Abstract
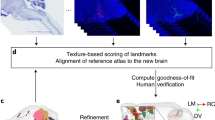
Associating specific gene activity with functional locations in the brain results in a greater understanding of the role of the gene. To perform such an association for the over 20,000 genes in the mammalian genome, reliable automated methods that characterize the distribution of gene expression in relation to a standard anatomical model are required. In this work, we propose a new automatic method that results in the segmentation of gene expression images into distinct anatomical regions in which the expression can be quantified and compared with other images. Our method utilizes shape models from training images, texture differentiation at region boundaries, and features of anatomical landmarks, to deform a subdivision mesh-based atlas to fit gene expression images. The subdivision mesh provides a common coordinate system for internal brain data through which gene expression patterns can be compared across images. The automated large-scale annotation will help scientists interpret gene expression patterns at cellular resolution more efficiently.
Chapter PDF
Similar content being viewed by others
References
Waterston, R., and the Mouse Genome Sequencing Consortium: Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002)
Albrecht, U., Lu, H.C., Revelli, J.P., Xu, X.C., Lotan, R., Eichele, G.: Studying Gene Expression on Tissue Sections Using In Situ Hybridization. In: Adolph, K.W. (ed.) Human Genome Methods, pp. 93–119. CRC Press, Boca Raton (1997)
Carson, J.P., Thaller, C., Eichele, G.: A transcriptome atlas of the mouse brain at cellular resolution. Curr. Opin. Neurobiol. 12, 562–565 (2002)
Ju, T., Warren, J., Eichele, G., Thaller, C., Chiu, W., Carson, J.: A geometric database for gene expression data. In: Eurographics Symposium on Geometry Processing, Aachen, Germany, June 2003, pp. 166–176 (2003)
Warren, J., Weimer, H.: Subdivision Methods for Geometric Design: A Constructive Approach. Morgan Kaufmann Publishers, San Francisco (2002)
Kakadiaris, I.A., Bello, M., Arunachalam, S., Kang, W., Ju, T., Warren, J., Carson, J., Chiu, W., Thaller, C., Eichele, G.: Landmark-driven, atlas-based segmentation of mouse brain tissue images containing gene expression data. In: Proc. 7th International Conference on Medical Image Computing and Computer-Assisted Intervention, Rennes, France, pp. 192–199 (2004)
Vapnik, V.N.: The Nature of Statistical Learning Theory. Springer, Heidelberg (2000)
Laws, K.: Textured Image Segmentation. PhD thesis, USC (1980)
Aksoy, S., Haralick, R.: Feature normalization and likelihood-based similarity measures for image retrieval. Pattern Recognition Letters 22, 563–582 (2001)
Guyon, I., Elisseeff, A.: An introduction to variable and feature selection. Journal of Machine Learning Research 3, 1157–1182 (2003)
Fayyad, U.M., Irani, K.B.: On the handling of continuous-valued attributes in decision tree generation. Machine Learning 8, 87–102 (1992)
Schölkopf, B., Smola, A.: Learning with Kernels. MIT Press, Cambridge (2002)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2005 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Bello, M. et al. (2005). Hybrid Segmentation Framework for Tissue Images Containing Gene Expression Data. In: Duncan, J.S., Gerig, G. (eds) Medical Image Computing and Computer-Assisted Intervention – MICCAI 2005. MICCAI 2005. Lecture Notes in Computer Science, vol 3749. Springer, Berlin, Heidelberg. https://doi.org/10.1007/11566465_32
Download citation
DOI: https://doi.org/10.1007/11566465_32
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-29327-9
Online ISBN: 978-3-540-32094-4
eBook Packages: Computer ScienceComputer Science (R0)