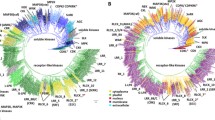
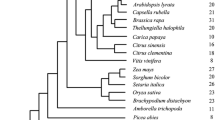
The Arabidopsis Genome Contains 61 Kinesins, The Largest Kinesin Family Of Any Fully Sequenced And Annotated Genome. Characterizing Each Member Of A Large Protein Family Is A Difficult And Laborious Task, Not Only Due To The Size, But Also The Inherent Diversification And/Or Redundancy Associated With Gene Duplication. An Alternative Approach To Better Understand The Arabidopsis Kinesin Family Is To Study The Domain Composition Of These Proteins. Domains Are Defined As The Structural And Functional Units Of Proteins. Understanding The Domains And Domain Structure Of An Unknown Protein Often Allows For A Reliable Prediction Of Function When Experimental Evidence Is Unavailable. Kinesins Share A Highly Conserved Motor Domain With Low Sequence Similarity Outside Of This Domain, And Many Members Contain Additional Protein Domains In The Nonconserved Regions. New Arrangements Of Domains, By Means Of Genetic Duplication, Replication, Or Loss, Result In Novel Proteins With Unique Functions. Using Both The Arabidopsis And Other Sequenced Genomes, It Was Found That Several Kinesins Have Protein Domain Arrangements Specific To Different Lineages, Providing Compelling Evidence For The Evolution Of Kinesins With Novel Functions. A Phylogenic Analysis Of Plant-Specific Kinesin Domains, As Well As A Comparative Review Of The Molecular/ Biochemical Functions Of These Domains, Is Presented In This Chapter. Combined, They Provide An Alternative Means To Understanding And Developing More Direct And Relevant Hypotheses Concerning Novel Kinesins.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
C. J. Lawrence, R. K. Dawe, K. R. Christie, D. W. Cleveland, S. C. Dawson, S. A. Endow, L. S. B. Goldstein, H. V. Goodson, N. Hirokawa, J. Howard, R. L. Malmberg, J. R. McIntosh, H. Miki, T. J. Mitchison, Y. Okada, A. S. N. Reddy, W. M. Saxton, M. Schliwa, J. M. Scholey, R. D. Vale, C. E. Walczak, and L. Wordeman, A standardized kinesin nomenclature, J. Cell Bio. 167(1), 19–22 (2004).
H. Miki, Y. Okada, and N. Hirokawa, Analysis of the kinesin superfamily: insights into structure and function, Trends Cell Biol. 15(9), 467–76 (2005).
D. N. Richardson, M. P. Simmons, and A. S. Reddy, Comprehensive comparative analysis of kinesins in photosynthetic eukaryotes, BMC Genomics 7, 18 (2006).
A. S. N. Reddy and I. Day, Kinesins in the Arabidopsis genome: A comparative analysis among eukaryotes, BMC Genomics 2(1), 2 (2001).
A. Bannigan, M. Lizotte-Waniewski, M. Riley, and T. I. Baskin, Emerging molecular mechanisms that power and regulate the anastral mitotic spindle of flowering plants, Cell Motil. Cytoskel. 65(1), 1–11 (2007).
H. Tanaka, M. Ishikawa, S. Kitamura, Y. Takahashi, T. Soyano, C. Machida, and Y. Machida, The AtNACK1/HINKEL and STUD/TETRASPORE/AtNACK2 genes, which encode functionally redundant kinesins, are essential for cytokinesis in Arabidopsis, Genes Cells 9(12), 1199–211 (2004).
S. Muller, S. Han, and L. G. Smith, Two kinesins are involved in the spatial control of cytokinesis in Arabidopsis thaliana, Curr. Biol. 16(9), 888–94 (2006).
Y. R. Lee, Y. Li, and B. Liu, Two Arabidopsis phragmoplast-associated kinesins play a critical role in cytokinesis during male gametogenesis, Plant Cell 19(8), 2595–605 (2007).
C. B. Chen, A. Marcus, W. X. Li, Y. Hu, J. P. V. Calzada, U. Grossniklaus, R. J. Cyr, and H. Ma, The Arabidopsis ATK1 gene is required for spindle morphogenesis in male meiosis, Development 129(10), 2401–2409 (2002).
J. C. Ambrose, W. Li, A. Marcus, H. Ma, and R. Cyr, A minus-end-directed kinesin with plus-end tracking protein activity is involved in spindle morphogenesis, Mol. Biol. Cell 16(4), 1584–1592 (2005).
D. G. Oppenheimer, M. A. Pollock, J. Vacik, D. B. Szymanski, B. Ericson, K. Feldmann, and M. D. Marks, Essential role of a kinesin-like protein in Arabidopsis trichome morphogenesis, PNAS 94, 6261–6266 (1997).
M. A. Jones, M. J. Raymond, and N. Smirnoff, Analysis of the root-hair morphogenesis transcriptome reveals the molecular identity of six genes with roles in root-hair development in Arabidopsis, Plant J. 45(1), 83–100 (2006).
D. B. Wetlaufer, Nucleation, rapid folding, and globular intrachain regions in proteins, PNAS 70(3), 697–701 (1973).
R. F. Doolittle, The multiplicity of domains in proteins, Ann. Rev. Biochem. 64, 287–314 (1995).
C. P. Ponting and R. R. Russell, The natural history of protein domains, Ann. Rev. Biophys. Biomol. Struct. 31, 45–71 (2002).
F. Jacob, Molecular tinkering in evolution, in Evolution from Molecules to Men, D.S. Bendell, Editor. 1983, Cambridge University Press: Cambridge, pp. 131–144.
R. F. Doolittle, Similar amino acid sequences: chance or common ancestry? Science 214(4517), 149–159 (1981).
A. N. Lupas, C. P. Ponting, and R. B. Russell, On the evolution of protein folds: are similar motifs in different protein folds the result of convergence, insertion, or relics of an ancient peptide world? J. Struct. Biol. 134(2–3), 191–203 (2001).
E. S. Lander, L. M. Linton, B. Birren, C. Nusbaum, M. C. Zody, J. Baldwin, K. Devon, K. Dewar, M. Doyle, W. FitzHugh, R. Funke, D. Gage, K. Harris, A. Heaford, J. Howland, L. Kann, J. Lehoczky, R. LeVine, P. McEwan, K. McKernan, J. Meldrim, J. P. Mesirov, C. Miranda, W. Morris, J. Naylor, C. Raymond, M. Rosetti, R. Santos, A. Sheridan, C. Sougnez, N. Stange-Thomann, N. Stojanovic, A. Subramanian, D. Wyman, J. Rogers, J. Sulston, R. Ainscough, S. Beck, D. Bentley, J. Burton, C. Clee, N. Carter, A. Coulson, R. Deadman, P. Deloukas, A. Dunham, I. Dunham, R. Durbin, L. French, D. Grafham, S. Gregory, T. Hubbard, S. Humphray, A. Hunt, M. Jones, C. Lloyd, A. McMurray, L. Matthews, S. Mercer, S. Milne, J. C. Mullikin, A. Mungall, R. Plumb, M. Ross, R. Shownkeen, S. Sims, R. H. Waterston, R. K. Wilson, L. W. Hillier, J. D.McPherson, M. A. Marra, E. R. Mardis, L. A. Fulton, A. T. Chinwalla, K. H. Pepin, W. R. Gish, S. L. Chissoe, M. C. Wendl, K. D. Delehaunty, T. L. Miner, A. Delehaunty, J. B. Kramer, L. L. Cook, R. S. Fulton, D. L. Johnson, P. J. Minx, S. W. Clifton, T. Hawkins, E. Branscomb, P. Predki, P. Richardson, S. Wenning, T. Slezak, N. Doggett, J. F. Cheng, A. Olsen, S. Lucas, C. Elkin, E. Uberbacher, M. Frazier, R. A. Gibbs, D. M. Muzny, S. E. Scherer, J. B. Bouck, E. J. Sodergren, K. C. Worley, C. M. Rives, J. H. Gorrell, M. L. Metzker, S. L. Naylor, R. S. Kucherlapati, D. L. Nelson, G. M. Weinstock, Y. Sakaki, A. Fujiyama, M. Hattori, T. Yada, A. Toyoda, T. Itoh, C. Kawagoe, H. Watanabe, Y. Totoki, T. Taylor, J. Weissenbach, R. Heilig, W. Saurin, F. Artiguenave, P. Brottier, T. Bruls, E. Pelletier, C. Robert, P. Wincker, D. R. Smith, L. Doucette-Stamm, M. Rubenfield, K. Weinstock, H. M. Lee, J. Dubois, A. Rosenthal, M. Platzer, G. Nyakatura, S. Taudien, A. Rump, H. Yang, J. Yu, J. Wang, G. Huang, J. Gu, L. Hood, L. Rowen, A. Madan, S. Qin, R. W. Davis, N. A. Federspiel, A. P. Abola, M. J. Proctor, R. M. Myers, J. Schmutz, M. Dickson, J. Grimwood, D. R. Cox, M. V. Olson, R. Kaul, N. Shimizu, K. Kawasaki, S. Minoshima, G. A. Evans, M. Athanasiou, R. Schultz, B. A. Roe, F. Chen, H. Pan, J. Ramser, H. Lehrach, R. Reinhardt, W. R. McCombie, M. de la Bastide, N. Dedhia, H. Blocker, K. Hornischer, G. Nordsiek, R. Agarwala, L. Aravind, J. A. Bailey, A. Bateman, S. Batzoglou, E. Birney, P. Bork, D. G. Brown, C. B. Burge, L. Cerutti, H. C. Chen, D. Church, M. Clamp, R. R. Copley, T. Doerks, S. R. Eddy, E. E. Eichler, T. S. Furey, J. Galagan, J. G. Gilbert, C. Harmon, Y. Hayashizaki, D. Haussler, H. Hermjakob, K. Hokamp, W. Jang, L. S. Johnson, T. A. Jones, S. Kasif, A. Kaspryzk, S. Kennedy, W. J. Kent, P. Kitts, E. V. Koonin, I. Korf, D. Kulp, D. Lancet, T. M. Lowe, A. McLysaght, T. Mikkelsen, J. V. Moran, N. Mulder, V. J. Pollara, C. P. Ponting, G. Schuler, J. Schultz, G. Slater, A. F. Smit, E. Stupka, J. Szustakowski, D. Thierry-Mieg, J. Thierry-Mieg, L. Wagner, J. Wallis, R. Wheeler, A. Williams, Y. I. Wolf, K. H. Wolfe, S. P. Yang, R. F. Yeh, F. Collins, M. S. Guyer, J.Peterson, A. Felsenfeld, K. A. Wetterstrand, A. Patrinos, M. J. Morgan, P. de Jong, J. J. Catanese, K. Osoegawa, H. Shizuya, S. Choi and Y. J. Chen, Initial sequencing and analysis of the human genome, Nature 409(6822), 860–921 (2001).
S. Liu, C. Zhang, and Y. Zhou, Domain graph of Arabidopsis proteome by comparative analysis, J. Proteome Res. 4(2), 435–444 (2005).
F. J. Kull, Motor proteins of the kinesin superfamily: structure and mechanism, Essays Biochem. 35, 61–73 (2000).
C. J. Lawrence, R. L. Malmberg, M. G. Muszynski, and R. K. Dawe, Maximum likelihood methods reveal conservation of function among closely related kinesin families, J. Mol. Evol. 54(1), 42–53 (2002).
R. D. Vale and R. J. Fletterick, The design plan of kinesin motors, Ann. Rev. Cell Dev. Biol. 13, 745–77 (1997).
E. P. Sablin, R. B. Case, S. C. Dai, C. L. Hart, A. Ruby, R. D. Vale, and R. J. Fletterick, Direction determination in the minus-end-directed kinesin motor ncd, Nature 395(6704), 813–816 (1998).
E. P. Sablin, F. J. Kull, R. Cooke, R. D. Vale, and R. J. Fletterick, Crystal structure of the motor domain of the kinesin-related motor ncd, Nature 380(6574), 555–559 (1996).
J. Nakayama, G. Xiao, K. Noma, A. Malikzay, P. Bjerling, K. Ekwall, R. Kobayashi, and S. I. Grewal, Alp13, an MRG family protein, is a component of fission yeast Clr6 histone deacetylase required for genomic integrity, Embo J. 22(11), 2776–2787 (2003).
J. Schultz, F. Milpetz, P. Bork, and C. P. Ponting, SMART, a simple modular architecture research tool: identification of signaling domains, PNAS 95(11), 5857–5864 (1998).
S. E. Abdel-Ghany, I. S. Day, M. P. Simmons, P. Kugrens, and A. S. Reddy, Origin and evolution of Kinesin-like calmodulin-binding protein, Plant Physiol. 138(3), 1711–1722 (2005).
T. Sakai, H. V. Honing, M. Nishioka, Y. Uehara, M. Takahashi, N. Fujisawa, K. Saji, M. Seki, K. Shinozaki, M. A. Jones, N. Smirnoff, K. Okada, and G. O. Wasteneys, Armadillo repeat-containing kinesins and a NIMA-related kinase are required for epidermal-cell morphogenesis in Arabidopsis, Plant J. 53(1), 157–171 (2008).
G. Yang, P. Gao, H. Zhang, S. Huang, and Z. L. Zheng, A mutation in MRH2 kinesin enhances the root hair tip growth defect caused by constitutively activated ROP2 small GTPase in Arabidopsis, PLoS ONE. 2(10), e1074 (2007).
M. Peifer, S. Berg, and A. B. Reynolds, A repeating amino acid motif shared by proteins with diverse cellular roles, Cell 76(5), 789–791 (1994).
E. Wieschaus and R. Riggleman, Autonomous requirements for the segment polarity gene armadillo during Drosophila embryogenesis, Cell 49(2), 177–184 (1987).
M. Hatzfeld, The armadillo family of structural proteins, Int. Rev. Cytol. — a Survey of Cell Biology, 186, 179–224 (1999).
S. Kim, H. I. Choi, H. J. Ryu, J. H. Park, M. D. Kim, and S. Y. Kim, ARIA, an Arabidopsis arm repeat protein interacting with a transcriptional regulator of abscisic acid-responsive gene expression, is a novel abscisic acid signaling component, Plant Physiol. 136(3), 3639–3648 (2004).
L. M. Palevitz, Cloning and Characterization of PAK, a Novel Plant Kinesin-Like Protein Involved in Guard Cell Development and Expressed in Other Differentiating Cell Types in the Brassicacease. 1999, Cornell: Ithaca, NY.
K. Tamura, J. Dudley, M. Nei, and S. Kumar, MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0, Mol. Biol. Evol. 24(8), 1596–1599 (2007).
J. C. Coates, L. Laplaze, and J. Haseloff, Armadillo-related proteins promote lateral root development in Arabidopsis, PNAS 103(5), 1621–1626 (2006).
T. Jimbo, Y. Kawasaki, R. Koyama, R. Sato, S. Takada, K. Haraguchi, and T. Akiyama, Identification of a link between the tumour suppressor APC and the kinesin superfamily, Nat. Cell Biol. 4(4), 323–327 (2002).
P. C. Salinas, Modulation of the microtubule cytoskeleton: a role for a divergent canonical Wnt pathway, Trends Cell Biol. 17(7), 333–342 (2007).
P. Tompa, Y. Emori, H. Sorimachi, K. Suzuki, and P. Friedrich, Domain III of calpain is a ca2+-regulated phospholipid-binding domain, Biochem. Biophys. Res. Comm. 280(5), 1333–1339 (2001).
A. Rodriguez, E. Samoff, M. G. Rioult, A. Chung, and N. W. Andrews, Host cell invasion by trypanosomes requires lysosomes and microtubule/kinesin-mediated transport, J. Cell Biol. 134(2), 349–362 (1996).
H. Mitsui, K Yamaguchi-Shinozaki, K. Shinozaki, K. Nishikawa, and H. Takahashi, Identification of a gene family (kat) encoding kinesin-like proteins in Arabidopsis thaliana and the characterization of secondary structure of KatA, Mol. Gen. Genet. 362–368 (1993).
K. Vancompernolle, M. Gimona, M. Herzog, J. Van Damme, J. Vandekerckhove, and V. Small, Isolation and sequence of a tropomyosin-binding fragment of turkey gizzard calponin, FEBS Lett. 274(1–2), 146–150 (1990).
M. Gimona, K. Djinovic-Carugo, W. J. Kranewitter, and S. J. Winder, Functional plasticity of CH domains, FEBS Lett. 513(1), 98–106 (2002).
P. Matsudaira, Modular organization of actin crosslinking proteins, Trends Biochem. Sci. 16(3), 87–92 (1991).
M. Gimona and S. J. Winder, Single calponin homology domains are not actin-binding domains, Curr. Biol. 8(19), R674–R675 (1998).
A. M. Bashour, A. T. Fullerton, M. J. Hart, and G. S. Bloom, IQGAP1, a Rac- and Cdc42-binding protein, directly binds and cross-links microfilaments, J. Cell Biol. 137(7), 1555–1566 (1997).
M. L. Preuss, D. R. Kovar, Y. R. Lee, C. J. Staiger, D. P. Delmer, and B. Liu, A plant-specific kinesin binds to actin microfilaments and interacts with cortical microtubules in cotton fibers, Plant Physiol. 136(4), 3945–3955 (2004).
A. J. Saurin, K. L. Borden, M. N. Boddy, and P. S. Freemont, Does this have a familiar RING?, Trends Biochem. Sci. 21(6), 208–214 (1996).
R. Gamsjaeger, C. K. Liew, F. E. Loughlin, M. Crossley, and J. P. Mackay, Sticky fingers: zinc-fingers as protein-recognition motifs, Trends Biochem. Sci. 32(2), 63–70 (2007).
Z. Hua and T. H. Kao, Identification and characterization of components of a putative petunia S-locus F-box-containing E3 ligase complex involved in S-RNase-based self-incompatibility, Plant Cell 18(10), 2531–2553 (2006).
M. A. Larkin, G. Blackshields, N. P. Brown, R. Chenna, P. A. McGettigan, H. McWilliam, F. Valentin, I. M. Wallace, A. Wilm, R. Lopez, J. D. Thompson, T. J. Gibson, and D. G. Higgins, Clustal W and Clustal X version 2.0, Bioinformatics 23(21), 2947–2948 (2007).
C. Maris, C. Dominguez, and F. H. Allain, The RNA recognition motif, a plastic RNA-binding platform to regulate post-transcriptional gene expression, FEBS J. 272(9), 2118–2131 (2005).
M. Vermel, B. Guermann, L. Delage, J. M. Grienenberger, L. Marechal-Drouard, and J. M. Gualberto, A family of RRM-type RNA-binding proteins specific to plant mitochondria, PNAS 99(9), 5866–5871 (2002).
G. Schuster and W. Gruissem, Chloroplast mRNA 3′ end processing requires a nuclear-encoded RNA-binding protein, Embo J. 10(6), 1493–1502 (1991).
M. Kloc, N. R. Zearfoss, and L. D. Etkin, Mechanisms of subcellular mRNA localization, Cell 108(4), 533–544 (2002).
Y. Kanai, N. Dohmae, and N. Hirokawa, Kinesin transports RNA: isolation and characterization of an RNA-transporting granule, Neuron 43(4), 513–525 (2004).
P. Enkhbayar, M. Kamiya, M. Osaki, T. Matsumoto, and N. Matsushima, Structural principles of leucine-rich repeat (LRR) proteins, Proteins 54(3), 394–403 (2004).
S. K. Hanks and T. Hunter, Protein kinases 6. The eukaryotic protein kinase superfamily: kinase (catalytic) domain structure and classification, Faseb J. 9(8), 576–596 (1995).
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2008 Springer Science + Business Media B.V.
About this paper
Cite this paper
Malcos, J.L., Cyr, R.J. (2008). Domain Complexity Of Plant Kinesins. In: Blume, Y.B., Baird, W.V., Yemets, A.I., Breviario, D. (eds) The Plant Cytoskeleton: a Key Tool for Agro-Biotechnology. NATO Science for Peace and Security Series C: Environmental Security. Springer, Dordrecht. https://doi.org/10.1007/978-1-4020-8843-8_17
Download citation
DOI: https://doi.org/10.1007/978-1-4020-8843-8_17
Publisher Name: Springer, Dordrecht
Print ISBN: 978-1-4020-8842-1
Online ISBN: 978-1-4020-8843-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)