Abstract
Accurate annotation of anatomical structures or pathological changes in microscopic images is an important task in computational pathology. Crowdsourcing holds promise to address this demand, but so far feasibility has only be shown for simple tasks and not for high-quality annotation of complex structures which is often limited by shortage of experts. Third-year medical students participated in solving two complex tasks, labeling of images and delineation of relevant image objects in breast cancer and kidney tissue. We evaluated their performance and addressed the requirements of task complexity and training phases. Our results show feasibility and a high agreement between students and experts. The training phase improved accuracy of image labeling.
This work was performed in the framework of SYSIMIT (FKZ:01ZX1308A), ILUMINATE (FKZ:031 B0006C), and SYSMIFTA (FKZ:031L0085A) funded by BMBF.
Nadine S. Schaadt and Anne Grote contributed equally to this work.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
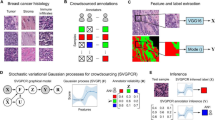
Crowdsourcing (CS) in digital pathology has been largely limited to less complex tasks such as identification of cancer cells [6, 11], scoring of cell nuclei based on immunohistochemistry (IHC) [2, 4, 6, 11], malaria diagnostics [9], and creation of training sets for convolutional neural networks [1, 5]. In general, intrinsically motivated contributors in voluntary CS perform better compared to paid “crowdworkers” [10]. Quality of CS depends on training and adaptation of task design to contributors’ background knowledge [3, 7].
In this paper, we investigate annotation of complex structures by medical students without pathology expertise but with profound understanding of anatomy and disease mechanism and a need to learn pathology as a strong incentive to recapitulate anatomy. We show that medical students can acquire skills to label images and delineate image objects in kidney and breast pathology, and discuss the influence of task complexity and training on CS approaches to produce high-quality annotations for machine learning.
2 Materials and Methods
2.1 Setting
We studied performance of a crowd of “educated” contributors: 142 third-year medical students, who were entering the curricular pathology course and thus had basic knowledge about microscopic anatomy but no expertise in pathology nor experience in annotating histological images.
We considered four independent experiments, each with 1–3 sessions on different days (Table 1). Each experiment started in a room equipped with computers with a short teaching session on relevant anatomical structures and pathological conditions, and explanations of the tools. The latter evolved from face-to-face lessons into a video tutorial, ready for use in experiment 4. The crowd were asked to work on two different tasks:
-
1.
Labeling of regions of interest (ROIs)
select one of several proposed categories for each of a set of images
Used tools: software developed for the project that displays the current image, a progress line, and radio buttons for each class
-
2.
Delineation of ROIs
draw the outlines of all objects of some well-defined classes and mark the class names in an image showing a tissue region
Used tools: Aperio ImageScope by Leica Microsystems (experiment 1), Cytomine [8] running on an own server (experiment 2–4)
The labeling task included an obligatory training phase in the beginning in which the correct solution was immediately shown to the participants and a test phase without feedback. Training for ROI delineation was introduced in experiment 4 as optional work on images with the possibility to switch on/off GT. Students received detailed feedback on both tasks after each session.
Images source were whole slide images (WSIs) from sections stained for H&E or IHC markers (ethical approval review board of Hannover Medical School).
2.2 Answer Aggregation and Evaluation
As final annotations, we aggregated individual statements as majority vote (MV: relative majority) or weighted vote (WV: weights calculated by training phase results of individuals). Equal votes result in unclassified objects and were counted as false negatives.
Two experts (one for each tissue type) provided annotations, such that there is a ground truth (GT) for each image to measure the performance of the crowd. To evaluate ROI labeling, we measured the accuracy averaged over each class,
where C is the set of classes, TP, TN, FP, and FN are the numbers of true positives, true negatives, false positives, and false negatives, respectively. This is compared to the expected value of random labeling, estimated as 1 / |C|.
In ROI delineation, due to comparably large tissue areas without occurrence of any considered class, we calculated the \(F_1\) score averaged over each class:
with the precision (PPV) and the recall (TPR)
both computed per class and averaged over all classes. This was calculated for each pixel that is part of the tissue.
3 Results and Discussion
3.1 Feasibility and Role of Task Complexity
We compared the crowd results with expert annotations for high-level structures in breast and kidney tissue in independent experiments confirming feasibility even for complex ROIs representing pathologically relevant tissue conditions.
ROI Labeling. Our results suggested that the class complexity has a stronger effect on crowd performance than the tissue type as mistakes occurred predominantly in the distinction of classes defined by complex object features (Table 2).
In experiment 1, automatically detected ROIs intended to show epithelial structures in normal breast tissue was categorized into: “lobule”, “duct”, “FP” (session 1), and additionally “lobule with extralobular ducts” (session 2 and 3).
In experiment 2, the crowd classified images from breast cancer cases, distinguishing between (1) “technical artefact”, (2) “invasive cancer”, (3) “intraepithelial neoplasia”, (4) “glandular epithelium”, and (5) “other anatomical structure”. As a single image could include normal and neoplastic structures, a more complex class definition was required. We used a hierarchical order such that the images should be classified by occurrence of highest order class. For example, if an image contained mainly glandular epithelium and some invasive tumor, then the image should be classified as “invasive tumor”. Accuracy of WV (weighted by training phase precision) was 0.976. Even the lowest accuracy for individuals was detectably higher than the corresponding probability by chance.
In experiment 3, kidney structures from biopsies were labeled into four types (“normal”, “pathologically changed”, “sclerotic”, “no” glomerulum). In both sessions, WV (session 1: 0.940, session 2: 0.832) was clearly higher than average but, some individuals outperformed the best combinations (Supp. Mat., Fig. 1). A particular challenge for the crowd was the class of “pathologically changed glomerula”, most likely because the class definition included semi-quantitative criteria such as hypercellularity, thickened Bowman’s capsule, mesangial sclerosis, collapse or retraction of the capillary tuft.
Experiment 4 considered four categories (“normal”, “partially sclerotic”, “sclerotic”, “no” glomerulum). Highest accuracy was achieved by WV (session 1: 0.973, session 2: 0.942). In the case of “partially sclerotic glomerula”, the precision was quite low for MV. Combining both classes “partially sclerotic glomerulum” and “sclerotic glomerulum” clearly increased the accuracy and precision.
ROI Delineation. To present the results, images are referred to as I, with experiment, session, and an id. For example, \(I_{ex3,se2,1}\) denotes the first image of the second session of the third experiment. If necessary, participant groups are referred to in the image name as G with group number. Figure 1 shows the difference of the MV to the reference for one image from each experiment. Table 3 shows the overall \(F_1\) scores for all experiments (further measures in Supp. Mat. Table 2–5) indicating general feasibility. The quality decreases with increasing complexity, number of classes, and image size. For structures with well-defined borders, such as glomerula (kidney) or lobules (breast), the borders of the objects have been drawn quite accurately in contrast to fractal-like outlines of tumor.
Difference of majority vote to ground truth (GT) in examples of kidney (A: \(I_{ex_1,se_1,1}\), C: \(I_{ex_3,se_1,G_{1},2}\)) and breast (B: \(I_{ex_2,se_1,2}\), D: \(I_{ex_4,se_1,1}\)) tissue. Green: agreement with GT, red: difference to GT. B: illustrates problems at tumor border. D: illustrates confused structures. (Color figure online)
Experiment 1 tested the crowd’s delineation performance on two image subsets representing renal tissue, using a duplex staining for immune cells (session 1) or immune cells and vascular endothelium (session 2). For delineating “glomerulum”, “artery”, and “tubulus”, the overall scores were distinctly lower for the second session than for the first session. Precision for the “artery” class in session 2 was markedly lower due to mislabeling of other blood vessels such as veins and smaller arterioles. To check how participants would be influenced by the provided classes, we used classes in session 2 that could potentially occur in kidney tissue but were not included in the specifically provided image. Several participants mistook narrow peritubular interstitial tissue for such a class (“collageneous tissue/septae”). We assume that in small, single images there is a tendency to annotate more objects in contrast to batches of larger images.
Experiment 2 tested a more complex setting for breast cancer and surrounding tissue. Classes e.g. included “invasive tumor”, “duct”, “lobule”, and “large blood vessel”. In most cases, the \(F_{1}\) scores of MV were better than the \(F_{1}\) scores on average. For the class “large blood vessel” in \(I_{ex_2,se_1,2}\), for example, the recall value was on average 0.472 and for the MV 0.826, without loss of precision. Some objects in this complex setting, however, were challenging. For example, blood vessels in \(I_{ex_2,se_2,1}\) and \(I_{ex_2,se_2,2}\) were missed by two thirds of the crowd. Common differences between MV and GT occurred in (1) individual variations in the object border delineation, most pronounced at the tumor border and (2) confusions between the visually similar structures (epithelial/epitheloid) “lobule”, “duct”, and “invasive tumor”.
Experiment 3 used eight WSIs of kidney tissue and focused on “glomerulum”, “artery”, and occasionally included “muscle”. We split the crowd into roughly equally sized groups \(G_{1}\) and \(G_{2}\). In each session, both groups worked on a common image (\(I_{ex_{3},se_{1},1}\) or \(I_{ex_{3},se_{2},1}\), stained for H&E) and additionally annotated one further image(s) stained for a macrophage marker. The class “glomerulum” had the highest scores. In five images, its MV precision was higher than 0.990, with virtually no FPs, and the outlines of the glomerula were close to GT.
In experiment 4, four images of breast cancer were used, with similar complexity to experiment 2, but with more participants. Classes were “duct”, “intra-epithelial neoplasia”, “tumor”, “lobule”, and “necrosis”. The MV results were in the same range as for experiment 2. The results of experiments 2 and 4 suggested that most objects could be found reliably already with a small crowd while some difficult objects could not be identified by most participants.
Overall, there seemed to be a role for certain pathological changes mimicking or hiding ROIs: In the renal images (experiment 1 and 3), sclerotic glomerula and arteries were sometimes confused and arteries were also frequently completely missed. In two images (experiment 2 and 4), lobules with heavy immune infiltration were missed by all participants.
Training phase effects in ROI labeling (A–C) and ROI delineation (D) A: Correlation between training and test accuracy for individuals (blue) and aggregations in experiment 3, session 1 (left) and session 2 (right). B: Changes of individual (lines) performance during experiment 4. C: Role of training phase length for difficult class “partially sclerotic glomerulum”. Shown is the difference between accuracy of the first 20 images and of images 21–40, 41–60, and 61–153. D: Influence of size of optional training image (blue: 80% of test image size, red: 40% of test image size) on \(F_1\) score. (Color figure online)
3.2 Role of Training Phase
ROI Labeling. For experiment 3, we compared the accuracy during the training phase, in which the correct label was shown to the participants immediately after their decision, with the test accuracy (Fig. 2A). Most students performed better during the test phase in both sessions, especially high-performer (based on test accuracy). Nevertheless, several results of the training phase were close to the test phase in session 1 (Spearman’s correlation coefficient: 0.41). To investigate a suitable size of the training phase, we varied them in experiment 4 (three student groups in each session: 20, 40, or 60 images). For this, we kept the same images in the same order. The number of correctly labeled images was similar with a trend to increase with increasing size of trainings phase (Fig. 2 (B)). Students that participated in both session 1 and 2 showed higher correctness in the second training phase compared to students first time participating. Figure 2 (C) shows that the second training phase did also not increase their accuracy for the most difficult class of partially sclerotic glomerula. We conclude out of this, that the training phase covering a broad variability of representatives for each class was helpful to increase the performance of the crowd.
ROI Delineation. The participants could annotate a “training image” with the option to see the GT in experiment 4. To measure the training effect, we compared two group of individuals that either received a small training image (40% of test image size, not all classes represented) or a large training image (80% of test size). No clear effect of the size on \(F_1\) score could be seen (Fig. 2D).
4 Conclusion
Our study shows general feasibility of CS for the annotation of complex histological images by participants with medical background, but without specific expert knowledge. To ensure annotation quality, it is necessary to design the tasks with well-defined objects and to include a sufficient training phase. Our approach can be adapted to individual project requirements and shows the importance of finding an adequate match between level of task complexity and previous knowledge of the crowd. Future work should focus on the comparison of “educated” contributors and nonexperts, and the usefulness of this type of noisy training data for machine learning.
References
Albarqouni, S., Baur, C., Achilles, F., Belagiannis, V., Demirci, S., Navab, N.: Aggnet: deep learning from crowds for mitosis detection in breast cancer histology images. IEEE Trans. Med. Imag. 35, 1313–1321 (2016)
Della Mea, V., Maddalena, E., Mizzaro, S., Machin, P., Beltrami, C.A.: Preliminary results from a crowdsourcing experiment in immunohistochemistry. Diagn. Pathol. 9, S6 (2014)
Hoßfeld, T., et al.: Best practices and recommendations for crowdsourced qoe-lessons learned from the qualinet task force crowdsourcing. In: QUALINET (2014)
Irshad, H., et al.: Crowdsourcing scoring of immunohistochemistry images: evaluating performance of the crowd and an automated computational method. Sci. Rep. 7 (2017)
Kim, E., Mente, S., Keenan, A., Gehlot, V.: Digital pathology annotation data for improved deep neural network classification. In: SPIE Medical Imaging, p. 101380D (2017)
Lawson, J., et al.: Crowdsourcing for translational research: analysis of biomarker expression using cancer microarrays. Br. J. Cancer 116, 237–245 (2017)
Liu, S., Xia, F., Zhang, J., Wang, L., Wang, L.: How crowdsourcing risks affect performance: an exploratory model. Manag. Decis. 54, 2235–2255 (2016)
Marée, R.: Collaborative analysis of multi-gigapixel imaging data using cytomine. Bioinformatics 32, 1395–1401 (2016)
Mavandadi, S., et al.: Distributed medical image analysis and diagnosis through crowd-sourced games: a malaria case study. PloS One 7, e37245 (2012)
Redi, J., Povoa, I.: Crowdsourcing for rating image aesthetic appeal: better a paid or a volunteer crowd? In: Proceedings of 2014 International ACM Workshop Crowdsourcing Multimedia, pp. 25–30. ACM (2014)
dos Reis, F.J.C., et al.: Crowdsourcing the general public for large scale molecular pathology studies in cancer. EBioMedicine 2, 681–689 (2015)
Acknowledgements
We thank all students for contribution; M. Temerinac-Ott, Icube; R. Schönmeyer, C. Vanegas, Definiens for help in data selection; G. Stiller, M. Behrends, Peter L. Reichertz Institute for Medical Informatics; and A.-K. Rieke for the video.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Switzerland AG
About this paper
Cite this paper
Schaadt, N.S., Grote, A., Forestier, G., Wemmert, C., Feuerhake, F. (2018). Role of Task Complexity and Training in Crowdsourced Image Annotation. In: Stoyanov, D., et al. Computational Pathology and Ophthalmic Medical Image Analysis. OMIA COMPAY 2018 2018. Lecture Notes in Computer Science(), vol 11039. Springer, Cham. https://doi.org/10.1007/978-3-030-00949-6_6
Download citation
DOI: https://doi.org/10.1007/978-3-030-00949-6_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-00948-9
Online ISBN: 978-3-030-00949-6
eBook Packages: Computer ScienceComputer Science (R0)