Abstract
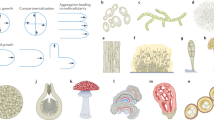
The conidia of airborne fungi are protected by a hydrophobic protein layer that coats the cell wall polysaccharides and renders the spores resistant to wetting and desiccation. A similar layer is presented on the outer surface of the aerial hyphae of some fungi. This layer serves multiple purposes, including facilitating spore dispersal, mediating the growth of hyphae into the air from moist environments, aiding host interactions in symbiotic relationships and increasing infectivity in pathogenic fungi. The layer consists of tightly packed, fibrillar structures termed “rodlets”, which are approximately 10 nm in diameter, hundreds of nanometres long and grouped in fascicles. Rodlets are an extremely stable protein structure, being resistant to detergents, denaturants and alcohols and requiring strong acids for depolymerisation. They are produced through the self-assembly of small, surface-active proteins that belong to the hydrophobin protein family. These small proteins are expressed by all filamentous fungi and are characterised by a high proportion of hydrophobic residues and the presence of eight cysteine residues. Rodlets are a form of the functional amyloid fibril, where the hydrophobin monomers are held together in the rodlets by intermolecular hydrogen bonds that contribute to a stable β-sheet core.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Aimanianda V, Bayry J, Bozza S, Kniemeyer O, Perruccio K, Elluru SR, Clavaud C, Paris S, Brakhage AA, Kaveri SV, Romani L, Latge JP (2009) Surface hydrophobin prevents immune recognition of airborne fungal spores. Nature 460(7259):1117–1121
Bayry J, Aimanianda V, Guijarro JI, Sunde M, Latge JP (2012) Hydrophobins–unique fungal proteins. PLoS Pathog 8(5):e1002700. https://doi.org/10.1371/journal.ppat.1002700
Beauvais A, Latge JP (2018) Special issue: fungal cell wall. J Fungi (Basel, Switzerland) 4(3). https://doi.org/10.3390/jof4030091
Beckerman JL, Ebbole DJ (1996) MPG1, a gene encoding a fungal hydrophobin of Magnaporthe grisea, is involved in surface recognition. Mol Plant-Microbe Interact: MPMI 9(6):450–456
Beever RE, Dempsey GP (1978) Function of rodlets on the surface of fungal spores. Nature 272(5654):608–610. https://doi.org/10.1038/272608a0
Beever RE, Redgewell RJ, Dempsey G (1979a) Purification and chemical characterization of the rodlet layer of Neurospora crassa conidia. J Bacteriol 140:1063–1070
Beever RE, Redgwell RJ, Dempsey GP (1979b) Purification and chemical characterization of the rodlet layer of Neurospora crassa conidia. J Bacteriol 140(3):1063
Bidochka MJ, Stleger RJ, Joshi L, Roberts DW (1995) The rodlet layer from aerial and submerged conidia of the entomopathogenic fungus beauveria-bassiana contains hydrophobin. Mycol Res 99:403–406
Butko P, Buford JP, Goodwin JS, Stroud PA, McCormick CL, Cannon GC (2001) Spectroscopic evidence for amyloid-like interfacial self-assembly of hydrophobin Sc3. Biochem Biophys Res Commun 280(1):212–215
Cho EM, Kirkland BH, Holder DJ, Keyhani NO (2007) Phage display cDNA cloning and expression analysis of hydrophobins from the entomopathogenic fungus Beauveria (Cordyceps) bassiana. Microbiology (Reading, England) 153(Pt 10):3438–3447. https://doi.org/10.1099/mic.0.2007/008532-0
Chuang E, Hori AM, Hesketh CD, Shorter J (2018) Amyloid assembly and disassembly. J Cell Sci 131(8):jcs189928. https://doi.org/10.1242/jcs.189928
Cole GT (1973) A correlation between rodlet orientation and conidiogenesis in Hyphomycetes. Can J Botany-Revue Can De Botanique 51:2413–2422
Cole GT, Sekiya T, Kasai R, Yokoyama T, Nozawa Y (1979) Surface ultrastructure and chemical composition of the cell walls of conidial fungi. Exp Mycol 3(2):132–156. https://doi.org/10.1016/S0147-5975(79)80025-0
Dagenais TR, Giles SS, Aimanianda V, Latge JP, Hull CM, Keller NP (2010) Aspergillus fumigatus LaeA-mediated phagocytosis is associated with a decreased hydrophobin layer. Infect Immun 78(2):823–829. https://doi.org/10.1128/iai.00980-09
Dague E, Alsteens D, Latgé J-P, Dufrêne YF (2008) High-resolution cell surface dynamics of germinating Aspergillus fumigatus conidia. Biophys J 94(2):656–660. https://doi.org/10.1529/biophysj.107.116491
De Simone A, Kitchen C, Kwan AH, Sunde M, Dobson CM, Frenkel D (2012) Intrinsic disorder modulates protein self-assembly and aggregation. Proc Natl Acad Sci USA 109(18):6951–6956. https://doi.org/10.1073/pnas.1118048109
de Vocht ML, Scholtmeijer K, van der Vegte EW, de Vries OM, Sonveaux N, Wosten HA, Ruysschaert JM, Hadziloannou G, Wessels JG, Robillard GT (1998) Structural characterization of the hydrophobin SC3, as a monomer and after self-assembly at hydrophobic/hydrophilic interfaces. Biophys J 74(4):2059–2068. https://doi.org/10.1016/s0006-3495(98)77912-3
de Vries OMH, Fekkes MP, Wösten HAB, Wessels JGH (1993) Insoluble hydrophobin complexes in the walls of Schizophyllum commune and other filamentous fungi. Arch Microbiol 159(4):330–335. https://doi.org/10.1007/BF00290915
Dempsey GP, Beever RE (1979) Electron microscopy of the rodlet layer of Neurospora crassa conidia. J Bacteriol 140(3):1050–1062
Dyer PS (2002) Hydrophobins in the lichen symbiosis. New Phytol 154(1):1–4. https://doi.org/10.1046/j.1469-8137.2002.00387.x
Fan H, Wang X, Zhu J, Robillard GT, Mark AE (2006) Molecular dynamics simulations of the hydrophobin SC3 at a hydrophobic/hydrophilic interface. Proteins: Struct Funct Bioinform 64(4):863–873. https://doi.org/10.1002/prot.20936
Fernandez J, Orth K (2018) Rise of a cereal killer: the biology of Magnaporthe oryzae biotrophic growth. Trends Microbiol 26(7):582–597. https://doi.org/10.1016/j.tim.2017.12.007
Gandier J-A, Langelaan DN, Won A, O’Donnell K, Grondin JL, Spencer HL, Wong P, Tillier E, Yip C, Smith SP, Master ER (2017) Characterization of a Basidiomycota hydrophobin reveals the structural basis for a high-similarity Class I subdivision. Sci Rep 7:45863–45863. https://doi.org/10.1038/srep45863
Gardner JS, Hess WM, Tripathi RK (1983) Surface rodlets of Tilletia indica teliospores. J Bacteriol 154(1):502–504
Gerin PA, Asther M, Sleytr UB, Rouxhet PG (1994) Detection of rodlets in the outer wall region of conidiospores of Phanerochaete chrysosporium. Can J Microbiol 40(5):412–416
Ghiorse WC, Edwards MR (1973) Ultrastructure of Aspergillus fumigatus conidia development and maturation. Protoplasma 76(1):49–59
Girardin H, Paris S, Rault J, Bellon-Fontaine MN, Latgé JP (1999) The role of the rodlet structure on the physicochemical properties of Aspergillus conidia. Lett Appl Microbiol 29(6):364–369. https://doi.org/10.1046/j.1472-765X.1999.00643.x
Grunbacher A, Throm T, Seidel C, Gutt B, Rohrig J, Strunk T, Vincze P, Walheim S, Schimmel T, Wenzel W, Fischer R (2014) Six hydrophobins are involved in hydrophobin rodlet formation in Aspergillus nidulans and contribute to hydrophobicity of the spore surface. PLoS ONE 9(4):e94546. https://doi.org/10.1371/journal.pone.0094546
Gunning AP, De Groot PWJ, Visser J, Morris VJ (1998) Atomic force microscopy of a hydrophobin protein from the edible mushroom Agaricus bisporus. J Colloid Interface Sci 201(2):118–126
Hakanpaa J, Linder M, Popov A, Schmidt A, Rouvinen J, Linder MB, Szilvay GR, Nakari-Setala T, Penttila ME (2006) Hydrophobin HFBII in detail: ultrahigh-resolution structure at 0.75 A. Acta Crystallogr D Biol Crystallogr 62 (Pt 4):356–367
Hakanpaa J, Paananen A, Askolin S, Nakari-Setala T, Parkkinen T, Penttila M, Linder MB, Rouvinen J (2004) Atomic resolution structure of the HFBII hydrophobin, a self-assembling amphiphile. J Biol Chem 279(1):534–539
Hallett IC, Beever RE (1981) Rodlets on the surface of Neurospora conidia. Trans Br Mycol Soc 77(3):662–665. https://doi.org/10.1016/S0007-1536(81)80124-6
Hess WM, Sassen MM, Remsen CC (1968) Surface characteristics of Penicillum conidia. Mycologia 60(2):290–303
Hess WM, Stocks DL (1969) Surface characteristics of Aspergillus conidia. Mycologia 61(3):560–571
Holder DJ, Keyhani NO (2005) Adhesion of the entomopathogenic fungus Beauveria (Cordyceps) bassiana to substrata. Appl Environ Microbiol 71(9):5260–5266. https://doi.org/10.1128/aem.71.9.5260-5266.2005
Honegger R (1991) Functional aspects of the lichen symbiosis. Annu Rev Plant Physiol Plant Mol Biol 42(1):553–578. https://doi.org/10.1146/annurev.pp.42.060191.003005
Jensen BG, Andersen MR, Pedersen MH, Frisvad JC, Sondergaard I (2010) Hydrophobins from Aspergillus species cannot be clearly divided into two classes. BMC Res Notes 3:344. https://doi.org/10.1186/1756-0500-3-344
Kershaw MJ, Wakley G, Talbot NJ (1998) Complementation of the Mpg1 mutant phenotype in Magnaporthe grisea reveals functional relationships between fungal hydrophobins. EMBO J 17(14):3838–3849. https://doi.org/10.1093/emboj/17.14.3838
Kollmer M, Close W, Funk L, Rasmussen J, Bsoul A, Schierhorn A, Schmidt M, Sigurdson CJ, Jucker M, Fändrich M (2019) Cryo-EM structure and polymorphism of Aβ amyloid fibrils purified from Alzheimer’s brain tissue. Nat Commun 10(1):4760. https://doi.org/10.1038/s41467-019-12683-8
Kwan AH, Macindoe I, Vukasin PV, Morris VK, Kass I, Gupte R, Mark AE, Templeton MD, Mackay JP, Sunde M (2008) The Cys3-Cys4 loop of the hydrophobin EAS is not required for rodlet formation and surface activity. J Mol Biol 382(3):708–720. https://doi.org/10.1016/j.jmb.2008.07.034
Kwan AH, Winefield RD, Sunde M, Matthews JM, Haverkamp RG, Templeton MD, Mackay JP (2006) Structural basis for rodlet assembly in fungal hydrophobins. Proc Natl Acad Sci USA 103(10):3621–3626. https://doi.org/10.1073/pnas.0505704103
Kwon-Chung KJ, Sugui JA (2013) Aspergillus fumigatus–what makes the species a ubiquitous human fungal pathogen? PLoS Pathog 9(12):e1003743–e1003743. https://doi.org/10.1371/journal.ppat.1003743
Lacroix H, Whiteford JR, Spanu PD (2008) Localization of Cladosporium fulvum hydrophobins reveals a role for HCf-6 in adhesion. FEMS Microbiol Lett 286(1):136–144. https://doi.org/10.1111/j.1574-6968.2008.01227.x
Latge JP (1999) Aspergillus fumigatus and aspergillosis. Clin Microbiol Rev 12(2):310–350
Latge JP, Bouziane H, Diaquin M (1988) Ultrastructure and composition of the conidial wall of Cladosporium cladosporioides. Can J Microbiol 34(12):1325–1329
Lee MJ, Sheppard DC (2016) Recent advances in the understanding of the Aspergillus fumigatus cell wall. J Microbiol (Seoul, Korea) 54(3):232–242. https://doi.org/10.1007/s12275-016-6045-4
Linder MB, Szilvay GR, Nakari-Setala T, Penttila ME (2005) Hydrophobins: the protein-amphiphiles of filamentous fungi. FEMS Microbiol Rev 29(5):877–896
Littlejohn KA, Hooley P, Cox PW (2012) Bioinformatics predicts diverse Aspergillus hydrophobins with novel properties. Food Hydrocolloids 27(2):503–516. https://doi.org/10.1016/j.foodhyd.2011.08.018
Lo VC, Ren Q, Pham CL, Morris VK, Kwan AH, Sunde M (2014) Fungal hydrophobin proteins produce self-assembling protein films with diverse structure and chemical stability. Nanomaterials (Basel, Switzerland) 4(3):827–843. https://doi.org/10.3390/nano4030827
Lugones LG, Bosscher JS, Scholtmeyer K, de Vries OM, Wessels JG (1996) An abundant hydrophobin (ABH1) forms hydrophobic rodlet layers in Agaricus bisporus fruiting bodies. Microbiology (Reading, England) 142(Pt 5):1321–1329. https://doi.org/10.1099/13500872-142-5-1321
Macindoe I, Kwan AH, Ren Q, Morris VK, Yang W, Mackay JP, Sunde M (2012) Self-assembly of functional, amphipathic amyloid monolayers by the fungal hydrophobin EAS. Proc Natl Acad Sci USA 109(14):E804–E811. https://doi.org/10.1073/pnas.1114052109
Mackay JP, Matthews JM, Winefield RD, Mackay LG, Haverkamp RG, Templeton MD (2001) The hydrophobin EAS is largely unstructured in solution and functions by forming amyloid-like structures. Structure (London, England: 1993) 9(2):83–91
Mankel A, Krause K, Kothe E (2002) Identification of a hydrophobin gene that is developmentally regulated in the ectomycorrhizal fungus Tricholoma terreum. Appl Environ Microbiol 68(3):1408–1413. https://doi.org/10.1128/aem.68.3.1408-1413.2002
Moonjely S, Keyhani NO, Bidochka MJ (2018) Hydrophobins contribute to root colonization and stress responses in the rhizosphere-competent insect pathogenic fungus Beauveria bassiana. Microbiology (Reading, England) 164(4):517–528. https://doi.org/10.1099/mic.0.000644
Morris VK, Kwan AH, Sunde M (2013) Analysis of the structure and conformational states of DewA gives insight into the assembly of the fungal hydrophobins. J Mol Biol 425(2):244–256. https://doi.org/10.1016/j.jmb.2012.10.021
Morris VK, Linser R, Wilde KL, Duff AP, Sunde M, Kwan AH (2012) Solid-state NMR spectroscopy of functional amyloid from a fungal hydrophobin: a well-ordered beta-sheet core amidst structural heterogeneity. Angew Chem Int Ed Engl 51(50):12621–12625. https://doi.org/10.1002/anie.201205625
Morris VK, Ren Q, Macindoe I, Kwan AH, Byrne N, Sunde M (2011) Recruitment of class I hydrophobins to the air: water interface initiates a multi-step process of functional amyloid formation. J Biol Chem 286(18):15955–15963. https://doi.org/10.1074/jbc.M110.214197
Niu B, Gong Y, Gao X, Xu H, Qiao M, Li W (2014) The functional role of Cys3–Cys4 loop in hydrophobin HGFI. Amino Acids 46(11):2615–2625. https://doi.org/10.1007/s00726-014-1805-0
Paris S, Debeaupuis JP, Crameri R, Carey M, Charles F, Prevost MC, Schmitt C, Philippe B, Latge JP (2003) Conidial hydrophobins of Aspergillus fumigatus. Appl Environ Microbiol 69(3):1581–1588
Pennacchio A, Cicatiello P, Notomista E, Giardina P, Piscitelli A (2018) New clues into the self-assembly of Vmh2, a basidiomycota class I hydrophobin. Biol Chem 399(8):895–901. https://doi.org/10.1515/hsz-2018-0124
Pham CL, Shanmugam N, Strange M, O’Carroll A, Brown JW, Sierecki E, Gambin Y, Steain M, Sunde M (2019) Viral M45 and necroptosis-associated proteins form heteromeric amyloid assemblies. EMBO reports 20(2). https://doi.org/10.15252/embr.201846518
Pham CLL, Kwan AH, Sunde M (2014) Functional amyloid: widespread in nature, diverse in purpose. In: Perrett S (ed) Amyloids in health and disease, vol 56. Essays in biochemistry, pp 207–219. https://doi.org/10.1042/bse0560207
Pham CLL, Rey A, Lo V, Soulès M, Ren Q, Meisl G, Knowles TPJ, Kwan AH, Sunde M (2016) Self-assembly of MPG1, a hydrophobin protein from the rice blast fungus that forms functional amyloid coatings, occurs by a surface-driven mechanism. Sci Rep 6:25288. https://doi.org/10.1038/srep25288
Pham CLL, Rodriguez de Francisco B, Valsecchi I, Dazzoni R, Pille A, Lo V, Ball SR, Cappai R, Wien F, Kwan AH, Guijarro JI, Sunde M (2018) Probing structural changes during self-assembly of surface-active hydrophobin proteins that form functional amyloids in fungi. J Mol Biol 430(20):3784–3801. https://doi.org/10.1016/j.jmb.2018.07.025
Pille A, Kwan AH, Cheung I, Hampsey M, Aimanianda V, Delepierre M, Latge JP, Sunde M, Guijarro JI (2015) (1)H, (13)C and (15)N resonance assignments of the RodA hydrophobin from the opportunistic pathogen Aspergillus fumigatus. Biomol NMR Assignments 9(1):113–118. https://doi.org/10.1007/s12104-014-9555-1
Ren Q, Kwan AH, Sunde M (2013) Two forms and two faces, multiple states and multiple uses: properties and applications of the self-assembling fungal hydrophobins. Biopolymers 100(6):601–612. https://doi.org/10.1002/bip.22259
Ren Q, Kwan AH, Sunde M (2014) Solution structure and interface-driven self-assembly of NC2, a new member of the class II hydrophobin proteins. Proteins 82(6):990–1003. https://doi.org/10.1002/prot.24473
Sallada ND, Dunn KJ, Berger BW (2018) A structural and functional role for disulfide bonds in a class II hydrophobin. Biochemistry 57(5):645–653. https://doi.org/10.1021/acs.biochem.7b01166
Sammer D, Krause K, Gube M, Wagner K, Kothe E (2016) Hydrophobins in the life cycle of the ectomycorrhizal basidiomycete Tricholoma vaccinum. PLoS ONE 11(12):e0167773. https://doi.org/10.1371/journal.pone.0167773
Scherrer S, De Vries OM, Dudler R, Wessels JG, Honegger R (2000) Interfacial self-assembly of fungal hydrophobins of the lichen-forming ascomycetes Xanthoria parietina and X. ectaneoides. Fungal Genet Biol FG & B 30(1):81–93. https://doi.org/10.1006/fgbi.2000.1205
Scherrer S, Haisch A, Honegger R (2002) Characterization and expression of XPH1, the hydrophobin gene of the lichen-forming ascomycete Xanthoria parietina. New Phytol 154(1):175–184. https://doi.org/10.1046/j.1469-8137.2002.00351.x
Seidl-Seiboth V, Gruber S, Sezerman U, Schwecke T, Albayrak A, Neuhof T, von Dohren H, Baker SE, Kubicek CP (2011) Novel hydrophobins from Trichoderma define a new hydrophobin subclass: protein properties, evolution, regulation and processing. J Mol Evol 72(4):339–351. https://doi.org/10.1007/s00239-011-9438-3
Seyedmousavi S, Bosco SMG, de Hoog S, Ebel F, Elad D, Gomes RR, Jacobsen ID, Jensen HE, Martel A, Mignon B, Pasmans F, Pieckova E, Rodrigues AM, Singh K, Vicente VA, Wibbelt G, Wiederhold NP, Guillot J (2018) Fungal infections in animals: a patchwork of different situations. Medical mycology 56 (suppl_1):165–187. https://doi.org/10.1093/mmy/myx104
Shams-Ghahfarokhi M, Aghaei-Gharehbolagh S, Aslani N, Razzaghi-Abyaneh M (2014) Investigation on distribution of airborne fungi in outdoor environment in Tehran, Iran. J Environ Health Sci Eng 12(1):54–54. https://doi.org/10.1186/2052-336x-12-54
Shanmugam N, Baker MODG, Ball SR, Steain M, Pham CLL, Sunde M (2019) Microbial functional amyloids serve diverse purposes for structure, adhesion and defence. Biophysical Reviews. https://doi.org/10.1007/s12551-019-00526-1
Skamnioti P, Gurr SJ (2008) Cutinase and hydrophobin interplay: a herald for pathogenesis? Plant Signaling Behav 3(4):248–250. https://doi.org/10.4161/psb.3.4.5181
Skamnioti P, Gurr SJ (2009) Against the grain: safeguarding rice from rice blast disease. Trends Biotechnol 27(3):141–150. https://doi.org/10.1016/j.tibtech.2008.12.002
Stivala A, Wybrow M, Wirth A, Whisstock JC, Stuckey PJ (2011) Automatic generation of protein structure cartoons with Pro-origami. Bioinformatics (Oxford, England) 27(23):3315–3316. https://doi.org/10.1093/bioinformatics/btr575
Stringer MA, Dean RA, Sewall TC, Timberlake WE (1991) Rodletless, a new Aspergillus developmental mutant induced by directed gene inactivation. Genes Dev 5(7):1161–1171. https://doi.org/10.1101/gad.5.7.1161
Sunde M, Kwan AH, Templeton MD, Beever RE, Mackay JP (2008) Structural analysis of hydrophobins. Micron 39(7):773–784
Takeo K (1976) Existence of a surface configuration on the aerial spore and aerial mycelium of micropolyspora. J Gen Microbiol 95(1):17–26. https://doi.org/10.1099/00221287-95-1-17
Talbot NJ, Ebbole DJ, Hamer JE (1993) Identification and characterization of MPG1, a gene involved in pathogenicity from the rice blast fungus Magnaporthe grisea. Plant Cell 5(11):1575–1590. https://doi.org/10.1105/tpc.5.11.1575
Talbot NJ, Kershaw MJ, Wakley GE, De Vries O, Wessels J, Hamer JE (1996) MPG1 encodes a fungal hydrophobin involved in surface interactions during infection-related development of Magnaporthe grisea. Plant Cell 8(6):985–999. https://doi.org/10.1105/tpc.8.6.985
Templeton MD, Greenwood DR, Beever RE (1995) Solubilization of neurospora crassa rodlet proteins and identification of the predominant protein as the proteolytically processed eas (ccg-2) gene product. Exp Mycol 19(2):166–169
Thau N, Monod M, Crestani B, Rolland C, Tronchin G, Latge JP, Paris S (1994) Rodletless mutants of Aspergillus fumigatus. Infect Immun 62(10):4380–4388
Trembley ML, Ringli C, Honegger R (2002) Hydrophobins DGH1, DGH2, and DGH3 in the lichen-forming basidiomycete Dictyonema glabratum. Fungal Genet Biol FG & B 35(3):247–259. https://doi.org/10.1006/fgbi.2001.1325
Valsecchi I, Dupres V, Michel JP, Duchateau M, Matondo M, Chamilos G, Saveanu C, Guijarro JI, Aimanianda V, Lafont F, Latge JP, Beauvais A (2019a) The puzzling construction of the conidial outer layer of Aspergillus fumigatus. Cell Microbiol 21(5):e12994. https://doi.org/10.1111/cmi.12994
Valsecchi I, Dupres V, Stephen-Victor E, Guijarro JI, Gibbons J, Beau R, Bayry J, Coppee J-Y, Lafont F, Latgé J-P, Beauvais A (2017) Role of hydrophobins in Aspergillus fumigatus. J Fungi (Basel, Switzerland) 4(1):2. https://doi.org/10.3390/jof4010002
Valsecchi I, Lai JI, Stephen-Victor E, Pillé A, Beaussart A, Lo V, Pham CLL, Aimanianda V, Kwan AH, Duchateau M, Gianetto QG, Matondo M, Lehoux M, Sheppard DC, Dufrene YF, Bayry J, Guijarro JI, Sunde M, Latgé J-P (2019b) Assembly and disassembly of Aspergillus fumigatus conidial rodlets. Cell Surface 5:100023. https://doi.org/10.1016/j.tcsw.2019.100023
van de Veerdonk FL, Gresnigt MS, Romani L, Netea MG, Latge JP (2017) Aspergillus fumigatus morphology and dynamic host interactions. Nat Rev Microbiol 15(11):661–674. https://doi.org/10.1038/nrmicro.2017.90
van Wetter MA, Wosten HA, Wessels JG (2000) SC3 and SC4 hydrophobins have distinct roles in formation of aerial structures in dikaryons of Schizophyllum commune. Mol Microbiol 36(1):201–210
Wang X, De Vocht ML, De Jonge J, Poolman B, Robillard GT (2002) Structural changes and molecular interactions of hydrophobin SC3 in solution and on a hydrophobic surface. Protein Sci 11(5):1172–1181
Wang X, Graveland-Bikker JF, de Kruif CG, Robillard GT (2004) Oligomerization of hydrophobin SC3 in solution: from soluble state to self-assembly. Protein Sci Publ Protein Soc 13(3):810–821
Wessels J, De Vries O, Asgeirsdottir SA, Schuren F (1991) Hydrophobin genes involved in formation of aerial hyphae and fruit bodies in Schizophyllum. Plant Cell 3(8):793–799. https://doi.org/10.1105/tpc.3.8.793
Wessels JGH, Kreger DR, Marchant R, Regensburg BA, De Vries OMH (1972) Chemical and morphological characterization of the hyphal wall surface of the basidiomycete Schizophyllum commune. Biochim et Biophys Acta (BBA)—Gen Subj 273(2):346–358. https://doi.org/10.1016/0304-4165(72)90226-7
Whiteford JR, Spanu PD (2001) The hydrophobin HCf-1 of Cladosporium fulvum is required for efficient water-mediated dispersal of conidia. Fungal Genet Biol 32(3):159–168. https://doi.org/10.1006/fgbi.2001.1263
Wong SSW, Aimanianda V (2017) Host soluble mediators: defying the immunological inertness of Aspergillus fumigatus conidia. J Fungi (Basel, Switzerland) 4(1). https://doi.org/10.3390/jof4010003
Wosten H, De Vries O, Wessels J (1993) Interfacial self-assembly of a fungal hydrophobin into a hydrophobic rodlet layer. Plant Cell 5(11):1567–1574. https://doi.org/10.1105/tpc.5.11.1567
Wosten HA, Scholtmeijer K (2015) Applications of hydrophobins: current state and perspectives. Appl Microbiol Biotechnol 99(4):1587–1597. https://doi.org/10.1007/s00253-014-6319-x
Wosten HA, Schuren FH, Wessels JG (1994) Interfacial self-assembly of a hydrophobin into an amphipathic protein membrane mediates fungal attachment to hydrophobic surfaces. EMBO J 13(24):5848–5854
Wosten HA, van Wetter MA, Lugones LG, van der Mei HC, Busscher HJ, Wessels JG (1999) How a fungus escapes the water to grow into the air. Curr Biol CB 9(2):85–88
Wosten HAB, de Vocht ML (2000) Hydrophobins, the fungal coat unravelled. Biochim Et Biophys Acta-Rev Biomembr 1469(2):79–86
Yang W, Ren Q, Wu YN, Morris VK, Rey AA, Braet F, Kwan AH, Sunde M (2013) Surface functionalization of carbon nanomaterials by self-assembling hydrophobin proteins. Biopolymers 99(1):84–94. https://doi.org/10.1002/bip.22146
Zampieri F, Wosten HAB, Scholtmeijer K (2010) Creating surface properties using a palette of hydrophobins. Materials (Basel, Switzerland) 3(9):4607–4625. https://doi.org/10.3390/ma3094607
Zhang S, Xia YX, Kim B, Keyhani NO (2011) Two hydrophobins are involved in fungal spore coat rodlet layer assembly and each play distinct roles in surface interactions, development and pathogenesis in the entomopathogenic fungus, Beauveria bassiana. Mol Microbiol 80(3):811–826. https://doi.org/10.1111/j.1365-2958.2011.07613.x
Zykwinska A, Guillemette T, Bouchara JP, Cuenot S (2014a) Spontaneous self-assembly of SC3 hydrophobins into nanorods in aqueous solution. Biochem Biophys Acta 1844 7:1231–1237. https://doi.org/10.1016/j.bbapap.2014.04.003
Zykwinska A, Pihet M, Radji S, Bouchara JP, Cuenot S (2014b) Self-assembly of proteins into a three-dimensional multilayer system: investigation of the surface of the human fungal pathogen Aspergillus fumigatus. Biochem Biophys Acta 1844 6:1137–1144. https://doi.org/10.1016/j.bbapap.2014.03.001
Acknowledgements
The authors acknowledge funding from the Australian Research Council over many years, for research into hydrophobin proteins, which has contributed to this work (DP0879121, DP120100756 and DP150104227) and support from the French-Australian Science and Technology (FAST) Program (FR110012). SRB is supported by the Australian Government in the form of a Research Training Program Scholarship. The authors wish to thank Chi Pham, Victor Lo, Vanessa Morris, Qin Ren, Jennifer Lai, Ingrid Macindoe and all other researchers and students working in the Sunde and Kwan Laboratories who have contributed to hydrophobin projects. We acknowledge the facilities and the scientific and technical assistance of Sydney Microscopy and Microanalysis at the Australian Centre for Microscopy and Microanalysis at the University of Sydney. We thank Chiara Neto for use of atomic force microscopy facilities.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Ball, S.R., Kwan, A.H., Sunde, M. (2019). Hydrophobin Rodlets on the Fungal Cell Wall. In: Latgé, JP. (eds) The Fungal Cell Wall . Current Topics in Microbiology and Immunology, vol 425. Springer, Cham. https://doi.org/10.1007/82_2019_186
Download citation
DOI: https://doi.org/10.1007/82_2019_186
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-49927-3
Online ISBN: 978-3-030-49928-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)