Abstract
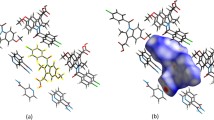
We present a novel approach to crystallographic ligand density interpretation based on Zernike shape descriptors. Electron density for a bound ligand is expanded in an orthogonal polynomial series (3D Zernike polynomials) and the coefficients from this expansion are employed to construct rotation-invariant descriptors. These descriptors can be compared highly efficiently against large databases of descriptors computed from other molecules. In this manuscript we describe this process and show initial results from an electron density interpretation study on a dataset containing over a hundred OMIT maps. We could identify the correct ligand as the first hit in about 30 % of the cases, within the top five in a further 30 % of the cases, and giving rise to an 80 % probability of getting the correct ligand within the top ten matches. In all but a few examples, the top hit was highly similar to the correct ligand in both shape and chemistry. Further extensions and intrinsic limitations of the method are discussed.
Chapter PDF
Similar content being viewed by others
Keywords
References
Perrakis, A., Morris, R., Lamzin, V.S.: Automated protein model building combined with iterative structure refinement. Nat. Struct. Biol. 6(5), 458–463 (1999)
Terwilliger, T.C.: Solve and resolve: automated structure solution and density modification. Methods Enzymol. 374, 22–37 (2003)
Adams, P.D., Grosse-Kunstleve, R.W., Hung, L.W., Ioerger, T.R., McCoy, A.J., Moriarty, N.W., Read, R.J., Sacchettini, J.C., Sauter, N.K., Terwilliger, T.C.: Phenix: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58(Pt 11), 1948–1954 (2002)
Cowtan, K.: The buccaneer software for automated model building. 1. tracing protein chains. Acta Crystallogr. D Biol. Crystallogr. 62(Pt 9), 1002–1011 (2006)
Morris, R.J.: Statistical pattern recognition for macromolecular crystallographers. Acta Crystallogr. D Biol. Crystallogr. 60(Pt 12 Pt 1), 2133–2143 (2004)
Greer, J.: Computer skeletonization and automatic electron density map analysis. Methods Enzymol. 115, 206–224 (1985)
Cowtan, K.: Fast fourier feature recognition. Acta Crystallogr. D Biol. Crystallogr. 57(Pt 10), 1435–1444 (2001)
Cowtan, K.: Fitting molecular fragments into electron density. Acta Crystallogr. D Biol. Crystallogr. 64(Pt 1), 83–89 (2008)
Morris, R.J., Perrakis, A., Lamzin, V.S.: ARP/wARP model-building algorithms. i. the main chain. Acta Crystallogr. D Biol. Crystallogr. 58(Pt 6 Pt 2), 968–975 (2002)
Morris, R.J., Perrakis, A., Lamzin, V.S.: ARP/wARP and automatic interpretation of protein electron density maps. Methods Enzymol. 374, 229–244 (2003)
Morris, R.J., Zwart, P.H., Cohen, S., Fernandez, F.J., Kakaris, M., Kirillova, O., Vonrhein, C., Perrakis, A., Lamzin, V.S.: Breaking good resolutions with ARP/wARP. J. Synchrotron. Radiat. 11(Pt 1), 56–59 (2004)
Adams, P.D., Gopal, K., Grosse-Kunstleve, R.W., Hung, L.W., Ioerger, T.R., McCoy, A.J., Moriarty, N.W., Pai, R.K., Read, R.J., Romo, T.D., Sacchettini, J.C., Sauter, N.K., Storoni, L.C., Terwilliger, T.C.: Recent developments in the phenix software for automated crystallographic structure determination. J. Synchrotron. Radiat. 11(Pt 1), 53–55 (2004)
Terwilliger, T.C.: Automated structure solution, density modification and model building. Acta Crystallogr. D Biol. Crystallogr. 58(Pt 11), 1937–1940 (2002)
Terwilliger, T.: Solve and resolve: automated structure solution, density modification and model building. J. Synchrotron. Radiat. 11(Pt 1), 49–52 (2004)
Cohen, S.X., Morris, R.J., Fernandez, F.J., Ben Jelloul, M., Kakaris, M., Parthasarathy, V., Lamzin, V.S., Kleywegt, G.J., Perrakis, A.: Towards complete validated models in the next generation of ARP/wARP. Acta Crystallogr. D Biol. Crystallogr. 60(Pt 12 Pt 1), 2222–2229 (2004)
Terwilliger, T.C., Grosse-Kunstleve, R.W., Afonine, P.V., Moriarty, N.W., Zwart, P.H., Hung, L.W., Read, R.J., Adams, P.D.: Iterative model building, structure refinement and density modification with the phenix autobuild wizard. Acta Crystallogr. D Biol. Crystallogr. 64(Pt 1), 61–69 (2008)
Joosten, K., Cohen, S.X., Emsley, P., Mooij, W., Lamzin, V.S., Perrakis, A.: A knowledge-driven approach for crystallographic protein model completion. Acta Crystallogr. D Biol. Crystallogr. 64(Pt 4), 416–424 (2008)
Terwilliger, T.C., Grosse-Kunstleve, R.W., Afonine, P.V., Moriarty, N.W., Adams, P.D., Read, R.J., Zwart, P.H., Hung, L.W.: Iterative-build omit maps: map improvement by iterative model building and refinement without model bias. Acta Crystallogr. D Biol. Crystallogr. 64(Pt 5), 515–524 (2008)
Morris, R.J., Bricogne, G.: Sheldrick’s 1.2 Å rule and beyond. Acta Crystallogr. D Biol. Crystallogr. 59(Pt 3), 615–617 (2003)
Morris, R.J., Blanc, E., Bricogne, G.: On the interpretation and use of < |E|2 > (d*) profiles. Acta Crystallogr. D Biol. Crystallogr. 60(Pt 2), 227–240 (2004)
Zwart, P.H., Langer, G.G., Lamzin, V.S.: Modelling bound ligands in protein crystal structures. Acta Crystallogr. D Biol. Crystallogr. 60(Pt 12 Pt 1), 2230–2239 (2004)
Evrard, G.X., Langer, G.G., Perrakis, A., Lamzin, V.S.: Assessment of automatic ligand building in arp/warp. Acta Crystallogr. D Biol. Crystallogr. 63(Pt 1), 108–117 (2007)
Terwilliger, T.C.: Maximum-likelihood density modification using pattern recognition of structural motifs. Acta Crystallogr. D Biol. Crystallogr. 57(Pt 12), 1755–1762 (2001)
Terwilliger, T.C., Klei, H., Adams, P.D., Moriarty, N.W., Cohn, J.D.: Automated ligand fitting by core-fragment fitting and extension into density. Acta Crystallogr. D Biol. Crystallogr. 62(Pt 8), 915–922 (2006)
Terwilliger, T.C., Adams, P.D., Moriarty, N.W., Cohn, J.D.: Ligand identification using electron-density map correlations. Acta Crystallogr. D Biol. Crystallogr. 63(Pt 1), 101–107 (2007)
Mooij, W.T.M., Hartshorn, M.J., Tickle, I.J., Sharff, A.J., Verdonk, M.L., Jhoti, H.: Automated protein-ligand crystallography for structure-based drug design. Chem. Med. Chem. 1(8), 827–838 (2006)
Wlodek, S., Skillman, A.G., Nicholls, A.: Automated ligand placement and refinement with a combined force field and shape potential. Acta Crystallogr. D Biol. Crystallogr. 62(Pt 7), 741–749 (2006)
Aishima, J., Russel, D.S., Guibas, L.J., Adams, P.D., Brunger, A.T.: Automated crystallographic ligand building using the medial axis transform of an electron-density isosurface. Acta Crystallogr. D Biol. Crystallogr. 61(Pt 10), 1354–1363 (2005)
Hattne, J., Lamzin, V.S.: Pattern-recognition-based detection of planar objects in three-dimensional electron-density maps. Acta Crystallogr. D Biol. Crystallogr. D64(Pt 8), 834–842 (2008)
Mak, L., Grandison, S., Morris, R.J.: An extension of spherical harmonics to region-based rotationally invariant descriptors for molecular shape description and comparison. J. Mol. Graph. Model 26(7), 1035–1045 (2008)
Sael, L., Li, B., La, D., Fang, Y., Ramani, K., Rustamov, R., Kihara, D.: Fast protein tertiary structure retrieval based on global surface shape similarity. Proteins (2008)
Grandison, S., Roberts, C., Morris, R.J.: The application of 3d zernike moments for the description of “model-free” molecular structure, functional motion, and structural reliability. J. Comput. Biol. 16(3), 487–500 (2009)
Cowtan, K.: The clipper C++ libraries for x-ray crystallography. IUCr Computing Commission Newsletter 2, 4–9 (2003)
Revol-Muller, C., Peyrin, F., Carrillon, Y., Odet, C.: Automated 3d region growing algorithm based on an assessment function. Pattern Recogn. Lett. 23(1-3), 137–150 (2002)
Canterakis, N.: 3D zernike moments and zernike affine invariants for 3D image analysis and recognition. In: Scandinavian Conference on Image Analysis (1999)
Novotni, M., Klein, R.: Shape retrieval using 3d zernike descriptors. Computer-Aided Design 36(11), 1047–1062 (2004)
Sael, L., La, D., Li, B., Rustamov, R., Kihara, D.: Rapid comparison of properties on protein surface. Proteins 73(1), 1–10 (2008)
Morris, R.J., Najmanovich, R.J., Kahraman, A., Thornton, J.M.: Real spherical harmonic expansion coefficients as 3d shape descriptors for protein binding pocket and ligand comparisons. Bioinformatics 21(10), 2347–2355 (2005)
Morris, R.J.: An evaluation of spherical designs for molecular-like surfaces. J. Mol. Graph. Model 24(5), 356–361 (2006)
Mathar, R.J.: Third order newton’s method for zernike polynomial zeros, arXiv: math.NA/0705.1329 (2007)
Mathar, R.J.: Zernike basis to cartesian transformations, arXiv: 0809.2368 math-ph (2008)
Kleywegt, G.J.: Crystallographic refinement of ligand complexes. Acta Crystallogr. D Biol. Crystallogr. 63(Pt 1), 94–100 (2007)
Funkhouser, T., Glaser, F., Laskowski, R.A., Morris, R.J., Najmanovich, R., Stockwell, G., Thornton, J.M.: Shape-based classification of bound ligands. In: Barber, S., Baxter, P.D., Mardia, K.V., Wells, R.E. (eds.) Quantitative Biology Shape Analysis and Wavelets, pp. 39–42 (2005)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2009 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Gunasekaran, P., Grandison, S., Cowtan, K., Mak, L., Lawson, D.M., Morris, R.J. (2009). Ligand Electron Density Shape Recognition Using 3D Zernike Descriptors. In: Kadirkamanathan, V., Sanguinetti, G., Girolami, M., Niranjan, M., Noirel, J. (eds) Pattern Recognition in Bioinformatics. PRIB 2009. Lecture Notes in Computer Science(), vol 5780. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-04031-3_12
Download citation
DOI: https://doi.org/10.1007/978-3-642-04031-3_12
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-04030-6
Online ISBN: 978-3-642-04031-3
eBook Packages: Computer ScienceComputer Science (R0)