Abstract
Symbolic Classification (SC), an offshoot of Genetic Programming (GP), can play an important role in any well rounded predictive analytics tool kit, especially because of its so called “WhiteBox” properties. Recently, algorithms were developed to push SC to the level of basic classification accuracy competitive with existing commercially available classification tools, including the introduction of GP assisted Linear Discriminant Analysis (LDA). In this paper we add a number of important enhancements to our basic SC system and demonstrate their accuracy improvements on a set of theoretical problems and on a banking industry problem. We enhance GP assisted linear discriminant analysis with a modified version of Platt’s Sequential Minimal Optimization algorithm which we call (MSMO), and with swarm optimization techniques. We add a user-defined typing system, and we add deep learning feature selection to our basic SC system. This extended algorithm (LDA++) is highly competitive with the best commercially available M-Class classification techniques on both a set of theoretical problems and on a real world banking industry problem. This new LDA++ algorithm moves genetic programming classification solidly into the top rank of commercially available classification tools.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
References
Berthold, M.R., Cebron, N., Dill, F., Gabriel, T.R., Kötter, T., Meinl, T., Ohl, P., Thiel, K., Wiswedel, B.: Knime-the konstanz information miner: version 2.0 and beyond. ACM SIGKDD Explorations Newsletter 11, 26–31 (2009)
Fisher, R.A.: The use of multiple measurements in taxonomic problems. Annals of Human Genetics 7, 179–188 (1936)
Friedman, J.H.: Regularized discriminant analysis. Journal of the American Statistical Association 84, 165–175 (1989)
Ingalalli, V., Silva, S., Castelli, M., Vanneschi, L.: A multi-dimensional genetic programming approach for multi-class classification problems. In: European Conference on Genetic Programming 2014, pp. 48–60. Springer (2014)
Karaboga, D., Akay, B.: A survey: algorithms simulating bee swarm intelligence. Artificial intelligence review 31(1–4), 61 (2009)
Korns, M.F.: A baseline symbolic regression algorithm. In: Genetic Programming Theory and Practice X. Springer (2012)
Korns, M.F.: Extreme accuracy in symbolic regression. In: Genetic Programming Theory and Practice XI, pp. 1–30. Springer (2014)
Korns, M.F.: Highly accurate symbolic regression with noisy training data. In: Genetic Programming Theory and Practice XIII, pp. 91–115. Springer (2016)
Korns, M.F.: An evolutionary algorithm for big data multiclass classification problems. In: Genetic Programming Theory and Practice XIV. Springer (2017)
Korns, M.F.: Evolutionary linear discriminant analysis for multiclass classification problems. In: Proceedings of the Genetic and Evolutionary Computation Conference Companion, pp. 233–234. ACM (2017)
Korns, M.F.: Genetic programming symbolic classification: A study. In: Genetic Programming Theory and Practice XV, pp. 39–52. Springer (2017)
McLachlan, G.: Discriminant analysis and statistical pattern recognition, vol. 544. John Wiley & Sons (2004)
Munoz, L., Silva, S., Trujillo, L.: M3gp–multiclass classification with gp. In: European Conference on Genetic Programming 2015, pp. 78–91. Springer (2015)
Platt, J.: Sequential minimal optimization: A fast algorithm for training support vector machines. Tech. Rep. MSR-TR-98-14, Microsoft Research (1998)
Acknowledgements
Our thanks to: Thomas May from Lantern Credit for assisting with the KNIME Learner training/scoring on all ten artificial classification problems.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Appendix: Artificial Test Problems
Appendix: Artificial Test Problems
-
T 1: y = argmax(D 1, D 2) where Y = 1, 2, X is 5000 × 25, and each D i is as follows:

-
T 2: y = argmax(D 1, D 2) where Y = 1, 2, X is 5000 × 100, and each D i is as follows:

-
T 3: y = argmax(D 1, D 2) where Y = 1, 2, X is 5000 × 1000, and each D i is as follows:
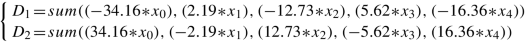
-
T 4: y = argmax(D 1, D 2, D 3) where Y = 1, 2, 3, X is 5000 × 25, and each D i is as follows:
$$\displaystyle \begin{aligned} \left\{ \begin{array}{ll} &D_1=sum((1.57*cos(x_0)),(-39.34*square(x_{10})),(2.13*(x_2/x_3)), \\&(46.59*cube(x_{13})),(-11.54*log(x_4))) \\&D_2=sum((-0.56*cos(x_0)),(9.34*square(x_{10})),(5.28*(x_2/x_3)),\\& (-6.10*cube(x_{13})),(1.48*log(x_4))) \\&D_3=sum((1.37*cos(x_0)),(3.62*square(x_{10})),(4.04*(x_2/x_3)),\\&(1.95*cube(x_{13})),(9.54*log(x_4))) \end{array} \right. \end{aligned}$$ -
T 5: y = argmax(D 1, D 2, D 3) where Y = 1, 2, 3, X is 5000 × 100, and each D i is as follows:
$$\displaystyle \begin{aligned}\left\{ \begin{array}{ll} &D_1=sum((1.57*sin(x_0)),(-39.34*square(x_{10})),(2.13*(x_2/x_3)),\\&(46.59*cube(x_{13})), (-11.54*log(x_4))) \\&D_2=sum((-0.56*sin(x_0)),(9.34*square(x_{10})),(5.28*(x_2/x_3)),\\&(-6.10*cube(x_{13})), (1.48*log(x_4))) \\&D_3=sum((1.37*sin(x_0)),(3.62*square(x_{10})),(4.04*(x_2/x_3)),\\&(1.95*cube(x_{13})),(9.54*log(x_4))) \end{array} \right.\end{aligned}$$ -
T 6: y = argmax(D 1, D 2, D 3) where Y = 1, 2, 3, X is 5000 × 1000, and each D i is as follows:
$$\displaystyle \begin{aligned}\left\{ \begin{array}{ll} &D_1=sum((1.57*tanh(x_0)),(-39.34*square(x_{10})),(2.13*(x_2/x_3)),\\&(46.59*cube(x_{13})), (-11.54*log(x_4))) \\&D_2=sum((-0.56*tanh(x_0)),(9.34*square(x_{10})),(5.28*(x_2/x_3)),\\&(-6.10*cube(x_{13})), (1.48*log(x_4))) \\&D_3=sum((1.37*tanh(x_0)),(3.62*square(x_{10})),(4.04*(x_2/x_3)),\\&(1.95*cube(x_{13})),(9.54*log(x_4))) \end{array} \right.\end{aligned}$$ -
T 7: y = argmax(D 1, D 2, D 3, D 4, D 5) where Y = 1, 2, 3, 4, 5, X is 5000 × 25, and each D i is as follows:
$$\displaystyle \begin{aligned}\left\{ \begin{array}{ll} &D_1=sum((1.57*cos(x_0/x_{21})),(9.34*((square(x_{10})/x_{14})*x_6)),\\&(2.13*((x_2/x_3)*log(x_8))), (46.59*(cube(x_{13})*(x_9/x_2))),\\&(-11.54*log(x_4*x_{10}*x_{15}))) \\&D_2=sum((-1.56*cos(x_0/x_{21})),(7.34*((square(x_{10})/x_{14})*x_6)),\\&(5.28*((x_2/x_3)*log(x_8))), (-6.10*(cube(x_{13})*(x_9/x_2))),\\&(1.48*log(x_4*x_{10}*x_{15}))) \\&D_3=sum((2.31*cos(x_0/x_{21})),(12.34*((square(x_{10})/x_{14})*x_6)),\\&(-1.28*((x_2/x_3)*log(x_8))), (0.21*(cube(x_{13})*(x_9/x_2))),\\&(2.61*log(x_4*x_{10}*x_{15}))) \\&D_4=sum((-0.56*cos(x_0/x_{21})),(8.34*((square(x_{10})/x_{14})*x_6)),\\&(16.71*((x_2/x_3)*log(x_8))), (-2.93*(cube(x_{13})*(x_9/x_2))),\\&(5.228*log(x_4*x_{10}*x_{15}))) \\&D_5=sum((1.07*cos(x_0/x_{21})),(-1.62*((square(x_{10})/x_{14})*x_6)),\\&(-0.04*((x_2/x_3)*log(x_8))),(-0.95*(cube(x_{13})*(x_9/x_2))),\\&(0.54*log(x_4*x_{10}*x_{15}))) \end{array} \right.\end{aligned}$$ -
T 8: y = argmax(D 1, D 2, D 3, D 4, D 5) where Y = 1, 2, 3, 4, 5, X is 5000 × 100, and each D i is as follows:
$$\displaystyle \begin{aligned}\left\{ \begin{array}{ll} &D_1=sum((1.57*sin(x_0/x_{11})),(9.34*((square(x_{12})/x_{4})*x_{46})),\\&(2.13*((x_{21}/x_3)*log(x_{18}))), (46.59*(cube(x_3)*(x_9/x_2))),\\&(-11.54*log(x_{14}*x_{10}*x_{15}))) \\&D_2=sum((-1.56*sin(x_0/x_{11})),(7.34*((square(x_{12})/x_{4})*x_{46})),\\&(5.28*((x_{21}/x_3)*log(x_{18}))), (-6.10*(cube(x_3)*(x_9/x_2))),\\&(1.48*log(x_{14}*x_{10}*x_{15}))) \\&D_3=sum((2.31*sin(x_0/x_{11})),(12.34*((square(x_{12})/x_{4})*x_{46})),\\&(-1.28*((x_{21}/x_3)*log(x_{18}))), (0.21*(cube(x_3)*(x_9/x_2))),\\&(2.61*log(x_{14}*x_{10}*x_{15}))) \\&D_4=sum((-0.56*sin(x_0/x_{11})),(8.34*((square(x_{12})/x_{4})*x_{46})),\\&(16.71*((x_{21}/x_3)*log(x_{18}))), (-2.93*(cube(x_3)*(x_9/x_2))),\\&(5.228*log(x_{14}*x_{10}*x_{15}))) \\&D_5=sum((1.07*sin(x_0/x_{11})),(-1.62*((square(x_{12})/x_{4})*x_{46})),\\&(-0.04*((x_{21}/x_3)*log(x_{18}))),(-0.95*(cube(x_3)*(x_9/x_2))),\\&(0.54*log(x_{14}*x_{10}*x_{15}))) \end{array} \right.\end{aligned}$$ -
T 9: y = argmax(D 1, D 2, D 3, D 4, D 5) where Y = 1, 2, 3, 4, 5, X is 5000 × 1000, and each D i is as follows:

-
T 10: y = argmax(D 1, D 2, D 3, D 4, D 5) where Y = 1, 2, 3, 4, 5, X is 5000 × 1000, and each D i is as follows:
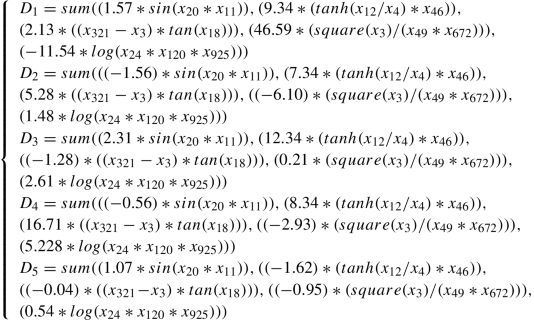
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Korns, M.F., May, T. (2019). Strong Typing, Swarm Enhancement, and Deep Learning Feature Selection in the Pursuit of Symbolic Regression-Classification. In: Banzhaf, W., Spector, L., Sheneman, L. (eds) Genetic Programming Theory and Practice XVI. Genetic and Evolutionary Computation. Springer, Cham. https://doi.org/10.1007/978-3-030-04735-1_4
Download citation
DOI: https://doi.org/10.1007/978-3-030-04735-1_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-04734-4
Online ISBN: 978-3-030-04735-1
eBook Packages: Computer ScienceComputer Science (R0)