Abstract
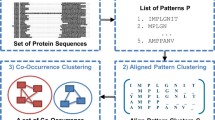
A basic task in protein analysis is to discover a set of sequence patterns that characterizes the function of a protein family. To address this task, we introduce a synthesized pattern representation called Aligned Pattern (AP) Cluster to discover potential functional segments in protein sequences. We apply our algorithm to identify and display the binding segments for the Cytochrome C. and Ubiquitin protein families. The resulting AP Clusters correspond to protein binding segments that surround the binding residues. When compared to the results from the protein annotation databases, PROSITE and pFam, ours are more efficient in computation and comprehensive in quality. The significance of the AP Cluster is that it is able to capture subtle variations of the binding segments in protein families. It thus could help to reduce time-consuming simulations and experimentation in the protein analysis.
Chapter PDF
Similar content being viewed by others
Keywords
References
Thompson, J.D., Higgins, D.G., Gibson, T.J.: Clustal w: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22(22), 4673–4680 (1994)
Notredame, C., Higgins, D.G., Heringa, J.: T-coffee: A novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 302(1), 205–217 (2000)
Subramanian, A.R., Kaufmann, A.M., Morgenstern, B.: Dialign-tx: greedy and progressive approaches for segment-based multiple sequence alignment. Algorithms Mol. Biol. 3, 6 (2008)
Durbin, R., Eddy, S.R., Krogh, A., Mitchison, G.: Biological Sequence Analysis: Probabilistic models of proteins and nucleic acids. Cambridge University Press (1998)
Wang, L., Jiang, T.: On the complexity of multiple sequence alignment. Journal of Computational Biology 1(4), 337–348 (1994)
Frith, M.C., Hansen, U., Spouge, J.L., Weng, Z.: Finding functional sequence elements by multiple local alignment. Nucleic Acids Res. 32(1), 189–200 (2004)
Bailey, T.L., Elkan, C.: Unsupervised learning of multiple motifs in biopolymers using expectation maximization. Machine Learning 21(1/2), 51–80 (1995)
Pisanti, N., Crochemore, M., Grossi, R., Sagot, M.F.: Bases of motifs for generating repeated patterns with wild cards. IEEE/ACM Transactions on Computational BIology and Bioinformatics 2(1), 40–50 (2005)
Lee, E.-S.A., Wong, A.K.C.: Synthesizing aligned random pattern digraphs from protein sequence patterns. In: Bioinformatics and Biomedicine Workshops (BIBMW), pp. 178–185 (2011)
Bairoch, A.: Prosite: a dictionary of sites and patterns in proteins. Nucleic Acids Research 19, 2241–2245 (1991)
Sigrist, C.J.A., Cerutti, L., de Castro, E., Langendijk-Genevaux, P.S., Bulliard, V., Bairoch, A., Hulo, N.: Prosite, a protein domain database for functional characterization and annotation. Nucleic Acids Res. 38(Database issue), 161–166 (2010)
Sonnhammer, E.L., Eddy, S.R., Durbin, R.: Pfam: A comprehensive database of protein domain families based on seed alignments. PROTEINS: Structure, Function, and Genetics 28, 405–420 (1997)
Finn, R.D., Mistry, J., Tate, J., Coggill, P., Heger, A., Pollington, J.E., Gavin, O.L., Gunasekaran, P., Ceric, G., Forslund, K., Holm, L., Sonnhammer, E.L., Eddy, S.R., Bateman, A.: The pfam protein families database. Nucleic Acids Research 211, D211–D222 (2010)
Peng, J., Schwartz, Elias, Thoreen, Cheng, Marsischky, Roelofs, et al.: A proteomics approach to understanding protein ubiquitination. Nature Biotechnology 21(8), 921–926 (2003)
Xu, P.P.: Characterization of polyubiquitin chain structure by middle-down mass spectrometry. Analytical Chemistry 80(9), 3438–3444 (2008)
Kirisako, T., Kamei, K., Kato, M., Fukumoto, Kanie, Sano, Tokunaga: A ubiquitin ligase complex assembles linear polyubiquitin chains. The EMBO Journal 25(20), 4877–4887 (2006)
Kim, H., Kim, Lledias, Kisselev, S., Skowyra, Gygi, Goldberg: Goldberg: Certain pairs of ubiquitin-conjugating enzymes (e2s) and ubiquitin-protein ligases (e3s) synthesize condegradable forked ubiquitin chains containing all possible isopeptide linkages. The Journal of Biological Chemistry 282(24), 17375–17386 (2007)
Ikeda, F.: Dikic: Atypical ubiquitin chains: new molecular signals. ’protein modifications: Beyond the usual suspects’ review series. EMBO Reports 9 (6), 536–542 (2008)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Lee, ES.A., Zhuang, D., Wong, A.K.C. (2012). Aligning Discovered Patterns from Protein Family Sequences. In: Shibuya, T., Kashima, H., Sese, J., Ahmad, S. (eds) Pattern Recognition in Bioinformatics. PRIB 2012. Lecture Notes in Computer Science(), vol 7632. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-34123-6_22
Download citation
DOI: https://doi.org/10.1007/978-3-642-34123-6_22
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-34122-9
Online ISBN: 978-3-642-34123-6
eBook Packages: Computer ScienceComputer Science (R0)