Abstract
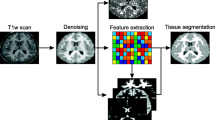
This study presents a novel automatic approach for the identification of anatomical brain structures in magnetic resonance images (MRI). The method combines a fast multiscale multi-channel three dimensional (3D) segmentation algorithm providing a rich feature vocabulary together with a support vector machine (SVM) based classifier. The segmentation produces a full hierarchy of segments, expressed by an irregular pyramid with only linear time complexity. The pyramid provides a rich, adaptive representation of the image, enabling detection of various anatomical structures at different scales. A key aspect of the approach is the thorough set of multiscale measures employed throughout the segmentation process which are also provided at its end for clinical analysis. These features include in particular the prior probability knowledge of anatomic structures due to the use of an MRI probabilistic atlas. An SVM classifier is trained based on this set of features to identify the brain structures. We validated the approach using a gold standard real brain MRI data set. Comparison of the results with existing algorithms displays the promise of our approach.
Chapter PDF
Similar content being viewed by others
Keywords
- Support Vector Machine
- Gray Matter
- Magnetic Resonance Image Data
- Candidate Segment
- Magnetic Resonance Image Segmentation
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
References
Pham, D., Xu, C., Prince, J.: Current methods in medical image segmentation. Annual Review of Biomedical Engineering 2, 315–337 (2000)
Rajapakse, J., Kruggel, F.: Segmentation of MR images with intensity inhomogeneities. IVC 16(3), 165–180 (1998)
Shattuck, D.W., Sandor-Leahy, S.R., Schaper, K.A., Rottenberg, D.A., Leahy, R.M.: Magnetic resonance image tissue classification using a partial volume model. Neuroimage 13(5), 856–876 (2001)
Bezdek, J.C., Hall, L.O., Clarke, L.P.: Review of MRI segmentation techniques using pattern recognition. Medical Physics 20(4), 1033–1048 (1993)
Sonka, M.M., Fitzpatrick, J.M. (eds.): Handbook of Medical Imaging. SPIE (2000)
Zijdenbos, A., Dawant, B.: Brain segmentation and white matter lesion detection in MRI. Critical Reviews in Biomedical Engineering 22 (1994)
Zeng, X., Staib, L.H., Schultz, R.T., Duncan, J.S.: Segmentation and measurement of the cortex from 3D MRI using coupled surfaces propagation. IEEE MI (1999)
Pham, D., Prince, J.: Robust unsupervised tissue classification in MRI. IEEE Biomedical Imaging: Macro to Nano (2004)
Wells, W.M., Grimson, W., Kikinis, R., Jolesz, F.A.: Adaptive segmentation of MRI data. IEEE MI 15, 429–442 (1996)
Zhang, Y., Brady, M., Smith, S.: Segmentation of brain MRI through a hidden markov random field model and the expectation-maximization algorithm. IEEE Medical Imaging 20(1), 45–57 (2001)
Van-Leemput, K., Maes, F., Vandermeulen, D., Colcher, A., Suetens, P.: Automated segmentation of MS by model outlier detection. IEEE MI 20, 677–688 (2001)
Akselrod-Ballin, A., Galun, M., Gomori, J.M., Fillipi, M., Valsasina, P., Brandt, A., Basri, R.: An integrated segmentation and classification approach applied to multiple sclerosis analysis. In: CVPR (2006)
Sharon, E., Brandt, A., Basri, R.: Fast multiscale image segmentation. In: CVPR, pp. 70–77 (2000)
Galun, M., Sharon, E., Basri, R., Brandt, A.: Texture segmentation by multiscale aggregation of filter responses and shape elements. In: ICCV, pp. 716–723 (2003)
Brandt, A., McCormick, S., Ruge, J. (eds.): Algebraic multigrid (AMG) for automatic multigrid solution with application to geodetic computations. Inst. for Computational Studies, POB 1852, Fort Collins, Colorado (1982)
Frackowiak, S., Friston, K., Frith, C., Dolan, R., Price, C., Zeki, S., Ashburner, J., Penny, W. (eds.): Human Brain Function. Academic Press, London (2003)
Vapnik, V. (ed.): The Nature of Statistical Learning Theory. Springer, Heidelberg (1995)
Burkhardt, J.: A markov chain monte carlo algorithm for the segmentation of t1-mr-images of the head. Diploma thesis, Technische Universitat Munchen (2003)
Gerig, G., Jomier, M., Chakos, M.: Valmet: A new validation tool for assessing and improving 3D object segmentation. In: Niessen, W.J., Viergever, M.A. (eds.) MICCAI 2001. LNCS, vol. 2208, pp. 516–523. Springer, Heidelberg (2001)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2006 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Akselrod-Ballin, A., Galun, M., Gomori, M.J., Basri, R., Brandt, A. (2006). Atlas Guided Identification of Brain Structures by Combining 3D Segmentation and SVM Classification. In: Larsen, R., Nielsen, M., Sporring, J. (eds) Medical Image Computing and Computer-Assisted Intervention – MICCAI 2006. MICCAI 2006. Lecture Notes in Computer Science, vol 4191. Springer, Berlin, Heidelberg. https://doi.org/10.1007/11866763_26
Download citation
DOI: https://doi.org/10.1007/11866763_26
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-44727-6
Online ISBN: 978-3-540-44728-3
eBook Packages: Computer ScienceComputer Science (R0)