Abstract
This chapter focuses on planning field campaigns and data collection relevant to plant biodiversity. Particular emphasis is placed on sampling spectra of plants across scales, from the leaf to the canopy and airborne level, considering the issue of matching ecological data with spectra. The importance of planning is highlighted from the perspective of the long-term sustainability of a project, which includes using and contributing to the development of standards for project documentation and archiving. These issues are critical to biodiversity researchers involved in data collection in situ and via remote sensing (RS).
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
15.1 Introduction
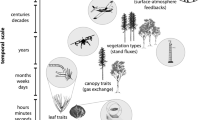
Field spectrometry is the measurement of spectral properties of (Earth) surface features in the environment (McCoy 2005) and is particularly relevant to plant biodiversity detection. Generally, molecular composition and arrangement and scattering properties of the measured media influence the spectral response (Goetz et al. 1985). Spectral characteristics of plants depend on chemical, structural, anatomical, and morphological leaf characteristics and whole plant traits, including plant height and shape, canopy architecture, branching structure, and the distribution of foliage within canopies (Cavender-Bares et al. 2017; Ustin and Gamon 2010; Serbin and Townsend, Chap. 3; Ustin and Jacquemoud, Chap. 14). Plant spectra provide a wealth of information about how plants use nutrients, light, and water; how these resources are shared within plant communities; and how patterns of resource use influence ecosystem functions and processes, including nutrient and water cycling and the provisioning of resources and habitat for other trophic levels. Spectroscopy of vegetation and plant biodiversity is part of the larger field of biophysics, which uses theories and methods of physics, such as optics, to understand biological systems.
For a long time, field campaigns were regarded as relatively unimportant for remote sensing (RS) studies. Using the term “ground data” or, more generally, “surface reference data” instead of “ground truth” has been suggested, since the latter implies that field data can be easily collected and are relatively “error-free” (Justice and Townshend 1981). In RS studies, ground data still are mainly collected for accuracy assessments or the validation of map products. Most often, land cover or vegetation maps are produced through image interpretation (Bartholomé and Belward 2005; Bicheron et al. 2008). Although the importance of accuracy assessments has been pointed out in the literature (Stehman 2001; Justice and Townshend 1981; Johannsen and Daughtry 2009), validation of map products usually means using high-resolution RS images to validate coarser resolution maps (Congalton et al. 2014). Map validations are often based on agreement among random points (Stehman 2001)—i.e., the extent to which land cover classes that random points fall into match the investigator’s interpretation of land cover visible in an image. While this procedure makes sense for global map products distinguishing few vegetation classes, local map products clearly benefit from the collection of ground data for validation. However, the importance of ground reference data goes far beyond map accuracy assessments. In fact, ground reference data are essential for remote sensing of plant biodiversity. Collecting ground reference data during spectral field campaigns provides a great opportunity to bridge the gap between RS science and ecology, two fields that are uniquely positioned to together develop methods to assess biodiversity across large spatial scales, continuously and in a detailed way. These assessments are needed to provide information about the current status of ecosystems; to predict the distribution of biodiversity, ecosystem function, and ecosystem processes into the future; and to counteract detrimental changes in ecosystems associated with global change. One reason to advocate for field campaigns is that remotely sensed images, which provide information pixel by pixel, always obscure part of the information on the ground, with the amount of hidden information depending on pixel size (Atkinson 1999). In order to understand the information provided by remotely sensed images of vegetation, it is critical to study the spectral characteristics of plants, their links to plant traits, and their influence on ecosystem properties at the sub-pixel level, because spectral variation is progressively lost when spectra of individual plants and non-vegetation features blend together at increasing spatial resolutions (Atkinson 1999).
This chapter deals mainly with planning field work and the collection of vegetation spectra with field spectrometers on the ground, which can subsequently be linked to other ecological data and/or RS data to investigate biological phenomena. Data collection for airborne spectroscopy is discussed as well, while other RS methods such as unmanned aerial systems (UASs), towers, and trams are covered in more detail in Gamon et al. (Chap. 16). Focus is also placed on data organization and management, particularly because these aspects of planning tend to receive less attention than, e.g., planning of sample collection, yet they are critical to a successful field campaign.
This chapter was written in full awareness that “good practices” are ever-evolving. The relative importance of, and acquisition methods for, ground data, including ecological data, depends on the research question, on the project goals, as well as on study scale, spectroscopic methods and RS data used, budget, time, site accessibility, and the personnel and their training (Justice and Townshend 1981). Likewise, spectral processing standards evolve and software goes out of date quickly. The examples included in this chapter are intended to illustrate what worked in particular situations and to point out pitfalls to avoid; many other approaches are as valuable. Flexibility (knowledge about different techniques and tools, a plan B, etc.) is important for adjusting to particular circumstances and challenges. The “best practice” is probably to learn about several “good practices”; read protocols; talk to field ecologists, data administrators, geographic information system (GIS) professionals, programmers, and communication experts; and get some hands-on experience. A selection of excellent protocols is available from Australia’s Terrestrial Ecosystem Research Network (TERN , http://www.auscover.org.au/wp-content/uploads/AusCover-Good-Practice-Guidelines_web.pdf), the Field Spectroscopy Facility at UK’s Natural Environment Research Council (NERC , http://fsf.nerc.ac.uk/resources/guides/), the US National Ecological Observatory Network (NEON, http://data.neonscience.org/documents) , the Global Airborne Observatory (GAO, https://gao.asu.edu/spectranomics), and the Canadian Airborne Biodiversity Observatory (CABO, http://www.caboscience.org), among others. For more in-depth coverage of particular topics, see texts on the general principles of RS (e.g., Warner et al. 2009) and RS of vegetation (e.g., Jones and Vaughan 2010; Thenkabail et al. 2012), field methods in RS (e.g., McCoy 2005), spatial statistics (e.g., Stein et al. 2002), vegetation sampling (e.g., Bonham 2013), and plant trait measurements (e.g., Perez-Harguindeguy et al. 2013).
15.1.1 Why Plan? The Data Life Cycle
Central to every field campaign are the research questions and proposed explanations outlined in the form of testable hypotheses. It seems natural that planning the science (What data do we need to tackle our questions? What methods are available?) and planning the logistics (Where do we collect data and when? What resources do we need?) often rank above planning data organization and communication. However, starting a project with a data management plan (DMP) has a series of advantages. A DMP integrates several planning aspects in a structured way; it ensures the long-term sustainability of a project and its data, which is important not only because sustainability furthers scientific advancement (e.g., through data sharing and the reuse of data in meta-analysis) but also because it provides accountability for spending resources on research. DMPs are usually required in research proposals and make, through self-defined standards on data acquisition, data formats, documentation, and archiving, scientific work, including collaborations, more effective.
Funding sources often have their own guidelines about the structure and content of a DMP. Although only some of them might be required or relevant for a particular project, common components include:
-
Data collection and documentation: description of the types, formats, and volumes of data and samples and other materials collected, observed, or generated during a project, including existing data sources; description of the methods for data collection, observation, and generation, including derivative data; standards for ensuring data quality, including repeated measurements, sampling design, naming conventions, version control, and folder structure; description of the documentation standards for data and metadata format and content; and the software used for analyses
-
Ethical, legal, and security issues: details regarding the protection of privacy, confidentiality, security, and intellectual property rights, including information about access, use, reuse, and distribution rights; the time of data storage; possible changes to these rights over time; and strategies for settling disagreements
-
Archiving: description of storage needs for data, samples, and other research products; plans for long-term preservation, access, and security, including details on the parties and organizations involved; backup strategies; selection criteria for long-term storage; community standards for documentation; and file formats
A DMP covers all aspects of the data life cycle (Corti et al. 2014), including the following phases:
-
Discovery and planning: designing the research project and planning data management; planning data collection and consent for data sharing; outlining processing protocols and templates; and developing strategies for discovering existing data sources
-
Data collection: collecting data, including observations, measurements, recordings, experimentations, and simulations; capturing and creating metadata; and acquiring existing third-party data
-
Data processing and analysis: entering, digitizing, transcribing, and translating data and metadata; checking, validating, cleaning, and anonymizing data, where necessary; deriving, describing, and documenting data and metadata; analyzing and interpreting data; producing research outputs; authoring publications; citing data sources; and managing and storing data
-
Publishing and sharing: establishing copyright of data; creating discoverable metadata and user documentations; publishing, sharing, and distributing data and metadata; managing access to data; and archiving
-
Long-term management: migrating data to best format and suitable media; backing up and storing data; gathering and producing metadata and documentation; and preserving and curating data
-
Reusing data: conducting secondary analysis; undertaking follow-up research and conducting research reviews; scrutinizing findings; and using data for teaching and learning
Compiling a DMP, establishing guidelines for data and metadata collection and documentation, and outlining data use policies early in the planning phase is good practice. Starting discussions about how to organize data during or after data collection is a difficult task; reorganizing file structures, renaming files, and explaining and setting up new data structures will rarely be a top priority once data collection has started, and new data sets are ready to work with. Many organizations and agencies have their own standards (e.g., https://ngee-arctic.ornl.gov/data-policies) and can provide a good starting point when thinking about one’s own.
Communication strategies are another planning aspect that should not be overlooked (Sect. 15.2.2). Timely communication with site administrators is not only central to receiving permits and critical information; it also brings opportunities for public engagement during fieldwork, which is one of the most publicly visible parts of the scientific process. Even if site visits last for only a couple of hours, planning is important . Interactions with the public can happen at any time, so being prepared to answer questions and give a brief project overview in plain language, and perhaps having a flyer ready to hand out, can provide valuable opportunities for science communication. The support of stakeholders, such as site managers, local communities, and authorities, not only is important for a successful research project but also plays a critical role in determining the degree to which ecological research enters in public discourse and ultimately results in broader impact. Moreover, fieldwork brings opportunities for connecting researchers from different disciplines, which can aid in developing a common language, lead to new collaborations, and make projects more effective. Good research plans and communication strategies increase the chances for fruitful exchanges.
From the perspective of a project’s feasibility in terms of time, personnel, and budget, proper planning allows field campaigns to stick to their schedule (which is important because ecological processes change over time) and to the collection of data that are relevant for answering particular questions (it is easy to keep bolting on new measurements that slow down and jeopardize the main focus of a study). Moreover, adjusting to particular situations and handling challenges becomes easier when a detailed plan and the reasons behind it are clearly communicated to the research team. Clarity on the daily responsibilities and the project aims also help to keep research teams motivated.
15.1.2 Spectral Models and Scales of Measurement
Models are simplified descriptions of some aspect of the world and usually how it works (Fleishman and Seto 2009; Horning et al. 2010). Modeling is a multistep iterative process to formulate, by abstraction and idealization, a representation of reality (conceptual model), specify it mathematically (mathematical model), and “solve it,” which usually involves translating the math into computer code (computational model; Dahabreh et al. 2017). Models are used to test hypotheses, to assess relationships between response variables and factors that influence them, to investigate interactions between parts of a system, to make predictions about how a system will likely behave in the future, and to test how well models calibrated with data from the past fit current conditions (also known as hindcasting). Ideally, a model describes the full extent of the phenomenon of interest, but in practice, there are limits to the variables that can be determined in any given study. These limits can be formally described by model boundaries, which are as any ordering/bordering system an attempt to say something about (or attain power over) what is incompletely understood (or under-controlled; Jones 2009; Szary 2015). For the purpose of this chapter, model boundaries illustrate the situations under which a model is likely valid to some degree; ideally, likelihood and degree of validity are mathematically established. Model boundaries can be biological (e.g., specific to ecosystems, species, life stages), physical (e.g., specific to latitudinal, geological, hydrological, and topographical extents, or specific in time and place), or political (e.g., specific to regions or countries). Further, models can be classified as reductionist or system-based, quantitative or conceptual, correlative or mechanistic, static or dynamic, and hybrids thereof (Horning et al. 2010). Model boundaries and modeling approaches should be determined early in the planning phase and reported. Some model limitations are likely beyond the researcher’s control, such as instances in which data can only be acquired within certain political units. However, the choice of model to describe a particular system should be made deliberately; and the modeling approach should determine data collection, and not vice versa.
The data needed to investigate a phenomenon of interest with spectroscopy depend not only on the research question, the modeling approach, and model boundaries but also on the aim of the analysis (e.g., model calibration, validation, interpretation) and the level of spectral data acquisition (leaf-level spectroscopy, proximal, airborne, satellite RS; see Sect. 15.3., Gamon et al. Chap. 16). Data for model calibration and validation should match the conceptual model’s boundaries (e.g., the model’s temporal and spatial scale) and the modeling approach. For example, while quantitative and correlative models are ideally based on relatively uncorrelated or orthogonal variables, conceptual and mechanistic models ideally include all variables relevant for a particular study system. Drawing inferences from models and applying them to make predictions are only justifiable when model accuracy has been assessed (Horning et al. 2010); a model can give a very accurate description of a particular system, but one would not know until its accuracy is assessed.
Model calibration describes the process of determining the values of parameters so that model outputs fit the observed data. Internal validation refers to testing a model’s ability to explain the data used to populate the model. One common method for this is cross-validation. During cross-validation, the data set is split into calibration and (internal) validation data, the calibration data are used to fit the model, the model coefficients are applied to the validation data, and predicted and measured values from the validation data are statistically compared to assess model fit. Usually the data are split repeatedly, and model statistics (and often model parameters) are averaged across the number of splits. In k-fold cross-validation, k indicates the number of random data subsets or splits. One subset is omitted from model calibration and used for validation, and the process is repeated until all subsets have been left out once. For small data sets, leave-one-out cross-validation is particularly useful; here, only one sample is omitted from model calibration and used for validation, and the process is repeated until all samples have been left out once. In contrast to internal validation, external validation refers to a model’s ability to predict observations not used for model development (Dahabreh et al. 2017), which is critical for evaluating model performance and transferability. External validation involves either leaving out samples from the internal calibration-validation process or collecting additional independent data, followed by the evaluation of agreement between model output and observed data without any attempt to modify model parameters to improve fit.
Spectral models can be categorized into empirical-statistical and physical models and combinations between the two (see Verrelst et al. 2015). Empirical-statistical models are based on the relationship between the spectral behavior of certain spectral bands or the entire spectrum (the predictors or independent variables) and the vegetation characteristic(s) of interest (the dependent variable(s)). Empirical-statistical models generally use regression or clustering algorithms and aim to predict vegetation characteristics or class membership from a population of spectral data that were not included in the modeling process. For calibrating empirical-statistical models, it is essential to collect representative field data (see Sect. 15.2.3). Generally, this means that for the ecosystem, time of year, and area of interest, data should cover the range of values for which predictions are intended to be made or the number of vegetation classes with suitable replication, as well as the range of environmental conditions present in that area. Sample size should be large enough and samples distributed evenly across the expected range of values, classes, and environmental gradients to allow samples to be left out from model development and enable external validation.
Regression techniques are generally used for modeling and predicting continuous vegetation characteristics, such as biomass and chemical or structural composition (Ustin et al. 2009; Serbin et al. 2014; Schweiger et al. 2015a, b; Couture et al. 2016), or relative proportions of vegetation properties, such as the abundances of species, plant functional types, or vegetation types (Schmidtlein et al. 2012; Lopatin et al. 2017; Fassnacht et al. 2016; Féret and Asner 2014; Schweiger et al. 2017). Univariate, multivariate, linear, and nonlinear regressions are common for modeling and predicting vegetation characteristics from few spectral bands or from spectral indices. Spectral indices are used to infer vegetation status, including plant stress, and ecosystem parameters, inducing productivity, from empirical or physical relationships between spectra and plant traits. Many spectral indices have been published (see, e.g., https://cubert-gmbh.com/applications/vegetation-indices/). The most widely used indices include the normalized difference vegetation index (NDVI; Rouse Jr et al. 1974; Tucker 1979), an indicator of vegetation greenness, and modified versions such as the soil-adjusted vegetation index (SAVI; Huete 1988) and the photochemical reflectance index (PRI; Gamon et al. 1992). The NDVI and its variants have been shown to correlate well with biomass, LAI, and the photosynthetic capacity of canopies. The PRI estimates light-use efficiency and can be used to estimate gross primary productivity (GPP) and assess environmental stress (Sims and Gamon 2002). Spectral indices can also be used directly to estimate certain environmental characteristics (Anderson et al. 2010; Pettorelli et al. 2011; Wang et al. 2016). For example, the NDVI has been used for predicting and mapping taxonomic diversity (e.g., Gould 2000), the rationale being the expected increase in ecological niches with increasing energy or resources in ecosystems (Brown 1981; Wright 1983; Bonn et al. 2004). In addition, advances in sensor technology allow capturing aboveground productivity in ecologically meaningful units, such as the annual amount, variation, and minimum of photosynthetically active radiation (fPAR), which has been found to explain global patterns of mammalian, amphibian, and avian diversity to a substantial degree (Coops et al. 2018). The idea behind spectral indices is that they are generally applicable and transferable; however, ground reference data are still needed to assess their accuracy. In addition, site-specific data are also needed to recalibrate spectral indices, for example, by selecting the optimal wavelengths for the sensor used and by estimating site-specific model coefficients, because index responses often vary with the particular context.
Latent variable methods, such as partial least squares regression (PLSR) and partial least squares discriminant analysis (PLSDA), were developed for chemometrics and specifically deal with the high degree of autocorrelation inherent in data with high spectral resolution (Wold et al. 1983; Martens 2001). PLSR is a standard method for modeling and predicting continuous vegetation characteristics and PLSDA for determining class membership from spectral data. Clustering methods, including principal component analysis (PCA), principal coordinate analysis (PCoA), and linear discriminant analysis (LDA), are frequently used to explore patterns, such as the degree to which species or plant functional types cluster separately from each other in spectral space. In addition, machine learning algorithms [e.g., random forest (RF) or support vector machines (SVNs)] and deep learning methods [e.g., convolutional neural networks (CNNs)] can be used for classification problems, including the identification of vegetation types based on physiognomic attributes (e.g., forest, shrubland, grassland, cropland), for the detection of plant pathogens and other stresses (Pontius et al. 2005; Herrmann et al. 2018), and for species detection (Clark et al. 2005; Fassnacht et al. 2016; Kattenborn et al. 2019).
Physical models are based on causal physical relationships between electromagnetic radiation and vegetation properties; in spectroscopy, these models are called radiative transfer models (RTMs). For leaf optical properties, the RTM PROSPECT (Jacquemoud and Baret 1990; Jacquemoud et al. 2009) models leaf reflectance and transmittance based on leaf chlorophyll a and b content, “brown pigment” content, equivalent water thickness, leaf dry matter content, and a leaf structure parameter indicating mesophyll thickness and density. For modeling optical properties of canopies, leaf-level RTMs can be combined with canopy RTMs (Jacquemoud et al. 2009), which incorporate structural canopy parameters, including leaf area index (LAI, the ratio of leaf area to ground surface area), leaf inclination, a hot spot parameter (a function of the ratio of leaf size to canopy height), as well as soil reflectance and measurement characteristics, including sun and viewing angle. Frequently used canopy RTMs include SAIL (Verhoef 1984), GeoSAIL (Huemmrich 2001), and DART (Gastellu-Etchegorry et al. 2004). Although RTMs do not model all interactions between plants and light, because they cannot incorporate all characteristics of leaves and vegetation canopies that influence the spectral response, they are useful for simulating spectra and retrieving estimates about plant characteristics. In “forward mode,” RTMs simulate spectra from the vegetation parameters incorporated into the model, e.g., by using the expected range of values for chlorophyll content, SLA, and other parameters, in certain ecosystems or for certain species. In “backward mode,” RTMs estimate vegetation characteristics from spectra (Vohland and Jarmer 2008; Weiss et al. 2000). Model inversion is usually done by running an RTM in forward mode and systematically varying the input parameters using lookup tables (LUT) with different trait combinations, until the measured spectral signal is sufficiently well approximated. The most plausible input parameter combinations can then be averaged to provide estimates of vegetation traits. In principle, RTM inversion can be conducted without ground reference data collected on-site (e.g., when input parameters can be sourced from plant trait databases, such as TRY; Kattge et al. 2011) or determined based on expert knowledge. However, inversion of RTMs is generally an ill-posed problem in the sense that there is not a single solution but rather multiple solutions to model inversions (i.e., multiple input parameters can yield the same output spectra). Ground reference data are important for RTMs because they can be used to limit the ranges of possible input values (Combal et al. 2003) and are essential for model validation.
15.2 Planning Field Campaigns
This section includes thoughts about data organization (Sect. 15.2.1) and communication (Sect. 15.2.2), before covering the planning of data collection in more detail (Sect. 15.2.3).
15.2.1 Data Organization
Data organization schemes help define and implement guidelines to make project management and collaborations more efficient and ensure long-term project sustainability and the reproducibility of research (https://ropensci.github.io/reproducibility-guide/). Guidelines for folder structure, file names, documentation, file formats, data sharing and archiving, version control, and data backups are all part of data organization. When archiving is handled by a third party, researchers need to consider how to structure data and metadata to match external requirements. Generally, it is good practice to work backward and start with identifying where project data will be stored long term and which data and metadata standards will make long-term storage possible and data sets discoverable and reusable later. For instance, it is important to use community standards for taxon names, units, and keywords and to store data in file formats that are nonproprietary (open), unencrypted, and in common use by the research community. The US Library of Congress has released a recommended file format statement (http://www.loc.gov/preservation/resources/rfs/) and provides detailed format description documents for different categories of digital data, including data sets, images, and geospatial data. Once data characteristics for long-term storage are known, one can define short-term storage structures and the workflow leading from raw data collection, to cleaned-up data, to preliminary and final results and products. Data backups should match the original data structure so recovery requires only a few minutes. Ideally, backups are done automatically, continuously, and incrementally, and data history is preserved. Data recovery should be tested on a regular basis. Additionally, it is important to check how long data backups are being sustained. When storage space is limited, it makes sense to use a time-dependent structure, such as keeping daily backups for a year, biweekly backups for 3 years, and monthly backups thereafter. Several resources provide details about good data management practices, including the Oak Ridge National Laboratory Distributed Active Archive Center (https://daac.ornl.gov/datamanagement/) and the rOpenSci initiative (https://ropensci.github.io/reproducibility-guide/).
Fundamentals of data organization include (see, e.g., Cook et al. 2018):
-
Definition of file/folder content: Keep similar measurements in one data file or folder (e.g., if the documentation/metadata for data are the same, then the data products should all be part of one data set).
-
Variable definition: Describe the variable name and explicitly state the units and formats in the metadata; use commonly accepted names, units, and formats and provide details on the standards used (e.g., SI units, ISO standards, nomenclature standards); use format consistently throughout the file; use a consistent code (e.g., −9999) for missing values; and use only one variable per measurement (e.g., avoid reporting coordinates in more than one coordinate system or time in several time zones).
-
Consistent data organization: Do not change or rearrange columns in the original data; include header rows (first row should contain file name, data set title, author, date, and companion file names); use column headings to describe the content of each column; include one row for variable names and one for variable units; and make sure either each row in a file represents a complete record (with columns representing all variables that make up the record) or each variable is placed in an individual row (e.g., for relational databases).
-
Stability of file formats: Avoid proprietary formats, and prefer formats encoding information with a lossless algorithm (e.g., text, comma/tab-separated values, SQL, XML, HTML, TIFF, PNG, GIF, WAV, postscript formats).
-
Descriptive file names: Use descriptive, unique file names; use ASCII characters only and avoid spaces (e.g., start with ISO date, followed by descriptive file name: 20180430_siteA_plotB_vegSurvey); remember that file names are not a replacement for metadata; explain naming structure of files in metadata; organize files logically; and make sure directory structure and file names are both human- and machine-readable (check operating or database system limitations on file name length and allowed characters).
-
Processing information: Consider including information on software or programming language and version; provide well-documented code and information about data transformation.
-
Quality checks: Ensure that data are delimited and lined up in proper columns; check that there are no missing values (blank cells); scan for impossible and anomalous values; perform and review statistical summaries; and map location data.
-
Documentation: Document content of data set; reason for data collection; investigator; current contact person; time, location, and frequency of data collection; spatial resolution of data; sampling design; measurement protocol and methods used, including references; processing information; uncertainty, precision, accuracy, and known problems with the data set; processing information; assumptions regarding spatial and temporal representativeness; data use and distribution policy; ancestors and offspring of data set (including references to publications).
-
Data protection: Create backup copies often and without user interference (automatically, continuously, incrementally); three copies (original, on-site external, off-site) are ideal; test restoring information; and use checksums to ensure that copies are identical.
-
Data preservation: Preserve well-structured data files with variables, units, and values defined; documentation and metadata records; materials from project wiki/websites; files describing the project, protocols, and field sites (including photos); and project proposal (at least parts) and publications in open-access archives. Check platform standards for data archiving beforehand.
The best way to organize data depends on the project, size of the team, and degree of interaction among team members, among other things. It is good practice to think about ways to organize data early in the project-planning phase and to include at least the core project team in these discussions. However, differences in personal work styles can be a challenge for reaching agreements; the larger the team, the more difficult this becomes. In such cases, top-down approaches to data organization can be a good option, especially ones that have been tested before. Laying out data organization schemes at an early project stage and inviting people’s feedback at this stage is good practice. Clearly, there should be room for discussion during later project stages as well (particularly when the existing organization scheme is not working as expected), but generally adjustments become more complicated the longer projects are running. Data organization schemes are intended to make daily workflows, data exchange, and data archiving easier; they should not cause an extra workload. Research teams are much more likely to adapt a particular organizing scheme when it is simple and intuitive, and everyone is much more likely to stick to a system when its benefits are obvious. In the end, even the best organization structure fails when no one is following it.
Box 15.1 An Example of Folder Structure
It can be advantageous for research groups and institutions to implement common data standards and a file structure that forms the backbone of a data organization scheme and does not have to be discussed for every new project. Data standards also promote reproducible research. One option is to set up file directories separated by content. The example below structures a higher-level directory (e.g., the project directory) into docu_work, docu_pub, orig_data, data_work, data_pub, gis_work, gis_pub, maps, and printout folders. The key feature of this structure is the distinction between work folders, containing work in progress and intermediate results, and pub folders, containing the final versions and results. After the completion of a project, the orig_data, maps, printout, and all pub folders are archived (publicly and internally), while all work folders get deleted from a public or shared drive. Backups are kept at a certain frequency for a certain amount of time (e.g., daily backups for a year, biweekly backups for 3 years, monthly backups for 10 years), and personal copies can of course be kept as long as necessary. A folder structure like this makes archiving data easy because at each project stage it is clear which data sets, documents, and products will be preserved. The pub, orig_data, and maps folders should contain everything a person without knowledge about the project would need to repeat the analysis, including basic project background and workflow descriptions in the docu_pub folder, but nothing unnecessary, such as intermediate results. One testable reproducibility goal could be that a person with appropriate analytical skills but without any information about the project would be able to re-create and explain a main result, including the rationale behind the analysis, after 1 workday without any external help. For GIS heavy projects, it may make sense to separate analysis, results, and products based on geospatial data from those using other data sources. In this example, map products are reproducible based on data from gis_pub and the layouts found in the maps folder.
Short Description of Contents and Management of Example Folder Structure
Folder type | Content description and management |
|---|---|
docu folders | These are for the proposal, project descriptions, documentation of workflows, planning documents, minutes of meetings, manuscripts, photos, etc. File names could, for example, start with the ISO date followed by a descriptive name. Files are usually organized into subfolders. Typical file formats include .doc, .txt., .pdf, and .tiff. The docu_work folder is deleted after the project is finished. The docu_pub folder contains the final versions in a non-editable format, such as .pdf. It should contain the essentials of the project background and all workflows needed to repeat the analysis; detailed descriptions should either be left out or clearly flagged, e.g., as “additional information.” The docu_pub folder also includes information about the use of corporate or project identity styles (use of logos, colors, fonts, etc.). This folder gets archived after the project is finished |
orig_data folder | This folder includes raw data acquired during the project. These data never get changed. “Read only” permission is advisable; metadata files describing the data sets are critical. It is good practice to check backup copies when new data sets are added. From here data can be copied to the data_work folder, e.g., if the format needs to be changed or different data sets are being combined into a master data set. This folder can contain proprietary file formats, in which case it is important to include details about the software and version used to access files. This folder gets archived after the project is finished |
data folders | These are where analyses happen. The folders usually contain subfolders, e.g., for code, data input, and data output. Work copies of data copied from orig_data are saved here. Metadata describing any changes to the original data are important, including variable transformations, references for methodology, software, and version used. Typical file formats are .csv and .txt. The data_work folder includes preliminary results and is deleted after the project is finished. All analysis steps are being documented in the docu folders. The data_pub folder contains final scripts, final results, compiled master data sets, etc. This folder gets archived after the project is finished |
gis folders | These are similar to data folders but for geospatial data. This folder contains geo data that have been modified from the original data. Processing details are included in the metadata; original data remain in orig_data. Typical file formats include geotiff, .tiff, and .shp. The gis_work folder is for intermediate steps and is deleted after the project is finished. The gis_pub folder is for final results and gets archived |
maps folder | This folder is for data associated with maps, layouts, and styles. It gets archived after the project is finished. All paths in maps should refer to data in the data_pub or orig_data folders when the project is finished |
printout folder | This folder contains final products including publications, maps, posters, and presentations, usually in a non-editable format. This folder gets archived after the project is finished |
15.2.2 Communication
Communicating plans for fieldwork and applying for necessary permits in a timely manner avoids unnecessary complications. Essentially, the earlier researchers get in touch with site administrators, the better. Regulations vary, but obtaining necessary permits can take months, especially in areas with a high protection status. Often, research proposals are evaluated by a panel. However, it is good practice to get in touch with site administrators before writing a detailed proposal because local regulations might influence project plans, including changes to the location, timing, and duration of data collection; the equipment used; and the number of people on site. In addition, it is good practice to figure out logistics, such as transportation of people and equipment, early in the planning phase. Early communication provides time to understand the rationale behind a research plan and to adjust the plan appropriately. Often, it is not until details are questioned that it becomes clear which aspects of a research plan are critical and which can be handled somewhat more flexibly.
For research projects planned on or close to Indigenous lands, it is critical to inquire early in the planning process about local procedures and ethical guidelines. Generally, site administrators would be the first points of contact and should be aware of how to communicate the research objectives to the Indigenous communities and judge the level of involvement required (e.g., between short- and long-term studies). However, it is important for researchers to initiate this inquiry and to seek additional guidance if needed. For both short- and long-term projects, familiarizing oneself with ethical frameworks for research with Indigenous communities and/or on Indigenous lands (e.g., Claw et al. 2018) is good practice.
15.2.3 Planning Data Collection
Remote sensing of plant biodiversity typically includes the comparison of ecological or spectral data collected on the ground with remotely sensed images, often through a model that considers scale effects. Two aspects are critical to data collection: (i) sampling representative areas and/or individual plants that can be aligned with the imagery and (ii) sampling at high spatial accuracy and a level of precision that matches the sensor. With these two points in mind, the following sections discuss area selection (Sect. 15.2.3.1); value ranges (Sect. 15.2.3.2), which are important for model representativeness and thus also of interest for the validation of physical models; and sampling design (Sect. 15.2.3.3), with a particular focus on empirical-statistical analyses. Data collection is typically formalized in a sampling plan. A sampling plan describes data acquisition, recording, and processing (Domburg et al. 1997) and includes the first elements of the data life cycle (see Sect. 15.1.1). Consequently, the plan will likely include decisions about the area selection and variables measured, logistical constraints, sampling and analysis methods, sampling design, sampling protocols, estimation of measurement accuracy/precision, and operational costs.
Sampling is a method of selection from a larger population carried out to reduce the time and cost of examining the entire population (Justice and Townshend 1981). In the case of sampling plant biodiversity at a particular field site, we are generally selecting individual plants to represent local populations of a set of species that capture the range of functional and phylogenetic variation in a site or represent the dominant species. Data collection balances accuracy and representativeness against time and budget. Two questions are central to planning data collection: (i) Which population(s) is (are) best suited to answer the specific research question(s)? and (ii) Is (are) the population(s) adequately represented by the sampling scheme? During the early planning phase, it is important to get a sense of which environmental factors cause variation in the samples to be collected and variables to be measured (Johannsen and Daughtry 2009) and to choose research sites accordingly. This requires researchers to familiarize themselves with the conditions at the site. It is also critical to define at an early planning stage how the data will be analyzed and to choose the sampling design and sample size accordingly (see Sect. 15.2.3.3). Improper randomization in particular can lead to biased conclusions based on inappropriate assumptions (De Gruijter 1999).
15.2.3.1 Area Selection
It is rarely possible to describe all aspects of an ecological phenomenon of interest in a single study. Model boundaries help clarify the conditions under which a model is likely valid (see Sect. 15.1.2). Research areas represent conceptual model boundaries, including ecosystem(s), plant community(ies), species, and geographic location. Although it is good practice to formulate model boundaries first and pick research areas accordingly, in practice model boundaries usually need to be adjusted after selecting a research area to reflect the conditions at the site. Formulating the model first and refining it during the planning process help clarify a study’s limitations; stating them clearly is important for the analysis and further synthesis work.
Once model boundaries have been formulated, it is important to investigate which environmental conditions influence the response and explanatory variables in the study system and how they are spatially distributed. Generally, it is critical to cover the range of environmental conditions, both biotic and abiotic, for which model inferences are being made, including the diversity and distribution of vegetation communities, plant species, successional gradients, soil types, soil moisture and nutrient gradients, aspects, slopes, land uses, microclimatic conditions, animal communities, pathogens, and other factors determining environmental heterogeneity in a study area. Accounting for the variation of every factor might not be possible, but many environmental factors are correlated. If possible, it is good practice to investigate the covariance structure of environmental factors based on previous studies and existing data and to focus on a few factors that are expected to have the most effect on the phenomenon of interest. Data collection might need to be limited to a smaller area than anticipated due to a high degree of environmental heterogeneity and/or time constraints, which affects the range of conditions for which conclusions can be drawn. However, it is generally advantageous to work with sound models for small areas with a limited degree of environmental variation than to work with weaker models for larger areas. Testing model predictions outside the model boundaries can provide important insights regarding model transferability, the comparability of ecological conditions, and differences and similarities in ecosystem function between areas.
Maps, local sources, and other research groups can provide important information regarding environmental variation in a research area. Again, it is helpful to contact site administrators early to gain access to resources and build connections to other research groups. Local administrators can often give advice regarding the timing and location of sample collection, including practical considerations such as accessibility. Visiting a potential research area can be extremely helpful during the planning phase. Often, it is easier to discuss issues and concerns in person, and seeing the conditions on site facilitates decision-making. Joining another research group for some time in the field or accompanying a person who knows the area well provides a great way to get to know an area. Covering the heterogeneity of a research area is important for all aspects of remote sensing of plant biodiversity, including collecting spectral references for image processing and sampling ground data (such as vegetation spectra and vegetation samples) for model calibration, validation, and interpretation. For example, empirical line correction (ELC, Kruse et al. 1990, Broge and Leblanc 2001), a common method for correcting atmospheric and instrument influences on remotely sensed data, utilizes ground measurements of invariant surfaces (e.g., pavements, rocky outcrops, snow, water, calibration tarps) that can be readily identified in the image and/or for which accurate location data has been acquired. As for any empirical method, the performance of ELC depends on the representativeness and accuracy of the input data, and model transferability is limited. Thus, target surfaces for ELC should ideally be distributed across the area of interest (e.g., located across all flight lines) and cover differences in altitude, slope, and aspect (see Sect. 15.3.3.2).
Sampling biodiversity in a way that fits remotely sensed data means incorporating the heterogeneity of a research area but also requires thinking about the size and shape of sampling units. Generally, sampling units should be delineated to encompass areas with similar environmental conditions. Remotely sensed imagery are usually raster data, so ground measurements need to represent areas rather than points. The optimal size of the sampling units on the ground depends on the spatial heterogeneity and spatial resolution of the imagery. Sampling units that are smaller or the same size as the pixels in the imagery are usually unrepresentative (Justice and Townshend 1981). Pixel shifts are a common consequence of image processing, and averaging RS data across several pixels is common practice for noise reduction. As a general guideline, the minimal dimensions of a representative sampling unit can be calculated as A = P ∗ (1 + 2 L) (Justice and Townshend 1981), with P being the pixel dimensions of the image and L the accuracy of image alignment in number of pixels. For example, if the spatial resolution of an image is 3 m and a one-pixel shift is expected to occur during image processing, the minimal size of an internally homogeneous sampling area would be 3∗(1 + 2 ∗ 1) = 9 m × 9 m. This makes it possible to capture similar environmental conditions even when the image pixel that should align with the sampling unit’s center pixel has shifted one pixel in either direction or when averaging the center pixel and its neighboring pixels (Fig. 15.1). In this particular context, internal homogeneity does not mean that the sampling unit can only consist of one particular feature but rather that all features should be evenly distributed throughout the unit. In other words, to be considered internally homogeneous, a sampling unit does not have to consist of a single plant species of one particular age or size class; it can consist of different species and individuals as long as their spatial distribution is comparable among the pixels within that unit. For example, the center pixel in Fig. 15.1a is not representative for the sampling unit because species abundance varies among the nine pixels; the sampling area is internally heterogeneous. In contrast, the center pixel in Fig. 15.1b is representative for the sampling unit because species abundance is similar among the nine pixels; the sampling area is internally homogeneous.
(a, b) Internally heterogeneous and homogeneous sampling units. (Adapted from Justice and Townshend 1981)
Another way to determine the minimum size of sampling units for RS studies are structural cells, which are defined as area units that are large enough to fully capture the variation within one particular feature on the ground, such as the variation in the spatial distribution of individual plant species within a plant community or the variation in the terrain characteristics within a particular topographic feature (Grabau and Rushing 1968). If the size of a structural cell is larger than the image pixel plus its accuracy buffer, structural cells should be preferred; their size can vary depending on the variation of the environmental feature of interest. It is also worth mentioning that pixels as seen by a sensor are not square but elliptic and that surrounding pixels contribute substantially to the signal detected per focus pixel (Inamdar et al. 2020). Theoretically, elliptic or hexagonal sampling units, representing shapes that are frequent in nature, should capture local environmental conditions better than square plots. However, since remotely sensed images are usually subsampled to make pixels quadratic, a case for square sampling plots can be made. As pointed out earlier (Sect. 15.1.2), drawing hard boundaries around any natural feature is notoriously flawed because gradients are the norm and abrupt changes the exception. Thus, it is good practice to specifically sample ecotones and other transition zones if possible—or, if not, to acknowledge that a model might not be representative for transition zones when they are not sampled. Generally, a sampling unit can be considered adequately described when measurements within that unit cover the variation of the characteristic of interest. Thus, it is not necessary to sample entire sampling units when they are internally homogeneous (Fig. 15.1b). For example, when the plant species composition in every 1 m2 in a 9 m × 9 m research plot closely resembles that of every other 1 m2, it is sufficient to conduct a species inventory within 1 m2, ideally in the plot center. Likewise, it can be expected that the chemical composition of the biomass clipped in a 1 m × 20 cm strip in the central 1 m2 is representative for the chemical composition of the vegetation in the entire 9 m × 9 m plot, given that the clip strip captures the variation in species composition, height, and age-class distribution within the central 1 m2.
The accuracy and precision of the surveying equipment used for measuring plot coordinates is another aspect to consider when determining the minimal size of homogeneous sampling units; ideally, measurement errors should be estimated under field conditions and added to the minimal size of the sampling unit. Additionally, edge effects influence remotely sensed data. Sampling units should be placed sufficiently far from landscape features, such as open soil, gravel, snow, water, large rocks, roads, footpaths, bridges, and trees (when working in grasslands), that influence the spectral properties of adjacent areas. Depending on the time of day of image acquisition, shadow effects from tall objects such as trees, mountains, or buildings need to be taken into account as well.
15.2.3.2 Range of Values
For the remote sensing of biodiversity, the range of conditions (time of year, value range, species, and environmental context) used to calibrate a model should cover the range of conditions for which inferences or predictions should be made. For example, extrapolating beyond the range of values relies on the assumption that the estimated relationship holds beyond the investigated range. This cannot be assumed without additional information, because nonlinearities (e.g., saturating curves) are common in ecological data, especially when covering large areas and multiple environmental gradients and ecosystems.
For modeling continuous data with regression-style empirical-statistical approaches, the sampling design should cover the expected range of values in the area of interest with a sufficient number of evenly distributed samples. Predictions outside the calibrated range are not reliable because deviations from the 1:1 line between measured and predicted values (Fig. 15.2a) increase at the lower and upper ends of the distribution (Fig. 15.2b). During model validation the entire range of values should be covered as well, specifically paying attention to the value range most important for the research question(s). When the tails of the distribution are of interest for predictions, it is good practice to include a number of extreme values in the validation. These values can be used for updating the model, extending model validity beyond the previously calibrated range or beyond previously covered environmental contexts (Fig. 15.3c).
(a) A calibrated model (solid black line) generally deviates to some degree from the ideal 1:1 relationship (dashed black line) between measured and predicted values; (b) the deviation becomes more pronounced when predicting samples outside the calibrated range of values (red dots); (c) using the measured values of these samples (green dots) for calibrating a new model extends the range of values for which the model is valid (green solid line; note that in this case the samples are not evenly distributed such that model performance in the gaps of the value range, i.e., between black and green points, is unknown)
Examples of sampling designs: (a) simple random, (b) stratified random, (c) two-stage, (d) cluster, (e) systematic, and (f) spatial systematic sampling. (Adapted from De Gruijter 1999)
However, a larger range of values and environmental contexts is not automatically better. Empirical-statistical models are context specific. Transferability beyond the time of year, value range, species, and environmental context for which they are calibrated cannot be assumed without a test. The power of empirical-statistical methods lies in their ability to fit the data. Thus, empirical-statistical models should be calibrated for the range of values and the environmental conditions that are most relevant for answering a specific research question. Extending the calibrated range beyond the range of values for which predictions are being made generally decreases model accuracy for these values, as compared to a more narrowly defined model that fits that range.
For classification models, including empirical-statistical clustering and supervised classification methods, covering all classes of interest with a similar and sufficiently large number of samples is also important. However, the range of values/environmental conditions question is more nuanced. On the one hand, it can be advantageous to include all major classes as end-members for classification, even when not all of them are of interest for prediction. For example, when the aim is to differentiate broadleaf and needleleaf forest using remotely sensed imagery, it makes sense to include other classes present in the image, such as grasslands, roads, and water bodies, as well. The reason is that when broadleaf and needleleaf forest are the only two classes used for model calibration, the model will, when applied to the full image, try to assign grassland, road, and water pixels to these two forest types, decreasing model accuracy. On the other hand, too many extra classes can make it difficult for a model to differentiate among the classes of interest. If one is interested in differentiating two forest types, it would probably not make sense to use tree species as input classes, because a species differentiation model is likely overall less accurate than a model trained on just the two forest types of interest. Stepwise approaches to such classification problems are often helpful. First, one could differentiate broader classes such as vegetation, roads, and water bodies from each other; then, forest from grassland within the vegetation class; and finally different forest types within the forest class (for more details see textbooks on remote sensing of vegetation, e.g., Jones and Vaughan 2010; Thenkabail et al. 2012, and specific topics, such as on deep learning approaches to image classification, e.g., Cholet and Allair 2018).
Areas from within the calibrated value range are usually prioritized during model interpretation. Nevertheless, visits to areas with vegetation or other site characteristics at the edge of or beyond the calibrated range can be insightful regarding the limits of a model’s applicability. Field visits are ideal for investigating how environmental context and time of year influence model performance and for determining under which conditions a model performs well or poorly.
15.2.3.3 Sampling Design
A good approach to deciding on a sampling design and to planning data collection for remote sensing of plant biodiversity in general is to start at the end and think about (i) what type of modeling result or product would be most useful with respect to the research question (such as a hypothesis test at a given significance level or a map of a variable with a given accuracy), (ii) what kind of data analysis would lead to that result, (iii) what data properties are needed for the specific analysis, and (iv) how these data can be collected efficiently (De Gruijter 1999).
In a spatial context, a sampling design assigns a probability of selection to any set of points in a research area, while a sampling strategy is defined as the combination of sampling design and the estimator of the variable of interest, for which statistical quality measures (such as bias or variance) can be evaluated (De Gruijter 1999). Sampling designs can be model-based or design-based; the two approaches use different sources of randomness for sample selection and model inferences (Brus and DeGruijter 1993; Domburg et al. 1997). Harnessing as much information about spatial variation as possible, including maps of the study region and theory about spatial patterns, facilitates finding an efficient sampling design for both model- and design-based sampling strategies.
Model-based sampling is based on geostatistical theory and evaluates uncertainties by using a fixed set of sampling points while the pattern of the values of interest varies according to a defined random model of spatial variation (De Gruijter 1999). Model-based sampling strategies are, for example, used for kriging, a spatial interpolation method that uses measured point values to estimate unknown points on a surface. The ideal situation for using a model-based sampling scheme is when the desired result should be the prediction of values at individual points or of the entire spatial distribution of values in the research area (i.e., a map), when a large number of sample points can be afforded to calculate the variogram (~100–150 sample points, Webster and Oliver 1992), and, most importantly, when a reliable model of spatial variation is available, the spatial autocorrelation is high, and there is a strong association between the model of spatial variation and the variable of interest. The association between a geostatistical model and the variable of interest is particularly important, because the final inferences about the spatial distribution of the variable of interest are based on the model of spatial variation (De Gruijter 1999; Atkinson 1999). However, it is often difficult to decide if model assumptions are acceptable because several decisions for defining the spatial structure (e.g., about stationarity, isotropy, and the variogram) are subjective (Brus and DeGruijter 1993).
Ecological systems are often too complex to use model-based inference with much confidence (Theobald et al. 2007). Nevertheless, if a tight relationship between a geostatistical model and nature is expected, model-based sampling schemes are useful, for example, to find at a defined accuracy the optimal sampling grid orientation and spacing for kriging methods (see, e.g., Papritz and Stein 1999) and to define ideal locations for additional sampling points when some sampling points are predefined or spatially fixed. Moreover, model-based sampling encompasses methods such as convenience sampling (sampling at locations that are easy to reach) and purposive sampling (sampling sites chosen subjectively to represent “typical” conditions). Although no formal statement of representativeness can be made for these methods (Justice and Townshend 1981), and they are not appropriate for accuracy assessments (Stehman and Foody 2009), they can provide valuable information in a geostatistical context (for an introduction to geostatistics, see, e.g., Atkinson 1999, Chun and Griffith 2013).
Design-based sampling is based on classic sampling theory. It evaluates uncertainty by varying the sample points while the underlying values are unknown but fixed (De Gruijter 1999). Statistical inferences from design-based sampling are valid, regardless of spatial variation and patterns of spatial autocorrelation, because no assumptions about spatial structure are being made. Design-based sampling schemes can be classified depending on how randomizations are restricted. Two or more designs can be combined (De Gruijter 1999; Fig. 15.3):
-
Simple random sampling (Fig. 15.3a): No restriction is placed on randomization; all sample points are selected with equal probability and independently from each other.
-
Stratified random sampling (Fig. 15.3b): The area is divided into subareas (strata; small squares), and simple random sampling is performed in each stratum. This reduces the variance at the same sampling effort or the sampling effort at the same variance. Strata can be based on maps of environmental parameters (soil types, vegetation types, aspect, etc.) and can have any shape. Cost functions can be included for determining sample size. Generally, more points are sampled in larger, more variable, or cheaper to sample strata.
-
Two-stage sampling (Fig. 15.3c): The area is divided into subareas (also called principle units, PUs), but only a random subset of these subareas is sampled; within a subarea, sample points are selected with equal probability. This clustering of points is more time-efficient but less precise than simple random sampling.
-
Cluster sampling (Fig. 15.3d): Predefined sets of points (clusters) are sampled. The starting point of each cluster is selected at random; the geometry of the cluster is independent of the starting point (e.g., transects with equidistant points extending in opposite, predefined directions from the starting point). The regularity of the clusters makes sampling more time-efficient but less precise than simple random sampling.
-
Systematic sampling (Fig. 15.3e): Similar to cluster sampling, a predefined set of points is selected at random, but only one cluster is selected (e.g., a random grid); interference with periodic variations can be avoided by combining systematic sampling with a random element (e.g., two-stage sampling combined with cluster sampling).
-
Spatial systematic sampling (Fig. 15.3f): Randomization restrictions are used at the coordinate level; the area is split into strata, and one point is selected at random. The points in the other strata are not selected independently but follow a specific model (e.g., a Markov chain).
It is good practice to conduct a sensitivity analysis for estimating the sample size needed to detect differences in the parameter of interest with the desired level of confidence (Johannsen and Daughtry 2009). The sample size needed to estimate a statistical property with a chosen probability depends on the sampling scheme, the desired error rate, and the variation of the ecosystem property of interest (which can be approximated from existing data or a pilot study or based on literature values and experience). Details for estimating sample sizes for the sampling designs mentioned above are given by De Gruijter (1999). However, error rates of spectroscopic models of vegetation characteristics also depend on the measurement accuracies of vegetation and spectral data and the tightness of the association between the property of interest and spectral data. For example, as a rule of thumb, the smaller the amount of the chemical compound of interest and the less precise the laboratory method used to determine that compound, the more samples will be needed for building a sound model. Similarly, for classification models, the number of samples needed to differentiate classes with a desired accuracy will depend on intra- and interclass variation or the distinctiveness of classes. In other words, when projecting samples from different classes into spectral (or more generally, feature) space, model accuracy for class differentiation depends on the number of classes, the spread of the distribution of values within classes, and the distance among the class centroids. As before, the strength of the relationship between classes and their spectral characteristics and measurement accuracies should be taken into account when deciding on sample sizes. As a rule of thumb, a minimum of 50 samples per class and 75–100 samples per class for more than 12 categories or areas larger than 4000 km2 has been suggested (Congalton and Green 1999), but fewer samples can provide sufficient accuracy when classes are relatively dissimilar. Tracking error propagation is important for assessing the performance of spectral models (Singh et al. 2015, Wang et al. 2019, Serbin and Townsend, Chap. 3); ideally, the accuracy of laboratory analysis should be included in error assessments, as well.
For empirical-statistical models that combine vegetation characteristic and spectral measurements, stratified random sampling is often a good choice. Stratifying a research area based on ecologically relevant environmental variation helps cover the heterogeneity of a research area and the range of values of the vegetation characteristics of interest. A range of methods for automating sampling designs are available for R, including the packages spsurvey (Kincaid and Olsen 2016), spcosa (Walvoort et al. 2010), spatstat (Baddeley and Turner 2005), and spatialEco (Evans 2017), and ArcGIS, including the Geospatial Modeling Environment (Beyer 2010) and the Reverse Randomized Quadrant-Recursive Raster algorithm (RRQRR, Theobald et al. 2007). However, it can be difficult to automate sampling design completely, especially in natural ecosystems with limited accessibility. Moreover, for studies with a RS component, it can be difficult to select research plots that are internally homogeneous and located far enough from objects that influence the spectral signal of neighboring pixels (see Sect. 15.2.3.1) automatically. Under such circumstances, a mix of automated sampling based on GIS data and informed decision-making (convenience/purposive sampling) can be a good option. For example, information about environmental factors and gradients influencing the vegetation characteristics of interest and other relevant information about the study area, such as accessibility and travel time, can be used as input into a GIS and used as strata. Random points per stratum can be created automatically and used, for example, to define larger polygons within which the exact location of research plots is determined in the field. When vegetation characteristics are expected to vary along gradients, cluster sampling of plots at predefined intervals along these transects is a good choice—but again, it might be necessary to adjust these distances to avoid objects influencing the spectral signal of the plots or to find internally homogeneous areas. In this context, areas can be considered “internally homogeneous” when their biotic and abiotic characteristics are comparable, which means that they can actually show a high degree of small-scale heterogeneity (e.g., situations changing every 5 cm) as long as this small-scale heterogeneity creates a similar mosaic at the measurement scale (e.g., 1 m2 is comparable to the adjacent 1 m2; see Sect. 15.2.3.1). It is important to report the reasons for deviating from common sampling schemes in the methods.
When working at the level of individual plants, sampling random points within research plots makes it possible to capture interindividual variation, which can be important, for example, when scaling functional traits of individual plants to plot-level estimates (Wang et al. 2019; Serbin and Townsend, Chap. 3). Random sampling combined with species identification can also be used as an alternative to detailed botanical inventories because species frequencies approximate fractional cover when a sufficiently large number of points are sampled within a plot. For approximating fractional cover, it is good practice to choose random points (e.g., using the point frame method; Heady and Rader 1958; Jonasson 1988) and not random individuals within plots, to avoid overrepresenting species with more lateral growth. When botanical inventories are available, stratified random sampling within plots with plant species as strata followed by abundance weighting based on species fractional cover or biomass is a good way for capturing vegetation composition and for scaling traits of individuals to plot-level estimates. In plant communities where species abundances are unequally distributed, it is important to think about the pros and cons of sampling all species vs. sampling the most abundant species and of sampling all species at the same frequency vs. sampling more abundant species at higher frequencies (Table 15.1).
15.3 Field Data Collection
Spectra of plants can be acquired across scales (see Gamon et al., Chap. 16), including at the leaf level, using proximal RS techniques (e.g., handheld spectrometers, robotic systems, UASs) , airborne instruments, and satellite systems. For leaf-level studies, it is often of interest to collect information about taxonomic identity (species or clade), functional type (e.g., based on life form, growth form, dispersal type), functional traits (e.g., based on samples for chemical or structural analysis, growth measurements), developmental stage, and stress symptoms (e.g., signs of disease, herbivory, drought). For canopy-level studies, it is common to collect information about community composition and cover, spatial arrangement (or clustering), gap fractions, plant and canopy architecture (e.g., leaf area index, leaf angle distribution, branching structure, stem diameter, stratification), community biomass, and community traits. Additional data often collected together with vegetation spectra include soil characteristics (e.g., chemistry, water content), elevation, slope, and aspect. Important metadata include time and precise location, observer, nomenclature used, and photos, from which, for example, cover fractions can be estimated. Ideally data are recorded digitally to avoid the time and sources of error associated with transcriptions, and it is good practice to develop and test protocols for standardized data collection. Information should always be recorded as precisely as possible. For example, in grasslands it would be unnecessary to record vegetation height in classes because recording vegetation height at the cm level takes about the same amount of time, and classes can always be aggregated later if needed. Working together with other research groups can make it more efficient to collect additional data. This requires coordination at an early planning stage.
Offering educational opportunities might be part of the mission of a research area, and site administrators might be able to help with hiring students or technicians. However, it is advisable to focus on collecting the most ecologically relevant data, using well-trained personnel and sound methods, including appropriate sampling design and large enough sampling size, rather than collecting various kinds of data of poorer quality. For studies with a RS element, it is important to acquire accurate and precise coordinates of research plots and/or individuals to match their locations to the image data. Triangulation can be used to estimate plot coordinates from ground control points, and relative positions of individuals within plots can be estimated from plot coordinates. The level of accuracy and precision needed depends on the spatial resolution of the imagery, but professional surveying equipment can be needed. Again, early planning is important, because finding rental equipment can become difficult during peak season. Purchasing insurance for expensive equipment might be advisable. Research areas might have periodic surveying campaigns. Including research plots in such campaigns is a great option but requires marking plots temporarily; posts made out of a light but rot-resistant wood (e.g., larch, spruce) are well suited for this.
The following sections give some examples about spectral data acquisition at different levels of measurement; for details on the collection of ecological, non-spectral data, see textbooks on ecological methods (e.g., Sala et al. 2000; van der Maarel and Franklin 2012). As mentioned in the introduction (Sect. 15.1), the choice of methods will depend on the research question, the site conditions, equipment, and personnel available, among other things. There are many good protocols available (see Sect 15.1); familiarizing oneself with a couple of options and their advantages and limitations and testing them under specific scenarios is generally good practice.
15.3.1 Leaf-Level Spectroscopy
A typical setup for leaf-level spectroscopy consists of a spectrometer, light source, fiber-optic cable, leaf clip or integrating sphere, and user interface. Leaf-level spectrometers can be classified into VNIR instruments, usually covering the visible to the beginning of the near-infrared (NIR) portion of the electromagnetic spectrum (~ 350–1000 nm), and full-range instruments, covering additional wavelengths in the NIR and the shortwave-infrared (~350–2400 nm). Generally, VNIR instruments use a silicon array detector, which does not require cooling, making VNIR instruments relatively light and easy to carry. Full-range instruments use additional indium gallium arsenide (InGaAs) photodiodes, which require cooling, to detect the longer wavelengths in the less energetic infrared part of the spectrum, making instruments heavier and less stable. The conditions at the field site should be kept in mind when choosing an instrument. If the spectrometer needs to be carried for longer times and does not come with its own backpack, some extra effort is required to figure out a good packing solution, especially for the fiber-optic cable, which can be easily damaged. It is good practice to check with the instrument companies if warranties are still valid when instruments are transported without their shipping cases; additional insurance might be worth considering.
A number of leaf clips are commercially available; some are easier for one person to handle and/or better suited to measure narrow leaves, such as conifer needles and grass blades, than others, and there is room for design improvements (e.g., using 3D printers). Leaf clip measurements can be used to calculate reflectance (the ratio of detected to incident light), which most instruments do internally, while measurements with integrating spheres can be used to calculate reflectance, transmittance, and absorptance. Leaf clip measurements are generally faster. Ideally, leaves should cover the entire field of view of the sensor; special protocols are available for narrow leaves (Noda et al. 2013). It is important to note that measurements with different setups and among different instruments cannot be directly compared (Hovi et al. 2017). One relatively laborious way to make measurements comparable (e.g., to include data from different instruments in one study) would be estimating empirical transfer functions. This requires measuring the same leaf samples with the instrument setups to be compared. Empirical transfer functions can be estimated for each wavelength and applied to transform measurements from one spectrometer and setup to the other, given that measurement conditions are comparable between model development and application.
Typically, measurements start with assembling the spectrometer, fiber-optic cable, light source, and leaf clip or integrating sphere, turning on the instrument and light source, and giving them some time (e.g., around 15–30 minutes) to warm up and stabilize. Meanwhile, the user interface can be connected to the spectrometer and folders can be organized. For instruments that are operated from a bench, it is important to find a stable position; ideally, neither the instrument nor the fiber-optic cable should be moved between measurements. The same applies to instruments operated from a backpack; the setup should be as consistent and stable as possible. To avoid damage to the fibers, it is important to avoid bending fiber-optic cables, including at connections between the fibers and the instrument. Generally, it is good practice to use and transport fiber-optic cables as stretched out as possible. A good option is coiling longer fiber-optic cables loosely while allowing for enough play at connections between cable and instruments and between cable and leaf clip/integrating sphere to keep angles around 180°. It is important to keep the fiber-optic cable away from branches when walking through vegetation, because it can be snagged and broken.
One way to test if the instrument has warmed up and is stabilized is to measure an invariant surface, such as a reflectance standard. Before starting sample measurements, it is time to take reference measurements, including measurements of so-called dark current (the background signal from the instrument), which some instruments take automatically, and white references (materials that approximate Lambertian surfaces, which reflect light at all angles equally or are perfectly diffuse). Dark current measurements correct for instrument noise, while white reference measurements determine the light entering an instrument and allow the calculation of reflectance (and transmittance). White references are usually made of polytetrafluoroethylene, better known by its commercial name Spectralon. They are available in different sizes and shapes; some leaf clips have built-in white references, but they are difficult to keep clean. It is important to keep white references as clean as possible, because even small traces of dirt and oil affect the spectral signal. Reference panels should only be held by their sides (touching the surface should be avoided), and they should be covered after each measurement. Depending upon usage and field conditions, frequent cleaning according to the manufacturer’s instructions may be necessary.
Generally, a white reference reading should be made before the first measurement and whenever conditions (e.g., temperature, the arrangement of the fiber-optic cable, instrument, or lamp settings) change. However, it might be more practical or accurate to take white reference measurements at regular intervals, such as every 10 minutes, for each sample, or after a certain number of measurements, and to take additional measurements when needed. It is good practice to plot the reflectance spectrum of each white reference measurement and to save the spectrum. The reflectance of the white reference should be around 100% for all wavelengths except for the beginning and end of the spectrum, which are generally noisier; deviations from 100% or excessive noise can indicate a dirty panel, or issues with the cooling system, lamp, instrument setup, or a low battery. Measurements should be taken immediately after sample collection, because leaves dry out quickly. For most purposes, if samples need to be stored before measurement, they should be kept cool, moist, and dark and measured as soon as possible. It is good practice to keep the intensity of the light source at the minimum needed for a good signal-to-noise ratio, because leaves can get burned by the lamp. Defining a threshold for a “good” measurement, for example, based on the reflectance at the so-called NIR shoulder, which is the highest point of the reflectance curve at the beginning of the NIR, can be helpful. Spectral measurements should be made under dry conditions; instruments can be damaged by water, and water films alter the spectral characteristics of leaves.
Measurement protocols should specify how many leaves per plant to measure and which leaves to select. Generally, this means clarifying if a study deals with “ideal” or “average” plant individuals and if an entire plant should be characterized or only certain layers, such as the top canopy. Measuring mature, healthy, sunlit leaves is a good strategy for characterizing species or functional groups. For studies dealing with disease detection, asymptomatic and symptomatic leaves should be measured, ideally at different stages of the disease. Selecting leaves at random, including all ages, canopy layers, and stress levels, can make sense when aiming to relate leaf chemistry to ecosystem processes or when scaling leaf-level chemistry to plot-level estimates. If the aim is to characterize entire individuals, leaves from all canopy layers can be included, with the number of measurements per layer reflecting plant size, growth form, and architectural complexity. However, when leaf-level spectra are being matched to spectra acquired with RS, it makes sense to select only leaves from the layer that is captured by the sensor (i.e., from the top of the canopy). If possible, measuring the midvein should be avoided. Measuring different spots on the same leaf is also usually unnecessary, at least for small- to medium-sized leaves, because spectral variation at the leaf level is generally small. However, it is important to check the quality of every spectrum. Ideally, quality checks are done immediately after each measurement; bad measurement can be flagged for subsequent filtering, which considerably reduces preprocessing time.
15.3.2 Proximal Canopy-Level Spectroscopy
Proximal canopy spectra can be sampled with handheld spectrometers, robotic systems, and UASs. One important differentiation is between nonimaging and imaging systems. Nonimaging spectrometers integrate over a defined amount of time the spectral reflectance of an illuminated area; the output is one spectral curve per measurement. Instruments used for leaf-level spectroscopy (Sect. 15.3.1) fall into this category. The same instruments can be used in a handheld mode or mounted on a platform to sample spectra at the proximal canopy level, and the reach of instruments can be expanded using long fiber-optic cables attached to a beam. Creative solutions include mounting a spectrometer on a bike and using it as a mobile platform (see “reflectomobile” in Milton et al. 2009). Imaging spectrometers sample spectra in a spatially resolved fashion. The collected data are commonly represented as data cubes, with x- and y-axes representing the spatial extent of the imaged area and the z-axis representing the spectral response measured across the wavelength range (Vane and Goetz 1988). Commonly used systems are push-broom and whisk-broom imagers, which are usually operated from a moving platform. Alternatively, the imaged scene can move while the imaging spectrometer scans the samples, which is, for example, possible using conveyor-belt-like setups in the laboratory.
It is important to know the footprint, or ground field of view (GFOV), of remote sensing instruments. For nonimaging systems, the footprint equals the measured area on the ground; for imaging systems, it determines image and pixel size. The footprint depends on the field of view (FOV) of the sensor and the distance between sensor and measured object (h), and it is calculated as GFOV = tan(FOV/2) ∗ h ∗ 2 (Fig. 15.4). Foreoptic lenses can be used to narrow or expand an instrument’s FOV, which is particularly relevant for handheld nonimaging systems. For spectrometers operated using robotic systems or UASs, the footprint is typically regulated by adjusting the height of the sensor above the ground; the farther away, the larger the GFOV, imaged area, and pixel size.
Handheld spectrometers and accessories need to be sturdy and easy to carry, particularly when collecting data over longer periods of time and in areas that cannot be reached by car. Spectrometers covering the VNIR range of the electromagnetic spectrum are usually small, such that neck straps securely attached to the instrument are often sufficient. Full-range spectrometers are heavier and typically need to be fit into a backpack, which means that cable connections have to be robust; the entrance point of the fiber-optic cable in particular needs to be flexible, yet stable. Furthermore, it is important that the fiber-optic cable can be stored securely during transportation to avoid damaging the fibers and that the instrument is properly ventilated to avoid overheating. Changing the orientation of the fiber-optic cable changes the amount of light reaching the detector. Thus, it is good practice to ensure that the entire setup can be assembled easily in the same way every time. For spectrometers that are being used outside, the exterior should be made of materials that do not easily overheat when exposed to sunlight; polymeric surface films that provide radiative cooling (Zhai et al. 2017) could reduce the problem of overheating.
Often it is easier to collect data with handheld spectrometers in two-person teams, one person carrying the instrument and measuring and the other operating the computer and checking the data. However, it can be advantageous to have a system in place allowing one person to operate the spectrometer alone when needed. Vendor trays are a good option for carrying laptops while taking measurements. Small handheld devices can be very useful for collecting data, but their small screens make data checks difficult, and it can be impractical to name and rename files on small devices. Instrument software using voice control would be an advantage in this regard. Furthermore, when one person is operating a portable spectrometer, the white reference needs to be securely stored and in reach of the user. It is also important that backpacks fit comfortably, which means adjustable straps, cushioning, and ventilation. As with leaf-level spectrometers, instrument and light source should be switched on at least 15–30 minutes before data collection to allow the instrument to warm up and stabilize. Like other passive RS systems, handheld spectrometers should ideally be operated under stable illumination conditions, which is rarely possible. However, clear sky and no haze are a good place to start. Reference measurements (white reference and, if needed, dark current) should be taken before the first measurement and whenever illumination or temperature changes (after breaks, when adjusting the backpack, changing the sampling position, etc.). Again, it is good practice to take references at regular intervals, e.g., every 10 minutes, because of changing solar angle, ambient temperature, and sky conditions and to take additional references when needed. During measurements, the entrance optic of handheld systems should be positioned as far away from the body as possible (i.e., by stretching out the arm holding the fiber cable) to avoid measuring one’s own shadow. In flat terrain, measurements should be taken in nadir position (i.e., with the fiber optic pointing directly down). In steep terrain, it is advantageous to point the fiber optic at a right angle toward the slope. It is good practice to wear nonreflective clothing and shoes in dark shades as stray reflected light off clothing can affect the spectral measurements.
At midlatitudes, the best time for measurements, given clear sky, is around solar noon (i.e., when the sun is at its highest point in the sky), whereas in the Arctic and the Tropics, time of day is usually less important than sky conditions. Generally, at midlatitudes, spectral measurements should be taken when the sun elevation angle (the angle measured from the horizon to the center of the sun) exceeds 45°; angles exceeding around 50° are better. For example, a good time window for canopy spectroscopy (sun angle >45°) for Minneapolis, MN, on June 21 would be from around 10:00 am to 4:30 pm. At midlatitudes, the longest time window for sun elevation angles greater than certain angles occurs around summer solstice, while around the equator, it is around the equinox. Calculators for sun position can be found online, such as from the National Oceanic and Atmospheric Administration (NOAA, https://www.esrl.noaa.gov/gmd/grad/solcalc/).
15.3.3 Airborne Campaigns
This section covers flight planning and some aspects of reference data collection for image processing. Remote sensing of plant biodiversity typically involves matching ecological and spectral data collected on the ground to remotely sensed images. Key aspects to match ground and remote sensing data include choosing vegetation plots that match the spatial resolution of the imagery (i.e., plots that are representative for at least one pixel) and collecting accurate coordinate information at a relevant precision for the remote sensor; this and more is covered in Sect. 15.2.3 and the beginning of Sect. 15.3.
15.3.3.1 Flight Planning
Flight planning for airborne imaging spectroscopy campaigns starts with deciding on the best time window(s) for the flight(s). The ideal time depends on the research question, but generally it is when the phenomena of interest are most pronounced. For example, for modeling and mapping aboveground productivity, peak biomass could be a good time for acquiring airborne images; for differentiating plant species, early or late growing season could be the times of year when certain species are most distinctive; for modeling and mapping plant disease or plant stress, different symptoms could be expressed at specific times of the year.
Schedules for flight crews are usually tight; thus communicating ideal flight windows early (i.e., at least several months in advance) is important. It is also critical to communicate flight windows to site administrators as soon as possible (see Sect. 15.2.2). On the one hand, it might be important to limit disturbance (e.g., trampling or destructive sampling) at the site during the week(s) leading up to a flight and to have no people and/or equipment on-site during the time of the overflight. On the other hand, other research groups might be interested in data collection around the time of the overflight. Ideally, airborne data are collected under clear sky conditions and low humidity. It is good practice to take the typical weather conditions at the site into account and plan flights at a time of year with generally good sky conditions, if possible.
Typically, the next steps of flight planning include determining the desired pixel size and drawing the flight lines. The ideal pixel size depends, again, on the goal of the study and the study system. Image pixels can be larger for modeling biomass and chemical composition at the plant community level than for predicting functional traits or species identity of individuals. The desired pixel size (within the limits of instrument and platform) determines flight altitude and swath width. Airplanes need to fly lower and slower to acquire images with smaller pixels resulting in narrower flight lines. Thus, there is a trade-off between pixel size and the area covered with a single flight line or during one flight. Generally, flight lines should overlap 15–25% to ensure full area coverage. Flight lines can be stitched together in a process called mosaicking, but spectra from the same pixels from different flight lines vary, because of changing sun angles and atmospheric conditions over the course of the day. In part, these differences can be accounted for during atmospheric correction, but not perfectly. Given inevitable constraints, covering a research area in fewer flight lines or a single line and working with larger pixels could be an option, depending on the phenomenon of interest.
At midlatitudes, flights approximately ±2 h from solar noon are often ideal; solar noon times can be looked up, for example, on the NOAA website (https://www.esrl.noaa.gov/gmd/grad/solcalc/). When images are acquired around solar noon, flight lines are often oriented in a north-south direction to align the flight direction with the principle plane of the sun and to prevent the plane from casting a shadow on the image. However, in areas such as the Tropics, it might be better to fly in the morning or afternoon because of cloud formation during the middle of the day. In the morning, flight lines oriented southeast-northwest are a good option; in the afternoon, flight lines oriented southwest-northeast may be preferable. The sun azimuth angle (i.e., the angle between the sun’s position and north along the horizon: north equals 0°, east equals 90°, etc.) can be used as flight-line bearings. Additionally, it is seldom possible for flight crews to commit to specific dates and exact times, so the sun azimuth angle for the approximate time of overpass is generally fine as a bearing. Drawing separate sets of flight lines for different times of day is also an option. Again, information on sun azimuth angles for specific dates, times, and locations can be found online. During the time window of the light, it is often a judgment call if sky conditions are “good enough” for image acquisition; the costs of having the plane, air, and ground crews wait for a delay are important factors to consider. Generally, although cumulus clouds obscure parts of the image, they are preferable over cirrus or stratus clouds, which keep changing illumination conditions resulting in overall low-quality image data.
15.3.3.2 Reference Data Collection for Image Processing
In summary, the most important steps in image processing are:
-
Radiometric correction: Sensors record electromagnetic radiation in digital numbers (DNs). During radiometric correction, DNs are converted to at-sensor radiance using sensor- and pixel-specific radiation sensitivity coefficients. Information about the sun’s geometry, including Earth-sun distance and solar angle, is used to convert at-sensor radiance to top of the atmosphere (TOA) reflectance.
-
Spectral correction: Every pixel contains spectral information recorded at slightly different wavelengths, which are standardized to a common wavelength interval in this step.
-
Atmospheric correction: Atmospheric correction transforms TOA reflectance to surface reflectance using information about atmospheric conditions and aerosol properties at the time of image acquisition. This can be done with atmospheric radiative transfer models (e.g., MODTRAN, Berk et al. 1987), some of which are included in image processing software; dark subtraction, the subtraction of values from dark image pixels; or ELC (explained below).
-
Geometric correction (including geometric resampling, orthorectification, and georeferencing): Geometric resampling makes pixels square (initially they are elliptic). Orthorectification corrects image distortions caused by sensor tilt, flight altitude, and changes in surface terrain, creating planimetric images, which allow direct measurement of distances, areas, and angles. Geometric resampling and orthorectification require information about camera properties, the sensor position recorded by the inertial measurement unit (IMU), and an accurate digital elevation model (DEM), which provides information about terrain height (elevation above sea level). Georeferencing aligns images to a specific coordinate system. This is frequently done using ground control points (GCPs).
Ground reference data collected for image processing frequently include GCPs for georeferencing and reference spectra for image calibration/validation procedures and atmospheric correction with ELC. Generally, GCPs need to be easily identifiable in the acquired images; road intersections, corners of buildings, and trees are good choices. Accurate and precise coordinates of GCPs can be either determined from existing georeferenced imagery, in which case it is good practice to check if the features still exist, or measured on the ground. For ELC, the reflectance of large calibration targets on the ground is measured with a handheld spectrometer during the overflight. At-sensor radiance is transformed to ground-level reflectance by applying bandwise transformation coefficients estimated based on the difference between sensor and calibration target spectra (Smith and Milton 1999). Essentially, this subtracts atmospheric influences from the spectra recorded by the sensor. Calibration targets should meet several requirements. They need to be identifiable in the images; when in doubt, location data should be collected. Furthermore, calibration targets should allow the extraction of several pixels from the image, so they should be sufficiently large (e.g., targets measuring 7 × 7 pixels allow using 5 × 5 pixels from the image after removing the pixels at the edges). In addition, calibration targets should be Lambertian as possible (i.e., they should reflect light equally in all directions, independent of viewing angle). Ideally, calibration targets should include the range of values in the image, which means including targets with low and high reflectance. Good options are calibration tarps in different shades (e.g., white, gray, and black) that are as spectrally invariant as possible. Tarps can be made of boat canvas (e.g., acrylic-coated woven polyester) and should include grommets, so that they can be tightly pulled and secured with pegs. Calibration tarps should be placed in flat areas with short vegetation. Ideally, one set of tarps is placed in the center of each flight line and measured exactly at the time of overflight, such that each flight line can be corrected separately. However, when multiple sets of tarps are not an option, a single set of tarps can be placed in a central area instead. Other surfaces can be used for ELC, including water bodies, road intersections, rooftops, snow, and ice. Ideally, these targets are distributed throughout the image and are measured at the time of the overpass. However, it might be difficult to find already existing calibration targets that are spectrally stable and uniform; for example, the reflectance of lakes can vary substantially depending on the distance from the shore and currents; snow and ice are often less spectrally uniform than expected due to surface irregularities, dust, and algae growth; and asphalt concrete varies spectrally depending on the aggregate composition (bitumen varies less). If possible, limiting movement and traffic around calibration targets during measurements and image acquisition is advantageous for reducing the amount of dust and dirt (e.g., when using calibration tarps or roads), as well as limiting surface disturbance and shadow cast (e.g., when using snow). However, prioritizing calibration target quality over quantity is good practice. For example, measuring one set of spectrally stable tarps in a flat area in the center of an image is preferable over measuring several natural calibration targets that are likely not as spectrally uniform and might be difficult to locate precisely.
15.4 Conclusions
As ecologists and remote sensing scientists are coming together to develop methods for the continuous assessment and monitoring of plant biodiversity, connecting the local to the global scale, studies of species to communities and ecosystems, and ecological resources to human needs and values become more and more feasible. Field campaigns are essential to this effect, because only (the repeated process of generating) ecological knowledge and data (including spectral measurements of plants) make it possible to understand better what is happening to the ecosystems and species we care about and why. Thanks to the ecological data revolution, remote sensing and organismal data as well as climate, land use, and socioeconomic data are becoming increasingly publicly available. At the same time, bioinformatics and cyberinfrastructure promote innovative ways for data handling, storage, and visualization and for integrating and analyzing these data across scales. Harnessing this amount of data requires developing and documenting data standards that facilitate collaborations across disciplines, data integration across sites and scales, data discovery for meta-analysis, and model re-calibrations. This makes the planning of data collection with consideration of the data life cycle as important as the data collection itself. It is important in the face of our current and future challenges and opens a wealth of opportunities in biodiversity science.
References
Anderson TM, Hopcraft JGC, Stephanie E, Ritchie M, Grace JB, Olff H (2010) Landscape-scale analyses suggest both nutrient and antipredator advantages to serengeti herbivore hotspots. Ecology 91:1519–1529
Atkinson PM (1999) Spatial statistics. In: Stein A, Van Der Meer F, Gorte B (eds) Spatial statistics for remote sensing. Kluwer Academic Publishers, Dortrecht
Baddeley A, Turner R (2005) Spatstat: an r package for analyzing spatial point patterns. J Stat Softw 12:1–42
Bartholomé E, Belward AS (2005) Glc2000: a new approach to global land cover mapping from earth observation data. Int J Remote Sens 26:1959–1977
Berk A, Bernstein LS, Robertson DC (1987) Modtran: a moderate resolution model for lowtran. Spectral Sciences Inc, Burlington
Beyer HL (2010) Geospatial modelling environ
Bicheron P, Huc M, Henry C, Bontemps S, Lacaux J (2008) Globcover products description manual. European Space Agency, Paris
Bonham CD (2013) Measurements for terrestrial vegetation. Wiley, Hoboken
Bonn A, Storch D, Gaston KJ (2004) Structure of the species–energy relationship. Proc R Soc Lond Ser B Biol Sci 271:1685–1691
Broge NH, Leblanc E (2001) Comparing prediction power and stability of broadband and hyperspectral vegetation indices for estimation of green leaf area index and canopy chlorophyll density. Remote Sens Environ 76:156–172
Brown JH (1981) Two decades of homage to Santa Rosalia: toward a general theory of diversity. Am Zool 21:877–888
Brus DJ, Degruijter JJ (1993) Design-based versus model-based estimates of spatial means: theory and application in environmental soil science. Environmetrics 4:123–152
Cavender-Bares J, Gamon JA, Hobbie SE, Madritch MD, Meireles JE, Schweiger AK, Townsend PA (2017) Harnessing plant spectra to integrate the biodiversity sciences across biological and 1391 spatial scales. Am J Bot 104:966–969
Chollet F, Allaire JJ (2018) Deep learning with R, 1st edn. Manning Publications, Shelter Island, New York
Chun Y, Griffith DA (2013) Spatial statistics and geostatistics: theory and applications for geo- 1393 graphic information science and technology. Sage Publications, London
Clark M, Roberts D, Clark D (2005) Hyperspectral discrimination of tropical rain forest tree spe- 1395 cies at leaf to crown scales. Remote Sens Environ 96(3–4):375–398
Claw KG, Anderson MZ, Begay RL, Tsosie KS, Fox K, Garrison NA (2018) A framework for enhancing ethical genomic research with indigenous communities. Nat Commun 9(1):2957
Combal B, Baret F, Weiss M, Trubuil A, Mace D, Pragnere A, Myneni R, Knyazikhin Y, Wang L (2003) Retrieval of canopy biophysical variables from bidirectional reflectance: using prior information to solve the ill-posed inverse problem. Remote Sens Environ 84:1–15
Congalton RG, Green K (1999) Assessing the accuracy of remotely sensed data: principles and 1402 applications. Lewis Publishers, Boca Raton
Congalton RG, Gu J, Yadav K, Thenkabail P, Ozdogan M (2014) Global land cover mapping: a review and uncertainty analysis. Remote Sens 6:12070–12093
Cook RB, Wei Y, Hook LA, Vannan SK, Mcnelis JJ (2018) Preserve: protecting data for long-term use. In: Ecological informatics. Springer, Berlin, Heidelberg
Coops NC, Kearney SP, Bolton DK, Radeloff VC (2018) Remotely-sensed productivity clusters capture global biodiversity patterns. Sci Rep 8:16261
Corti L, Van Den Eynden V, Bishop L, Woollard M (2014) Managing and sharing research data: a guide to good practice. Sage Publications, London
Couture JJ, Singh A, Rubert-Nason KF, Serbin SP, Lindroth RL, Townsend PA (2016) Spectroscopic determination of ecologically relevant plant secondary metabolites. Methods Ecol Evol 7:1402–1412
Dahabreh IJ, Chan J, Earley A, Moorthy D, Avendano E, Trikalinos T, Balk E, Wong J (2017) Modeling and simulation in the context of health technology assessment: review of existing guidance, future research needs, and validity assessment. Technical report, Rockville: Agency for Healthcare Research and Quality (US). Available from: https://www.ncbi.nlm.nih.gov/books/NBK424024/
De Gruijter J (1999) Spatial sampling schemes for remote sensing. In: Spatial statistics for remote sensing. Springer, Berlin, Heidelberg
Domburg P, De Gruijter JJ, Van Beek P (1997) Designing efficient soil survey schemes with a knowledge-based system using dynamic programming. Geoderma 75:183–201
Evans JS (2017) Spatialeco. R package version 2:0–0
Fassnacht FE, Latifi H, Stereńczak K, Modzelewska A, Lefsky M, Waser LT, Straub C, Ghosh A (2016) Review of studies on tree species classification from remotely sensed data. Remote Sens Environ 186:64–87
Féret J-B, Asner GP (2014) Mapping tropical forest canopy diversity using high-fidelity imaging spectroscopy. Ecol Appl 24:1289–1296
Fleishman E, Seto KC (2009) Applications of remote sensing to ecological modeling [online]. Available: https://ncep.amnh.org/index.php/Detail/objects/36. Accessed 26 Nov 2018
Gamon J, Penuelas J, Field C (1992) A narrow-waveband spectral index that tracks diurnal changes in photosynthetic efficiency. Remote Sens Environ 41:35–44
Gamon JA, Cheng Y, Claudio H, Mackinney L, Sims DA (2006) A mobile tram system for systematic sampling of ecosystem optical properties. Remote Sens Environ 103:246–254
Gastellu-Etchegorry J, Martin E, Gascon F (2004) Dart: a 3d model for simulating satellite images and studying surface radiation budget. Int J Remote Sens 25:73–96
Goetz AFH, Vane G, Solomon JE, Rock BN (1985) Imaging spectrometry for earth remote sensing. Science 228:1147–1153
Gould W (2000) Remote sensing of vegetation, plant species richness, and regional biodiversity hotspots. Ecol Appl 10:1861–1870
Grabau WE, Rushing WN (1968) A computer-compatible system for quantitatively describing the physiognomy of vegetation assemblages. In: Land evaluation. Macmillan of Australia, Melbourne, pp 263–275
Heady HF, Rader L (1958) Modifications of the point frame. Rangeland Ecol & Manag/J Range Manag Arch 11:95–96
Herrmann I, Vosberg S, Ravindran P, Singh A, Chang H-X, Chilvers M, Conley S, Townsend P (2018) Leaf and canopy level detection of fusarium virguliforme (sudden death syndrome) in soybean. Remote Sens 10:426
Horning N, Robinson JA, Sterling EJ, Turner W, Spector S (2010) Linking remote sensing with modeling. In: Remote sensing for ecology and conservation: a handbook of techniques. Oxford University Press, Oxford
Hovi A, Forsström P, Mõttus M, Rautiainen M (2017) Evaluation of accuracy and practical applicability of methods for measuring leaf reflectance and transmittance spectra. Remote Sens 10:25
Huemmrich K (2001) The geosail model: a simple addition to the sail model to describe discontinuous canopy reflectance. Remote Sens Environ 75:423–431
Huete AR (1988) A soil-adjusted vegetation index (savi). Remote Sens Environ 25:295–309
Inamdar D, Kalacska M, Leblanc G, Arroyo-Mora JP (2020) Characterizing and mitigating sensor generated spatial correlations in airborne hyperspectral imaging data. Remote Sens 12:641
Jacquemoud S, Baret F (1990) Prospect: a model of leaf optical properties spectra. Remote Sens Environ 34:75–91
Jacquemoud S, Verhoef W, Baret F, Bacour C, Zarco-Tejada PJ, Asner GP, François C, Ustin SL (2009) Prospect+ sail models: a review of use for vegetation characterization. Remote Sens Environ 113:S56–S66
Johannsen CJ, Daughtry CS (2009) Surface reference data collection. In: Warner T, Nellis M, Foody G (eds) The SAGE handbook of remote sensing. Sage Publications, London
Jonasson S (1988) Evaluation of the point intercept method for the estimation of plant biomass. Oikos 52:101–106
Jones R (2009) Categories, borders and boundaries. Prog Hum Geogr 33:174–189
Jones HG, Vaughan RA (2010) Remote sensing of vegetation: principles, techniques, and applications. Oxford University Press, Oxford
Justice CO, Townshend JR (1981) Integrating ground data with remote sensing. In: Townshend JRG (ed) Terrain analysis and remote sensing. Allen & Unwin, London
Kattenborn T, Eichel J, Fassnacht FE (2019) Convolutional neural networks enable efficient, accurate and fine-grained segmentation of plant species and communities from high-resolution UAV imagery. Sci Rep 9:17656
Kattge J, Diaz S, Lavorel S, Prentice I, Leadley P, Bönisch G, Garnier E, Westoby M, Reich PB, Wright I (2011) Try–a global database of plant traits. Glob Chang Biol 17:2905–2935
Kincaid T, Olsen A (2016) Spsurvey: spatial survey design and analysis. R package version 3:3
Kruse F, Kierein-Young K, Boardman J (1990) Mineral mapping at cuprite, Nevada with a 63-channel imaging spectrometer. Photogramm Eng Remote Sens 56:83–92
Lopatin J, Fassnacht FE, Kattenborn T, Schmidtlein S (2017) Mapping plant species in mixed grassland communities using close range imaging spectroscopy. Remote Sens Environ 201:12–23
Martens H (2001) Reliable and relevant modelling of real world data: a personal account of the development of pls regression. Chemom Intell Lab Syst 58:85–95
McCoy RM (2005) Field methods in remote sensing. The Guilford Press, New York
Milton EJ, Schaepman ME, Anderson K, Kneubühler M, Fox N (2009) Progress in field spectroscopy. Remote Sens Environ 113:S92–S109
Noda HM, Motohka T, Murakami K, Muraoka H, Nasahara KN (2013) Accurate measurement of optical properties of narrow leaves and conifer needles with a typical integrating sphere and spectroradiometer. Plant Cell Environ 36:1903–1909
Papritz A, Stein A (1999) Spatial prediction by linear kriging. In: Spatial statistics for remote sensing. Springer, Berlin, Heidelberg
Perez-Harguindeguy N, Diaz S, Garnier E, Lavorel S, Poorter H, Jaureguiberry P, Bret-Harte M (2013) New handbook for standardised measurement of plant functional traits worldwide. Australian J 800:167–234
Pettorelli N, Ryan S, Mueller T, Bunnefeld N, Jedrzejewska B, Lima M, Kausrud K (2011) The normalized difference vegetation index (ndvi): unforeseen successes in animal ecology. Clim Res 46:15–27
Pontius J, Hallett R, Martin M (2005) Using aviris to assess hemlock abundance and early decline in the catskills, New York. Remote Sens Environ 97:163–173
Rouse JW Jr, Haas RH, Deering DW, Schell JA, Harlan JC (1974) Monitoring the vernal advancement and retrogradation (green wave effect) of natural vegetation. NASA/GSFC type III final report, Greenbelt. Available from: https://ntrs.nasa.gov/search.jsp?R=19750020419
Sala OE, Jackson RB, Mooney HA, Howarth RW (2000) Methods in ecosystem science: progress, tradeoffs, and limitations. Springer, New York
Schaepman ME (2007) Spectrodirectional remote sensing: from pixels to processes. Int J Appl Earth Obs Geoinf 9:204–223
Schmidtlein S, Feilhauer H, Bruelheide H (2012) Mapping plant strategy types using remote sensing. J Veg Sci 23:395–405
Schweiger AK, Risch AC, Damm A, Kneubühler M, Haller R, Schaepman ME, Schütz M (2015a) Using imaging spectroscopy to predict above-ground plant biomass in alpine grasslands grazed by large ungulates. J Veg Sci 26:175–190
Schweiger AK, Schütz M, Anderwald P, Schaepman ME, Kneubühler M, Haller R, Risch AC (2015b) Foraging ecology of three sympatric ungulate species - behavioural and resource maps indicate differences between chamois, ibex and red deer. Mov Ecol 3:6
Schweiger AK, Schütz M, Risch AC, Kneubühler M, Haller R, Schaepman ME (2017) How to predict plant functional types using imaging spectroscopy: linking vegetation community traits, plant functional types and spectral response. Methods Ecol Evol 8:86–95
Serbin SP, Singh A, Mcneil BE, Kingdon CC, Townsend PA (2014) Spectroscopic determination of leaf morphological and biochemical traits for northern temperate and boreal tree species. Ecol Appl 24:1651–1669
Sims DA, Gamon JA (2002) Relationships between leaf pigment content and spectral reflectance across a wide range of species, leaf structures and developmental stages. Remote Sens Environ 81:337–354
Singh A, Serbin SP, Mcneil BE, Kingdon CC, Townsend PA (2015) Imaging spectroscopy algorithms for mapping canopy foliar chemical and morphological traits and their uncertainties. Ecol Appl 25:2180–2197
Smith GM, Milton EJ (1999) The use of the empirical line method to calibrate remotely sensed data to reflectance. Int J Remote Sens 20:2653–2662
Stehman SV (2001) Statistical rigor and practical utility in thematic map accuracy assessment. 1525. Photogramm Eng Remote Sens 67:727–734
Stehman S, Foody G (2009) Accuracy assessment. In: Warner T, Nellis M, Foody G (eds) The SAGE handbook of remote sensing. Sage Publications, London
Stein A, Van Der Meer FD, Gorte B (2002) Spatial statistics for remote sensing. Kluwer Academic Publishers, Dordrecht
Szary ALA (2015) Boundaries and borders. In: The Wiley Blackwell companion to political geography
Thenkabail PS, Lyon JG, Huete A (2012) Hyperspectral remote sensing of vegetation. CRC Press, Boca Raton
Theobald DM, Stevens DL, White D, Urquhart NS, Olsen AR, Norman JB (2007) Using gis to generate spatially balanced random survey designs for natural resource applications. Environ Manag 40:134–146
Tucker CJ (1979) Red and photographic infrared linear combinations for monitoring vegetation. Remote Sens Environ 8:127–150
Ustin SL, Gamon JA (2010) Remote sensing of plant functional types. New Phytol 186:795–816
Ustin SL, Gitelson AA, Jacquemoud S, Schaepman ME, Asner GP, Gamon JA, Zarco-Tejada P (2009) Retrieval of foliar information about plant pigment systems from high resolution spectroscopy. Remote Sens Environ 113:S67–S77
van der Maarel E, Franklin J (2012) Vegetation ecology. Wiley-Blackwell, Chichester
Vane G, Goetz AFH (1988) Terrestrial imaging spectroscopy. Remote Sens Environ 24:1–29
Verhoef W (1984) Light scattering by leaf layers with application to canopy reflectance modeling: the sail model. Remote Sens Environ 16:125–141
Verrelst J, Camps-Valls G, Muñoz-Marí J, Rivera JP, Veroustraete F, Clevers JGPW, Moreno J (2015) Optical remote sensing and the retrieval of terrestrial vegetation bio-geophysical properties – a review. ISPRS J Photogramm Remote Sens 108:273–290
Vohland M, Jarmer T (2008) Estimating structural and biochemical parameters for grassland from spectroradiometer data by radiative transfer modelling (prospect+ sail). Int J Remote Sens 29:191–209
Walvoort DJ, Brus D, De Gruijter J (2010) An r package for spatial coverage sampling and random sampling from compact geographical strata by k-means. Comput Geosci 36:1261–1267
Wang R, Gamon J, Montgomery R, Townsend P, Zygielbaum A, Bitan K, Tilman D, Cavender-Bares J (2016) Seasonal variation in the ndvi–species richness relationship in a prairie grassland experiment (cedar creek). Remote Sens 8:128
Wang Z, Townsend PA, Schweiger AK, Couture JJ, Singh A, Hobbie SE, Cavender-Bares J (2019) Mapping foliar functional traits and their uncertainties across three years in a grassland experiment. Remote Sens Environ 221:405–416
Warner TA, Nellis MD, Foody GM (2009) The SAGE handbook of remote sensing. Sage Publications, London
Webster R, Oliver MA (1992) Sample adequately to estimate variograms of soil properties. J Soil Sci 43:177–192
Weiss M, Baret F, Myneni R, Pragnère A, Knyazikhin Y (2000) Investigation of a model inversion technique to estimate canopy biophysical variables from spectral and directional reflectance data. Agronomie 20:3–22
Wold S, Martens H, Wold H (1983) The multivariate calibration problem in chemistry solved by the pls method. In: Ruhe A, Kagstrom B (eds) Matrix pencils, lecture notes in mathematics. Springer, Heidelberg, pp 286–293
Wright DH (1983) Species-energy theory: an extension of species-area theory. Oikos 41:496–506
Zhai Y, Ma Y, David SN, Zhao D, Lou R, Tan G, Yang R, Yin X (2017) Scalable-manufactured randomized glass-polymer hybrid metamaterial for daytime radiative cooling. Science 355:1062–1066
Acknowledgments
AKS wishes to thank Susan L. Ustin and Shawn P. Serbin for reviewing this chapter and the National Institute for Mathematical Biology and Synthesis (NIMBioS) working group on remote sensing for stimulating discussions. This work was supported by the NSF/NASA Dimensions of Biodiversity grant “Linking remotely sensed optical diversity to genetic, phylogenetic and functional diversity to predict ecosystem processes” (DEB-1342872) to Jeannine Cavender-Bares.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2020 The Author(s)
About this chapter
Cite this chapter
Schweiger, A.K. (2020). Spectral Field Campaigns: Planning and Data Collection. In: Cavender-Bares, J., Gamon, J.A., Townsend, P.A. (eds) Remote Sensing of Plant Biodiversity. Springer, Cham. https://doi.org/10.1007/978-3-030-33157-3_15
Download citation
DOI: https://doi.org/10.1007/978-3-030-33157-3_15
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-33156-6
Online ISBN: 978-3-030-33157-3
eBook Packages: Earth and Environmental ScienceEarth and Environmental Science (R0)