphasized as a major source of uncertainty (Diniz-Filho et al. 2009; Qiao et al. 2015), by applying the ensemble framework, we found that uncertainties due to the modeling algorithm were low (<10%) for the four sets of environmental predictors and showed similar distribution estimates (Fig. 9.6e–h). These results suggest that our results are not biased by applying a particular algorithm. Interestingly, we found low correlations between uncertainty predictions under S-RS and WC comparisons (Table 9.4), which potentially could suggest an associated error due to the predictors used to build the models.
9.4 Perspectives
The field of ecological niche and species distribution modeling contributes significantly to our capacity to evaluate and describe the effect of geographical and environmental features on species distributions and has become in one of the most widely applied tools for the assessment of the impact of climate change and human activities on species and communities, biological invasions, epidemiology, and conservation biology (Peterson et al. 2011; Guisan et al. 2017). However, despite important advances in theory (Soberón 2007; Colwell and Rangel 2009; Soberón and Nakamura 2009; Peterson et al. 2011), methods, and algorithms (reviewed in Duarte et al. 2019; see also Warren et al. 2018) and practical applications (Guillera-Arroita et al. 2015; Cord et al. 2017; Sanín and Anderson 2018), most studies still rely on the use of interpolated climate data as environmental predictors (Saatchi et al. 2008; Waltari et al. 2014). In this article we compare the performance of environmental data derived from interpolated climate surfaces data (i.e., WorldClim) and S-RS products data (i.e., LAI and NDVI). Specifically, using live oaks as a case study, we show the advances and potential caveats in using S-RS data in describing and predicting species-environment relationships. Overall, our analyses show that S-RS products perform, as well as products from interpolated climate surfaces as environmental predictors (Tables 9.2, 9.3, and 9.4), and indeed present quite similar results for both species environmental suitability and macroecological patterns (Figs. 9.5 and 9.6), similar to Paz et al. (Chap. 11). However, they have the potential to provide more precise estimates of species distributions at higher spatial resolution.
In our example, we used different grain sizes for both data sets: WorldClim (10 and 2.5 arcmin or ~18.5 and ~ 4.5 km at the equator, respectively) and S-RS products (3 arcmin or ~5.5 km at the equator). Although changing grain size in the predictors might be anticipated to affect model performance, different lines of evidence indicate that model performance is not affected by grain resolution, but rather by species response to the environmental conditions in the study region (Guisan et al. 2007, see also Fig. 9.5). Our results show that enhancing spatial resolution of interpolated climatic data does not improve the spatial resolution at which species distributions can be accurately predicted. The quality of interpolated climate surfaces such as WorldClim, which depends on climatic stations as data sources, has been ignored as a source of uncertainty in studies of species-environment relationships—for example, Hijmans et al. (2005) used a variable number of weather stations for their interpolations, 47.554, 24.542, and 14.930 for precipitation, mean temperature, and maximum and minimum temperature, respectively—especially in the tropics (Fig. 9.2c), where weather stations are sparse (Hijmans et al. 2005; Soria-Auza et al. 2010). This source of uncertainty can be avoided using S-RS products, which have continuous (from daily to monthly) and quasi-global environmental information, including precipitation, temperature, and biophysical variables that represent different components of vegetation and ecosystems (Funk et al. 2015; Cord et al. 2017; Radeloff et al. 2019).
Fine and broad spatial and temporal scale data derived from S-RS, which have only been available in the last ~20 years (Turner 2014), can be used to improve the evaluation of species-environment relationships. A number of research avenues remain to be pursued to better understand the potential of S-RS data and their products in quantifying species ecological niches and estimating species distributions. For example, applying the same framework presented here to other species or clades (including vertebrates and invertebrates) or applying more complex frameworks (e.g., Peterson and Nakazawa 2008; Waltari et al. 2014) may shed light on the potential of S-RS products as predictors for the analysis of species-environment relationships. This is important because ENMs/SDMs are used as predictive models that can be extrapolated across space and time to forecast and monitor biodiversity under a changing global climate (Peterson and Nakazawa 2008; Warren 2012).
9.4.1 Should We Use S-RS Data for ENM/SDM?
Whether S-RS data should replace other environmental data in modeling niches and projecting species distribution depends on the modeling purposes (Peterson et al. 2011). In fact, modeling species niches and projecting distributions involves relating a set of species occurrences to relevant environmental predictors. In essence, ENM/SDM based only on climatic variables would tend to return broad predictions (Coudun et al. 2006, see also right panel in Fig. 9.5), particularly because climatic data are useful in describing macroecological patterns of species distributions and communities (Lin and Wiens 2017; Manzoor et al. 2018), while ENM/SDM based on S-RS data alone allows the discrimination of local features not captured by climatic information (Coudun et al. 2006; Saatchi et al. 2008; Radeloff et al. 2019). Nevertheless, it seems that overall statistical model accuracy in this example is not improved (Pearson et al. 2004; Thuiller 2004; see also Table 9.2). Given that climatic and S-RS data provide information at different spatial and temporal scales, a promising option would be to use both sources of environmental predictors to model species distributions to achieve “the best of both worlds” (Saatchi et al. 2008; Pradervand et al. 2013).
More accurate predictions of species distributions are critical for the development of conservation and management actions if we are to meet the challenges posed by global change (Coudun et al. 2006; Zimmermann et al. 2007; Cord et al. 2013). Our point here is to facilitate and demonstrate the potential for the use of S-RS data for predicting species distributions and modeling environmental niches. The results we show here and those of others (e.g., Saatchi et al. 2008; Waltari et al. 2014) indicate that S-RS data provide a valuable complement to other environmental variables for ENM/SDM.
Another potential and important research direction is the use of S-RS products that have high temporal resolution, such as LAI (Fig. 9.3b), as biophysical variables that represent ecosystem functions (Cord et al. 2013, 2017). These products allow the exploration of dynamics of vegetation growth and seasonality in vegetation function, fundamental features that characterize vegetation form and function (Myneni et al. 2002; Hobi et al. 2017). Here, using metrics derived from MODIS LAI in combination with other S-RS products (Table 9.1), we show that, using relevant biophysical variables, it is possible to predict distributions similar to those predicted from climate data alone (Fig. 9.5, Tables 9.2 and 9.3). In fact, a recent study (Simões and Peterson 2018) found that including biotic predictors can improve ENMs even while increasing model complexity, such that the combination of abiotic and biotic predictors improves model performance (Simões and Peterson 2018). To confirm this conclusion, substantial effort would be needed, including new methodological and conceptual approaches, to disentangle the real contribution of S-RS products—spatial and temporal features of S-RS products that improve statistical model performance—as predictors of species distributions. Nonetheless, our results highlight an advance on the use of relevant predictors for modeling species-environment relationships.
In addition, recent macroecological studies have used these products to relate annual vegetation productivity to continental and global patterns of species richness (Pigot et al. 2016; Hobi et al. 2017; Coops et al. 2018), providing spatially explicit support for the use of satellite data products in predicting biodiversity. These advances point to an exciting avenue for the study of the distribution and assembly of biological communities (Ferrier and Guisan 2006). For example, S-RS products can be used for the development of stacked species distribution models (S-SDM, see Fig. 9.6) that can be integrated into novel biodiversity modeling frameworks, such as Spatially Explicit Species Assemblage Modelling (SESAM, Guisan and Rahbek 2011) or the Hierarchical Modelling of Species Communities (HMSC, Ovaskainen et al. 2017), aimed at predicting composition and distribution of species and communities (Mateo et al. 2017).
Finally, the example presented here is meant to spur further theoretical, methodological, and empirical research aimed at developing a Global Biodiversity Observatory (Geller et al., Chap. 20). Explicit incorporation of biotic information into species-environment modeling may turn our focus away from the use only of climatic information toward the “complete” evaluation of the drivers that determine the species distributions (Fig. 9.1).
9.4.2 Enabling Large-Scale Biodiversity Change Detection
Since the last millennium, rising human population and activity have been major drivers of environmental change on Earth, with consequences for the distribution and abundance of biodiversity and associated ecosystem functioning (Tilman 1997; Tylianakis et al. 2008). Thus, improving large-scale biodiversity change detection is crucial to the development of effective policies that advance conservation and management of species and communities.
Such efforts are critical to enhancing efforts to develop a Global Biodiversity Observatory (Geller et al., Chap. 20; Jetz et al. 2016). Research interest in using S-RS has increased in recent years given its high potential for monitoring global biodiversity and detecting change (Turner 2014; Jetz et al. 2016). For example, it is possible to identify shifts in vegetation structure or to monitor the dynamics of the growing season of an entire region, or within a specific species geographical range (Fig. 9.4) using time series S-RS products such as LAI—half of the total green leaf area per unit of horizontal ground surface area (Xiao et al. 2014)—which has a temporal resolution of 8 days. This is particularly important given that ENM/SDM theory assumes that species’ niches are stable across time and space and that species and their environments are at pseudo-equilibrium, suggesting that species are occupying all suitable areas (Guisan and Thuiller 2005). However, the environment is dynamic and can change even at small scales; species ranges can thus expand and retract across time, varying within species lifetimes as well as over evolutionary timescales encompassing many generations. Long-term series of S-RS data products (i.e., spatial and temporal) supply remarkable opportunities for assessing and monitoring the state of the Earth’s surface and, combining with species-environment relationship modeling, provide new frontiers for the prediction of species distributions and species monitoring across time and space (Randin et al. 2020). Indeed, using biophysical variables derived from high-resolution S-RS products (i.e., LAI) allows the identification of geographic areas where species actually occur (Fig. 9.7) and thus has the potential for enhancing the predictions of a set of species that could occur in an area—species pool—that is used for species assignments from direct RS detection using hyperspectral data (see simulation in Fig. 7.8, Section 9.4.2 in Meireles et al., Chap. 7).
Mean LAI estimated at 8-day intervals averaged over 15 years (left panel). The overlaid continuous lines correspond to the geographical ranges of Q. virginiana and Q. oleoides obtained from BIEN database. Predicted distributions for Q. virginiana (top right panel) and Q. oleoides (bottom right panel) are based on S-RS products, which include the temporal variation in LAI shown in A. The triangles over the maps represent occurrence points used for calibration (where the authors have collected specimens), and the boxes represent a zoom over a specific area of the predicted species distributions. Note that high values of the predicted distributions coincide with the occurrence points
In addition, enhancing predictive models of the species expected to be present in a given geographic region can be coupled with other means of detecting which species are present based on spectroscopic imaging (Serbin et al. 2015; Bolch et al., Chap. 12; Meireles et al., Chap. 7), reducing the complexity of species identification algorithms. For example, imaging spectroscopy allows mapping of functional traits, by estimating vegetation traits for each pixel in an image (Wang et al. 2019; Asner et al. 2017; Martin, Chap. 5). Plant spectra obtained from imaging spectroscopy at different spatial resolutions can in turn be used to detect different aspects and traits—within- and between-species differences in morphology, foliar chemistry, life history strategies—of plant species (Ustin and Gamon 2010; Cavender-Bares et al. 2017; Schweiger et al. 2018) and the correct identification of different taxonomic levels from populations to species to clades (Cavender-Bares et al. 2016). Thus, the integration of spectral approaches with techniques for modeling species ecological niches has the potential to produce reliable information of species distributions and co-occurrence, filling current gaps about species-environment relationships at a range of spatial scales and levels of organization—from species to communities—increasing the accuracy of direct detection assignments, and enabling monitoring of changes in biodiversity, one of the premises for the sustainable management of the biosphere (Pinto-Ledezma and Rivero 2014; Fernández et al., Chap. 18).
References
Anderson RP (2012) Harnessing the world’s biodiversity data: promise and peril in ecological niche modeling of species distributions. Ann N Y Acad Sci 1260(1):66–80. https://doi.org/10.1111/j.1749-6632.2011.06440.x
Anderson RP (2013) A framework for using niche models to estimate impacts of climate change on species distributions. Ann N Y Acad Sci 1297(1):8–28. https://doi.org/10.1111/nyas.12264
Araújo MB, New M (2007) Ensemble forecasting of species distributions. Trends Ecol Evol 22(1):42–47. https://doi.org/10.1016/j.tree.2006.09.010
Asner GP, Martin RE, Knapp DE et al (2017) Airborne laser-guided imaging spectroscopy to map forest trait diversity and guide conservation. Science 355(6323):385–389. https://doi.org/10.1126/science.aaj1987
Berry S, Mackey B, Brown T (2007) Potential applications of remotely sensed vegetation greenness to habitat analysis and the conservation of dispersive fauna. Pacific Cons Bio 13(2):120. https://doi.org/10.1071/pc070120
Bivand R, Keitt T, Rowlingson B (2018) rgdal: bindings for the ‘geospatial’ data abstraction library. R package version 1.3-4. https://CRAN.R-project.org/package=rgdal.
Booth TH, Nix HA, Busby JR et al (2013) Bioclim: the first species studies. Divers Distrib 20(1):1–9. https://doi.org/10.1111/ddi.12144
Cavender-Bares J, Gonzalez-Rodriguez A, Pahlich A et al (2011) Phylogeography and climatic niche evolution in live oaks (Quercus series Virentes) from the tropics to the temperate zone. J Biogeogr 38(5):962–981. https://doi.org/10.1111/j.1365-2699.2010.02451.x
Cavender-Bares J, Gonzalez-Rodriguez A, Eaton DA et al (2015) Phylogeny and biogeography of the American live oaks (Quercus subsection Virentes): a genomic and population genetics approach. Mol Ecol 24(14):3668–3687. https://doi.org/10.1111/mec.13269
Cavender-Bares J, Meireles JE, Couture J et al (2016) Associations of leaf spectra with genetic and phylogenetic variation in oaks: prospects for remote detection of biodiversity. Remote Sens 8(3):221. https://doi.org/10.3390/rs8030221
Cavender-Bares J, Gamon JA, Hobbie SE et al (2017) Harnessing plant spectra to integrate the biodiversity sciences across biological and spatial scales. Am J Bot 104(7):966–969. https://doi.org/10.3732/ajb.1700061
Clifford P, Richardson S, Hemon D (1989) Assessing the significance of the correlation between two spatial processes. Biometrics 45(1):123–134. https://doi.org/10.2307/2532039
Colwell RK, Rangel TF (2009) Hutchinson’s duality: the once and future niche. Proc Natl Acad Sci U S A 106(Supplement_2):19651–19658. https://doi.org/10.1073/pnas.0901650106
Coops NC, Kearney SP, Bolton DK et al (2018) Remotely-sensed productivity clusters capture global biodiversity patterns. Sci Rep 8(1):16261. https://doi.org/10.1038/s41598-018-34162-8
Cord AF, Meentemeyer RK, Leitão PJ et al (2013) Modelling species distributions with remote sensing data: bridging disciplinary perspectives. J Biogeogr 40(12):2226–2227. https://doi.org/10.1111/jbi.12199
Cord AF, Brauman KA, Chaplin-Kramer R et al (2017) Priorities to advance monitoring of ecosystem services using earth observation. Trends Ecol Evol 32(6):416–428. https://doi.org/10.1016/j.tree.2017.03.003
Coudun C, Gégout JC, Piedallu C et al (2006) Soil nutritional factors improve models of plant species distribution: an illustration with Acer campestre (L.) in France. J Biogeogr 33(10):1750–1763. https://doi.org/10.1111/j.1365-2699.2005.01443.x
Diniz-Filho JAF, Bini L, Rangel TF et al (2009) Partitioning and mapping uncertainties in ensembles of forecasts of species turnover under climate change. Ecography 32(6):897–906. https://doi.org/10.1111/j.1600-0587.2009.06196.x
Duarte A, Whitlock SL, Peterson JT (2019) Species distribution modeling. In: Encyclopedia of Ecology, pp 189–198. https://doi.org/10.1016/b978-0-12-409548-9.10572-x
Elith J, Franklin J (2013) Species distribution modeling. In: Encyclopedia of biodiversity, pp 692–705. https://doi.org/10.1016/b978-0-12-384719-5.00318-x
Elton C (1927) Animal Ecology. Sedgwick and Jackson, London
Enquist BJ, Condit R, Peet RK et al (2016) Cyberinfrastructure for an integrated botanical information network to investigate the ecological impacts of global climate change on plant biodiversity. PeerJ Preprints 4:e2615v2. https://doi.org/10.7287/peerj.preprints.2615v2
Ferrier S, Guisan A (2006) Spatial modelling of biodiversity at the community level. J Appl Ecol 43(3):393–404. https://doi.org/10.1111/j.1365-2664.2006.01149.x
Franklin J (2010) Mapping species distribution: spatial inference and prediction. Cambridge University Press, Cambridge
Funk C, Peterson P, Landsfeld M et al (2015) The climate hazards infrared precipitation with stations—a new environmental record for monitoring extremes. Sci Data 2:150066. https://doi.org/10.1038/sdata.2015.66
Gaston KJ (2009) Geographic range limits: achieving synthesis. Proc R Soc Lond B Biol Sci 276(1661):1395–1406. https://doi.org/10.1098/rspb.2008.1480
Gillespie TW, Foody GM, Rocchini D et al (2008) Measuring and modelling biodiversity from space. Prog Phys Geogr 32(2):203–221. https://doi.org/10.1177/0309133308093606
Greenberg JA, Mattiuzzi M (2018) gdalUtils: wrappers for the geospatial data abstraction library (GDAL) utilities. R package version 2.0.1.14. https://CRAN.R-project.org/package=gdalUtils.
Grinnell J (1904) The Origin and Distribution of the Chest-Nut-Backed Chickadee. The Auk, 21(3), 364–382. https://doi.org/10.2307/4070199
Grinnell J (1917) The Niche-Relationships of the California Thrasher. The Auk, 34(4), 427–433. https://doi.org/10.2307/4072271
Guillera-Arroita G, Lahoz-Monfort JJ, Elith J et al (2015) Is my species distribution model fit for purpose? Matching data and models to applications. Glob Ecol Biogeogr 24(3):276–292. https://doi.org/10.1111/geb.12268
Guisan A, Thuiller W (2005) Predicting species distribution: offering more than simple habitat models. Ecol Lett 8(9):993–1009. https://doi.org/10.1111/j.1461-0248.2005.00792.x
Guisan A, Graham CH, Elith J et al (2007) Sensitivity of predictive species distribution models to change in grain size. Divers Distrib 13(3):332–340. https://doi.org/10.1111/j.1472-4642.2007.00342.x
Guisan A, Rahbek C (2011) SESAM - a new framework integrating macroecological and species distribution models for predicting spatio-temporal patterns of species assemblages. Journal of Biogeography, 38(8), 1433–1444. https://doi.org/10.1111/j.1365-2699.2011.02550.x
Guisan A, Thuiller W, Zimmermann NE (2017) Habitat suitability and distribution models. Cambridge University Press, Cambridge. https://doi.org/10.1017/9781139028271
Hijmans RJ, Phillips S, Leathwick J et al (2017) dismo: species distribution modeling. R package version 1.1-4. https://CRAN.R-project.org/package=dismo.
Hijmans RJ (2018) raster: Geographic Data Analysis and Modeling. R package version 3.0-7. https://CRAN.R-project.org/package=raster
Hijmans RJ, Cameron SE, Parra JL et al (2005) Very high resolution interpolated climate surfaces for global land areas. Int J Climatol 25(15):1965–1978. https://doi.org/10.1002/joc.1276
Hipp AL, Manos PS, González-Rodríguez A et al (2017) Sympatric parallel diversification of major oak clades in the Americas and the origins of Mexican species diversity. New Phytol 217(1):439–452. https://doi.org/10.1111/nph.14773
Hobi ML, Dubinin M, Graham CH et al (2017) A comparison of dynamic habitat indices derived from different MODIS products as predictors of avian species richness. Remote Sens Environ 195:142–152. https://doi.org/10.1016/j.rse.2017.04.018
Holdridge LR (1947) Determination of world plant formations from simple climatic data. Science 105(2727):367–368. https://doi.org/10.1126/science.105.2727.367
Hutchinson GE (1957) Concluding Remarks. Cold Spring Harbor Symposia on Quantitative Biology, 22(0), 415–427. https://doi.org/10.1101/sqb.1957.022.01.03
Jetz W, Cavender-Bares J, Pavlick R et al (2016) Monitoring plant functional diversity from space. Nat Plants 2(3):16024. https://doi.org/10.1038/nplants.2016.24
Karatzoglou A, Smola A, Hornik K et al (2004) Kernlab – an S4 package for Kernel methods in R. J Stat Softw 11(9):1–20. http://www.jstatsoft.org/v11/i09/.
Kissling WD, Dormann CF, Groeneveld J et al (2012) Towards novel approaches to modelling biotic interactions in multispecies assemblages at large spatial extents. J Biogeogr 39(12):2163–2178. https://doi.org/10.1111/j.1365-2699.2011.02663.x
Koehler K, Center A, Cavender-Bares J (2011) Evidence for a freezing tolerance-growth rate trade-off in the live oaks (Quercus series Virentes) across the tropical-temperate divide. New Phytol 193(3):730–744. https://doi.org/10.1111/j.1469-8137.2011.03992.x
Liaw A, Wiener M (2002) Classification and regression by randomForest. R News 2:18–22
Lin LH, Wiens JJ (2017) Comparing macroecological patterns across continents: evolution of climatic niche breadth in varanid lizards. Ecography 40(8):960–970. https://doi.org/10.1111/ecog.02343
Lobo JM, Jiménez-Valverde A, Hortal J (2010) The uncertain nature of absences and their importance in species distribution modelling. Ecography 33(1):103–114. https://doi.org/10.1111/j.1600-0587.2009.06039.x
Manzoor SA, Griffiths G, Lukac M (2018) Species distribution model transferability and model grain size – finer may not always be better. Sci Rep 8(1):7168. https://doi.org/10.1038/s41598-018-25437-1
Mateo RG, Mokany K, Guisan A (2017) Biodiversity models: what if unsaturation is the rule? Trends Ecol Evol 32(8):556–566. https://doi.org/10.1016/j.tree.2017.05.003
Milborrow S (2016) Earth: multivariate adaptive regression splines. R package version 4.4.4. Retrieved from https://CRAN.R-project.org/package=earth.
Myneni R, Hoffman S, Knyazikhin Y et al (2002) Global products of vegetation leaf area and fraction absorbed PAR from year one of MODIS data. Remote Sens Environ 83(1–2):214–231. https://doi.org/10.1016/s0034-4257(02)00074-3
Nix HA (1986) A biogeographic analysis of Australian elapid snakes. In: Longmore R (ed) Atlas of elapid snakes of Australia: Australian Flora and Fauna Series, vol 7. Bureau of Flora and Fauna, Canberra, pp 4–15
Ovaskainen O, Tikhonov G, Norberg A et al (2017) How to make more out of community data? A conceptual framework and its implementation as models and software. Ecol Lett 20(5):561–576. https://doi.org/10.1111/ele.12757
Pearson RG, Dawson TP, Liu C (2004) Modelling species distributions in Britain: a hierarchical integration of climate and land-cover data. Ecography 27(3):285–298. https://doi.org/10.1111/j.0906-7590.2004.03740.x
Peterson AT, Martínez-Meyer E, Soberón J et al (2011) Ecological niches and geographic distributions. Monographs in population biology, vol 49. Princeton University Press, Princeton
Peterson AT, Nakazawa Y (2008) Environmental data sets matter in ecological niche modelling: an example with Solenopsis invicta and Solenopsis richteri. Glob Ecol Biogeogr 17(1):135–144. https://doi.org/10.1111/j.1466-8238.2007.00347.x
Pettorelli N, Safi K, Turner W (2014a) Satellite remote sensing, biodiversity research and conservation of the future. Philos Trans R Soc Lond Ser B Biol Sci 369(1643):20130190–20130190. https://doi.org/10.1098/rstb.2013.0190
Pettorelli N, Laurance WF, O’Brien TG et al (2014b) Satellite remote sensing for applied ecologists: opportunities and challenges. J Appl Ecol 51(4):839–848. https://doi.org/10.1111/1365-2664.12261
Phillips LB, Hansen AJ, Flather CH (2008) Evaluating the species energy relationship with the newest measures of ecosystem energy: NDVI versus MODIS primary production. Remote Sens Environ 112(9):3538–3549. https://doi.org/10.1016/j.rse.2008.04.012
Pigot AL, Tobias JA, Jetz W (2016) Energetic constraints on species coexistence in birds. PLoS Biol 14(3):e1002407. https://doi.org/10.1371/journal.pbio.1002407
Pinto-Ledezma JN, Rivero ML (2014) Temporal patterns of deforestation and fragmentation in lowland Bolivia: implications for climate change. Clim Chang 127(1):43–54. https://doi.org/10.1007/s10584-013-0817-1
Pradervand JN, Dubuis A, Pellissier L et al (2013) Very high resolution environmental predictors in species distribution models. Prog Phys Geogr 38(1):79–96. https://doi.org/10.1177/0309133313512667
Qiao H, Soberón J, Peterson AT (2015) No silver bullets in correlative ecological niche modelling: insights from testing among many potential algorithms for niche estimation. Methods Ecol Evol 6(10):1126–1136. https://doi.org/10.1111/2041-210x.12397
R Core Team (2018) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Radeloff VC, Dubinin M, Coops NC et al (2019) The dynamic habitat indices (DHIs) from MODIS and global biodiversity. Remote Sens Environ 222:204–214. https://doi.org/10.1016/j.rse.2018.12.009
Randin CF, Ashcroft MB, Bolliger J et al. (2020) Monitoring biodiversity in the Anthropocene using remote sensing in species distribution models. Remote Sens Environ 239:111626 https://doi.org/10.1016/j.rse.2019.111626
Smith AB, Godsoe W, Rodríguez-Sánchez F et al (2018) Niche estimation above and below the species level. Trends Ecol Evol 34:260. https://doi.org/10.1016/j.tree.2018.10.012
Saatchi S, Buermann W, ter Steege H et al (2008) Modeling distribution of Amazonian tree species and diversity using remote sensing measurements. Remote Sens Environ 112(5):2000–2017. https://doi.org/10.1016/j.rse.2008.01.008
Sanín C, Anderson RP (2018) A framework for simultaneous tests of abiotic, biotic, and historical drivers of species distributions: empirical tests for north American Wood warblers based on climate and pollen. Am Nat 192(2):E48–E61. https://doi.org/10.1086/697537
Serbin SP, Singh A, Desai AR et al (2015) Remotely estimating photosynthetic capacity, and its response to temperature, in vegetation canopies using imaging spectroscopy. Remote Sens Environ 167:78–87. https://doi.org/10.1016/j.rse.2015.05.024
Simões MVP, Peterson AT (2018) Importance of biotic predictors in estimation of potential invasive areas: the example of the tortoise beetle Eurypedus nigrosignatus, in Hispaniola. PeerJ 6:e6052. https://doi.org/10.7717/peerj.6052
Soberón J (2007) Grinnellian and Eltonian niches and geographic distributions of species. Ecol Lett 10(12):1115–1123. https://doi.org/10.1111/j.1461-0248.2007.01107.x
Soberón J, Nakamura M (2009) Niches and distributional areas: concepts, methods, and assumptions. Proc Natl Acad Sci U S A 106(Supplement_2):19644–19650. https://doi.org/10.1073/pnas.0901637106
Soberón J, Peterson AT (2005) Interpretation of models of fundamental ecological niches and species’ distributional areas. Biodivers Inform 2. https://doi.org/10.17161/bi.v2i0.4
Soberón J, Osorio-Olvera L, Peterson AT (2017) Diferencias conceptuales entre modelación de nichos y modelación de áreas de distribución. Rev Mex Biodivers 88(2):437–441. https://doi.org/10.1016/j.rmb.2017.03.011
Sokal RR, Rohlf FJ (1995) Biometry: the principles and practice of statistics in biological research, 3rd edn. W. H. Freeman and Co., New York
Soria-Auza RW, Kessler M, Bach K et al (2010) Impact of the quality of climate models for modelling species occurrences in countries with poor climatic documentation: a case study from Bolivia. Ecol Model 221(8):1221–1229. https://doi.org/10.1016/j.ecolmodel.2010.01.004
Schweiger AK, Cavender-Bares J, Townsend PA et al (2018) Plant spectral diversity integrates functional and phylogenetic components of biodiversity and predicts ecosystem function. Nat Ecol Evol 2(6):976–982. https://doi.org/10.1038/s41559-018-0551-1
Thuiller W (2004) Patterns and uncertainties of species’ range shifts under climate change. Glob Chang Biol 10(12):2020–2027. https://doi.org/10.1111/j.1365-2486.2004.00859.x
Tilman D (1997) The influence of functional diversity and composition on ecosystem processes. Science 277(5330):1300–1302. https://doi.org/10.1126/science.277.5330.1300
Turner W (2014) Sensing biodiversity. Science 346(6207):301–302. https://doi.org/10.1126/science.1256014
Turner W, Spector S, Gardiner N et al (2003) Remote sensing for biodiversity science and conservation. Trends Ecol Evol 18(6):306–314. https://doi.org/10.1016/s0169-5347(03)00070-3
Tylianakis JM, Didham RK, Bascompte J et al (2008) Global change and species interactions in terrestrial ecosystems. Ecol Lett 11(12):1351–1363. https://doi.org/10.1111/j.1461-0248.2008.01250.x
Ustin SL, Gamon JA (2010) Remote sensing of plant functional types. New Phytol 186(4):795–816. https://doi.org/10.1111/j.1469-8137.2010.03284.x
Vallejos R, Osorio F (2014) Effective sample size of spatial process models. Spat Stat 9:66–92. https://doi.org/10.1016/j.spasta.2014.03.003
Wallace AR (1860) On the zoological geography of the Malay archipelago. Zool J Linnean Soc 4(16):172–184. https://doi.org/10.1111/j.1096-3642.1860.tb00090.x
Waltari E, Schroeder R, McDonald K et al (2014) Bioclimatic variables derived from remote sensing: assessment and application for species distribution modelling. Methods Ecol Evol 5(10):1033–1042. https://doi.org/10.1111/2041-210x.12264
Wang Z, Townsend PA, Schweiger AK et al (2019) Mapping foliar functional traits and their uncertainties across three years in a grassland experiment. Remote Sens Environ 221:405–416. https://doi.org/10.1016/j.rse.2018.11.016
Warren DL (2012) In defense of “niche modeling”. Trends Ecol Evol 27(9):497–500. https://doi.org/10.1016/j.tree.2012.03.010
Warren DL, Beaumont LJ, Dinnage R et al (2018) New methods for measuring ENM breadth and overlap in environmental space. Ecography 42:444. https://doi.org/10.1111/ecog.03900
West AM, Evangelista PH, Jarnevich CS et al (2016) Integrating remote sensing with species distribution models; mapping tamarisk invasions using the software for assisted habitat modeling (SAHM). J Vis Exp (116). https://doi.org/10.3791/54578
Wood SN (2006) Generalized additive models. Chapman and Hall/CRC, Boca Raton
Xiao Z, Liang S, Wang J et al (2014) Use of general regression neural networks for generating the GLASS leaf area index product from time-series MODIS surface reflectance. IEEE Trans Geosci Remote Sens 52(1):209–223. https://doi.org/10.1109/tgrs.2013.2237780
Zimmermann NE, Edwards TC, Moisen GG et al (2007) Remote sensing-based predictors improve distribution models of rare, early successional and broadleaf tree species in Utah. J Appl Ecol 44(5):1057–1067. https://doi.org/10.1111/j.1365-2664.2007.01348.x
Acknowledgments
We thank Julián Velasco for his advice with ENM/SDM analyses. J.N.P-L. was supported by the University of Minnesota College of Biological Sciences’ Grand Challenges in Biology Postdoctoral Program. We acknowledge the bioDISCOVERY community at the University of Zürich for important discussions and the NSF Research Coordination Network “Biodiversity Across Scales.”
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2020 The Author(s)
About this chapter
Cite this chapter
Pinto-Ledezma, J.N., Cavender-Bares, J. (2020). Using Remote Sensing for Modeling and Monitoring Species Distributions. In: Cavender-Bares, J., Gamon, J.A., Townsend, P.A. (eds) Remote Sensing of Plant Biodiversity. Springer, Cham. https://doi.org/10.1007/978-3-030-33157-3_9
Download citation
DOI: https://doi.org/10.1007/978-3-030-33157-3_9
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-33156-6
Online ISBN: 978-3-030-33157-3
eBook Packages: Earth and Environmental ScienceEarth and Environmental Science (R0)