Abstract
Rheumatoid factors (RFs) are autoantibodies that recognize the fragment crystallizable (Fc) region of immunoglobulin G (IgG). Genetically diverse RFs are produced in rheumatoid arthritis patients; however, in hematologic diseases, such as cryoglobulinemia and B cell lymphoma, RFs from a limited combination of heavy chain V-region genes and J-region genes are produced in large quantities and forms immune complexes with IgG. These genetically limited RFs have historically been used for the immunochemical characterization of RFs. Among them, RFs derived from the heavy-chain germline gene IGHV1-69 are the most common. Recently, the crystal structure of an IGHV1-69-derived RF named YES8c was elucidated in complex with human IgG1-Fc. Based on the structure and mutant analyses, a recognition mechanism for the autoantigen (IgG-Fc) common to IGHV1-69-derived RFs was proposed. This review summarizes the immunochemical character of the IGHV1-69-derived RFs, and then focuses on the recognition mechanism of the IGHV1-69-derived RFs, referring the structural features of the IGHV1-69-derived neutralizing antibodies.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
- Autoantibody
- B cell lymphoma
- Cryoglobulinemia
- Crystal structure
- Immunoglobulin-G (IgG)
- Molecular recognition
- Rheumatoid factor
1 Rheumatoid Factors (RF)
RFs are autoantibodies that recognize the fragment crystallizable (Fc) region of immunoglobulin G (IgG). RFs have been used as important diagnostic markers of rheumatoid arthritis (RA); an autoimmune disease characterized by the chronic joint inflammation leading to systemic inflammation (van Delft and Huizinga 2020). RFs are detected not only in RA but also in other inflammatory diseases or in the serum of some healthy individuals. In particular, a large amount of monoclonal RF (mainly IgM) is observed in the patients with some B-cell malignancies, such as mixed cryoglobulinemia (MC) and Waldenströms macroglobulinemia (WM) (Randen et al. 1992a). In these diseases, precipitation of a mixture of IgG and monoclonal IgM with RF activity, named cryoglobulin, is observed in the serum of the patient below body temperature. The cryoglobulin deposits on tissues such as small blood vessel and kidneys, causing inflammation and tissue injury (Muchtar et al. 2017). The abundant abnormal immunoglobulins secreted from malignant B cells are called “paraprotein”. Immunochemical studies of RFs, such as those to elucidate their molecular-recognition mechanisms, have evolved from studies using monoclonal IgM paraprotein with RF activity derived from B cell malignancies (Randen et al. 1992a).
There is a difference in genetic diversity and Fc-binding patterns between RA-derived RF and B cell lymphoma-derived RF. RFs derived from B cells isolated from RA patients are more diverse in terms of germline gene types than RFs derived from B cell malignancies (Pascual et al. 1990, 1992; Victor et al. 1991; Ermel et al. 1993; Mantovani et al. 1993; Youngblood et al. 1994). Additionally, RA-derived RFs exhibit various binding patterns for different IgG subclasses (IgG1, IgG2, IgG3, and IgG4) (Artandi et al. 1991, 1992), and its epitopes are thought to be widely distributed in the Fc region (Bonagura et al. 1993). By contrast, in B cell malignancies, such as MC and WM, monoclonal IgM-type RFs with specific idiotype combinations are frequently observed, and the binding pattern of the RFs is limited (Silverman et al. 1988; Kunkel et al. 1973).
2 RFs from a Common VH and VL Gene
In the serum of healthy individuals immunized with foreign antigen (mismatched red blood cells) (Thompson et al. 1994) and in patients who developed MC due to hepatitis C virus (HCV) infection (Charles et al. 2008), mucosal-associated lymphoid tissue-type lymphoma (Bende et al. 2005), HCV-associated lymphoma (De Re et al. 2002), or B cell chronic lymphocytic leukemia (Stamatopoulos et al. 2007), RFs with a combination of heavy and light chains derived from a specific germline gene are frequently observed (Bende et al. 2015). In such cases, the combination of a heavy chain with a cross-reactive idiotype (CRI) called Wa and a light chain with a CRI called 17.109 are the most frequent. Each of these heavy and light chains is derived from IGHV1-69 and IGKV3-20, respectively, which are frequently used V-region genes for RFs. RFs derived from a specific germline gene (especially H chain) having a specific length in the third complementarity determining region of the H chain (CDR-H3), and highly homologous to the RF are specifically referred to as “stereotypic RFs”. IGHV1-69 is the most major germline gene for stereotypic RFs; however, recently, stereotypic RFs derived from germline genes such as IGHV3-7 and IGHV4-59, have also been reported (Hoogeboom et al. 2013; Bende et al. 2016).
The heavy chain germline gene IGHV1-69 is used in the heavy chain of neutralizing antibodies against the following viruses and virulence factors: HCV (Kong et al. 2013; Chan et al. 2001), human immunodeficiency virus (HIV) (Luftig et al. 2006), influenza virus (Ekiert et al. 2009), Middle East respiratory syndrome coronavirus (MERS-CoV) (Ying et al. 2015), and the Staphylococcus aureus virulence factor NEAT (Yeung et al. 2016). The germline gene for the light chain is the kappa chain IGKV3-20, which normally pairs with the heavy chain IGHV1-69. Additionally, IGHV1-69-derived RF is reportedly highly associated with HCV-related lymphoma (Ivanovski et al. 1998), and notably, >80% of patients with MC are carriers of HCV (Agnello et al. 1992). Moreover, IgM antibody derived from IGHV1-69, which occurs in patients with HCV-related MC, reportedly exhibits RF activity with or without somatic mutation (Charles et al. 2013). These findings suggest that the antibody derived from IGHV1-69 harbors characteristics of molecular recognition related to RF activity.
Therefore, although IGHV1-69-derived RF is physiologically and pathologically important, details of the antigen-recognition mechanism have not been clarified. Previous reports suggest that IGHV1-69-derived RFs isolated from immunized healthy individuals, RA patients, or MC patients have a CDR-H3 with a length restriction of 12–15 residues but with diverse amino acid composition (Bende et al. 2005; Charles et al. 2013; Borretzen et al. 1995). On the other hand, most IGHV1-69-derived RFs recognize the CH2-CH3 elbow region of Fc as their epitope (Artandi et al. 1992). The question of whether IGHV1-69-derived RFs recognize this epitope in a common recognition mode, and whether the antibodies from this germline gene have structural features likely to exhibit RF activity, are interesting from a structural biology perspective.
3 Structure of the IGHV1-69-Derived Antibodies
As mentioned above, the IGHV1-69 gene-encoded antibodies are frequently observed as neutralizing antibodies against viruses and bacterial pathogens. Understanding the recognition mechanism of neutralizing antibodies provide important insights into rational vaccine design. So, what are the structural features of molecular recognition of this germline antibodies? This section summarizes the structure of IGHV1-69 antibodies other than RF. A systematic analysis of antigen-free Fab structures reveals that the structures of CDR-H1 of IGHV1-69 antibodies are substantially flexible, whereas the canonical structures of CDR-H2 have consistent conformations (Teplyakov et al. 2016). This study also reveals that heavy chains derived from IGHV1-69 gene form thermally stable Fab fragments.
What is common to neutralizing antibodies is that the germline sequence of IGHV1-69 originally has the properties of a neutralizing antibody, and becomes a neutralizing antibody without introducing many mutations. The structure of D5, a broadly neutralizing antibody for HIV-1, in complex with the glycoprotein gp41 shows that the hydrophobic CDR-H2 protrudes into the conserved gp41 hydrophobic pocket, which determines neutralizing ability (Luftig et al. 2006). The structure of m336, a neutralizing antibody for the MERS-CoV receptor-binding domain, reveals that the heavy chain is a major contributor to antigen binding, and that the V(D)J junctional and IGHV1-69 allele-specific residues play an important role in the neutralization of MERS-CoV (Ying et al. 2015). The structure of the neutralizing antibodies against Staphylococcus aureus virulence factor NEAT2 also reveals that the interaction by the germline-encoded CDR-H2 is critical for neutralization (Yeung et al. 2016). HCV is closely related to IGHV1-69 RFs. The structure of AR3C, a broadly neutralizing antibody for HCV, in complex with the envelope protein E2 reveals that the heavy chain is responsible for 86% of the antigen-antibody binding (Kong et al. 2013). The CDR-H2 of AR3C interacts with the hydrophobic cluster on E2 important for binding to the cell-surface CD81. The structures of the four independent neutralizing antibodies against HCV suggest the mechanism of the broad neutralization of the IGHV1-69-derived antibodies (Tzarum et al. 2019).
Among the IGHV1-69-derived neutralization antibody, the antibodies for influenzae virus are most reported. The structure of F10 and CR6261, broadly neutralizing antibodies against H5 hemagglutinin (HA), reveals that these antibodies block infection by inserting its heavy chain, which is the only contributor to the interaction, into a conserved pocket in the stem region (Ekiert et al. 2009; Sui et al. 2009). The germline-encoded CDR-H1 and CDR-H2 of the antibody CR6261 interact with the highly conserved stem region in HAs, demonstrating the broad neutralization and high affinity (Ekiert et al. 2009). Furthermore, only a small number of somatic mutations on germline residues enables strong neutralization, suggesting that selected immunoglobulin genes such as IGHV1-69 are adopted for neutralizing antibody (Lingwood et al. 2012). The IGHV1-69-derived neutralizing antibodies, 8M2 and 2G1, recognize the receptor binding region of H2 HA. The structure of 8M2 and 2G1 reveals that the Phe54 in CDR-H2 plugs a conserved cavity in the HA receptor binding site (Xu et al. 2013).
4 Structure of Determined RFs
The following sections focus on the molecular-recognition mechanisms of the RF derived from the germline gene IGHV1-69 based on the crystal structure of RF YES8c previously described (Fig. 1a) (Shiroishi et al. 2018). Before reporting the YES8c structure, the crystal structures of the two RFs had been revealed in complex with Fc (Corper et al. 1997; Duquerroy et al. 2007), providing interesting insights into the molecular-recognition mechanism and the origin of RFs. One is an IgM-type RF named RF-AN derived from an RA patient and that recognizes the CH2-CH3 cleft region of Fc, a major epitope recognized by many RFs (Fig. 1b). Unlike general antibodies, this RF has the unusual feature of binding Fc at the edge of the conventional paratope (antigen-recognition site) (Corper et al. 1997). The authors considered that due to its unique binding mode, an antibody originally against another antigen acquired binding ability to IgG-Fc at sites other than the original antigen-binding site by somatic mutation (Sutton et al. 2000). The other is a high-affinity IgM-type RF called RF61, which does not recognize the CH2-CH3 cleft but recognizes the CH3 domain near the C-terminus of Fc across two CH3 domains (Fig. 1b) (Duquerroy et al. 2007). The crystal structure of RF61 revealed a new RF epitope, as well as highlighted the diversity of RF epitopes. However, these two RFs were not stereotypic RFs and did not answer the questions presented in this review.
Crystal structure of YES8c in complex with IgG1-Fc and other Fc-binding proteins (a) Overall structure of the YES8c-Fab–Fc complex in an asymmetric unit. Fc and heavy and light chains are shown in pink, cyan, and green, respectively. In complex 2, atomic models of the constant region of YES8c could not be built due to weak electron density. (b) Overall structure of the RF-AN–Fc complex (PDB ID: 1ADQ) and the RF61–Fc complex (PDB ID: 2J6E). (c) Overall structure of other Fc-binding proteins in complex with Fc: minimized B domain of S. aureus protein A (PDB ID: 1L6X), Streptococcal protein G (PDB ID: 1FCC), neonatal Fc receptor (FcRn) (PDB ID: 1I1A), and TRIM21 (PDB ID: 2IWG)
5 Structural Basis for Molecular Recognition of the RF YES8c
5.1 RF YES8c
YES8c is a mono-reactive and IgM-type RF isolated from bone marrow B cells from RA patients and does not exhibit cross-reactivity with autoantigens other than IgG-Fc (Ezaki et al. 1991). This particular patient also had macroglobulinemia due to macroglobulin (IgM) produced by B cell lymphoma. The germline genes in the V, D, and J regions that constitute the heavy chain variable region of YES8c are IGHV1-69, IGHD2-2, and IGHJ4, respectively, and the germline genes in the V and J regions of the light chain are IGKV3-20 and IGKJ2, respectively. Therefore, YES8c is a stereotype RF derived from the IGHV1-69/IGKV3-20 germline gene combination common in B cell lymphomas.
5.2 Overall Structure of the YES8c−Fc Complex
We elucidated the crystal structure of the complex of the YES8c Fab fragment and human IgG1-Fc with a resolution of 2.8 Å (Shiroishi et al. 2018). Two YES8c−IgG1-Fc complexes were observed in the asymmetric unit of the crystal, with these two complexes nearly identical structures, except for the constant region (Fig. 1a), hence complex 1 is discussed below. YES8c bound to the CH2-CH3 elbow region of Fc similar to RF-AN (Fig. 1b) (Corper et al. 1997). Additionally, the crystal structure revealed that several proteins, such as protein A, protein G, FcRn, and tripartite motif-containing protein 21 (TRIM21) recognize the CH2-CH3 elbow (Fig. 1c) (James et al. 2007; Martin et al. 2001; Sauer-Eriksson et al. 1995; Deisenhofer 1981). Therefore, this region is thought to have an intrinsic physiochemical property that is easily recognized by other proteins (DeLano et al. 2000). Because the complex structure of protein A and Fc was elucidated long ago, protein A has been used to characterize the interaction of many RFs by the competitive-inhibition assay for Fc (Deisenhofer 1981; Sasso et al. 1988). The heavy chain of YES8c recognizes mainly the CH2−CH3 cleft, and its binding site overlaps with the site where protein A binds. Therefore, YES8c interaction was expected to be inhibited by protein A. Most regions of the light chain of YES8c were utilized for interaction with the CH3 domain.
One characteristic of the YES8c−Fc complex is the presence of a wider interaction interface than that of RF-AN and RF61 (Fig. 1b). The buried surface area involved in forming the YES8c−Fc complex is 1996–2101 Å2, which is larger than RF-AN (1458 Å2) and RF61 (1689 Å2). Additionally, the YES8c heavy chain has a larger interaction interface involved with the Fc interaction than the light chain, and the shape complementarity is also high, suggesting that the heavy chain plays a more central role in Fc recognition. Included in the large interaction interface are 22 amino acid residues on the YES8c side involved in Fc binding, which is a higher number of involved residues than RF-AN (9) and RF61 (14).
Here, we discuss structural differences with RF-AN, an RF that recognizes the CH2-CH3 elbow, as well as YES8c. The binding mode of RF-AN and YES8c differ, as the RF-AN antigen-recognition site deviates from the pseudo two-fold axis formed between VH and VL, and this RF also has the unusual feature of recognizing Fc at the edge of the general paratope (Fig. 2) (Corper et al. 1997). On the other hand, in YES8c, Fc is bound near the center of the pseudo two-fold axis of VH and VL, which is the normal binding mode of antibody (Fig. 2). Additionally, the binding mode of CDR-H3 located at the center of the antigen-binding site is also greatly different between RF-AN and YES8c. In RF-AN, the interaction area of CDR-H3 occupies 44% of the entire antibody binding site, and side chain residues of CDR-H3 (H-Arg96, H-Tyr98, and H-Val99), which is the key to the interaction, form a salt bridge, hydrogen bonds, and hydrophobic interactions with Fc, thereby greatly contributing to antigen recognition. This suggests that CDR-H3 plays a central role in the interaction between RF-AN and Fc. By contrast, the YES8c CDR-H3 occupies only 21% of the interaction area, and the interaction formed between CDR-H3 and Fc is not as strong, suggesting that regions other than YES8c CDR-H3 play a central role in Fc binding.
Comparison of the YES8c and RF-AN binding modes In YES8c, Fc and heavy and light chains are shown in magenta, cyan, and green, respectively, and in light pink, light blue, and light green, respectively, in RF-AN. CDRs are shown in orange in the both structures. The pseudo two-fold axis between VH and VL domains is shown as a dashed line
5.3 YES8c Subclass Specificity and Affinity
The binding specificity of RFs to the IgG subclass is largely divided into two groups: Ga (binding to IgG1, 2, and 4) and Pan (binding to all subclasses) (Bonagura et al. 1993). Binding experiments show that YES8c binds to both IgG1 and IgG4 subclasses (Shiroishi et al. 2018); therefore, YES8c belongs to a group with subclass specificity different from Ga and Pan. Here we discuss the binding specificity observed in YES8c according to the three-dimensional structure. Comparison of residues of the IgG3 epitope with other IgG subclasses reveals amino acid residue differences at positions 384, 422, 435, and 436. In particular, position 435 in Fc, which is the key to Ga specificity (Artandi et al. 1992), is a histidine in IgG1, IgG2, and IgG4 but an arginine in IgG3. According to a previous report, although RF-AN exhibits Ga specificity, the reason for its weak affinity with IgG3 cannot be clarified from the RF-AN crystal structure (Corper et al. 1997). The YES8c-Fc complex structure shows that arginine at position 435 would collide with H-Phe54 of YES8c, thereby generating subclass specificity. However, the reason for the reduced affinity with IgG2 remains unknown. Given the possibility that the three-dimensional structure of Fc is slightly different for each IgG subclass due to the influence of amino acid residues other than those at the interaction site, it is necessary to clarify the detailed structure of the subclass.
Similar to RF-AN, YES8c displays a low affinity for Fc. The affinity of YES8c Fab for IgG1 determined by surface plasmon resonance analysis is 160 ± 30 μM (Kd) (Shiroishi et al. 2018). Although this represents low affinity, these RFs are present in the form of IgM with 10 Fab “arms”, suggesting that they exhibit strong binding affinity to antigens along with high avidity effect. As noted, the YES8c−Fc complex has a large interface area, with many residues involved in the interaction; however, the distance of the hydrogen bonds between YES8c and Fc are relatively long, implying low bond strength. Moreover, the complex included no salt bridge, which is an interaction that generally promotes affinity, although the number of van der Waals interactions was similar to that in other RF (RF-AN and RF61) complexes. Therefore, the structure suggests that YES8c is loosely bound to Fc, which might also explain its low affinity for IgG1-Fc, despite the large interaction interface. Such a “loose” binding mode might be an important feature for IGHV1-69-derived RF recognition of common epitopes in Fc, regardless of the diversity of the CDR-H3 or the introduction of somatic hyper mutations (SMHs).
5.4 Length Limit and Amino Acid Diversity of the CDR-H3 in Fc Recognition
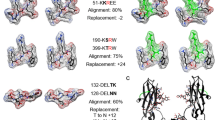
As previously reported, in IGHV1-69-derived RFs, the length of the CDR-H3 is limited to 12–15 residues, although some variation in amino acid composition is observed (Fig. 3a) (Bende et al. 2005, 2015; Borretzen et al. 1995). Based on the crystal structure of the YES8c–Fc complex, we proposed a structural model for allowing amino acid diversity in the CDR-H3 loop associated with Fc recognition by IGHV1-69-derived RFs (Shiroishi et al. 2018). In the structure of the YES8c–Fc complex the Leu432-His435 region protruding at the Fc elbow penetrates into the pocket formed between the RF VL and VH regions, and Asn434 interacts with the CDR-H3 from the inside of the loop (Fig. 3b), representing a characteristic feature of YES8c recognition. In particular, the area where the Leu432–His435 region interacts with Fc (~ 320 Å2) is large, occupying one third of the total interaction area. The Asn434 side chain is located near the center of the antigen-binding site and interacts with the CDR-H3 loop to primarily forms hydrogen bonds with CDR-H3 main-chain atoms (the N of H-Gly100 and the O of H-Thr100A). The CDR-H3 residues (especially from H-Ala99 to H-Pro100B) are subsequently located in the space formed between CH2-CH3 and without insertion into the binding interface (Fig. 3c). This binding mode allows accommodation of CDR-H3 in the space formed between CH2 and CH3 by only slight changes to the loop structure, regardless of amino acid composition. If the loop length is <12 residues, this will likely preclude formation of numerous interactions between CDR-H3 and Fc. On the other hand, a length > 15 residues would prevent CDR-H3 from fitting in the space between CH2 and CH3 or result in CDR-H3 covering the pocket between VL and VH, thereby interfering with interactions with the Leu432–His435 region and preventing RF activity.
Structural features of Fc recognition by YES8c (a) Alignment of the amino acid residues in the CDR-H3 of YES8c and the other IGHV1–69/IGKV3–20-derived RFs. The residues are numbered based on the Kabat numbering scheme. (b) The protruding region in Leu432-His435 region of Fc bound to YES8c. The heavy and light chains of YES8c are shown as cyan and green surfaces, respectively, and Fc is shown as magenta sticks. The CDR-H3 is shown in orange. (c) The interaction between the CDR-H3 and the CH2–CH3 cleft of Fc. The surfaces of CH2 and CH3 are shown in light pink and magenta, respectively. VH (excluding the CDR-H3) and VL domains are shown as cyan and green surfaces, respectively. The CDR-H3 is shown as orange sticks. (d) Alignment of the amino acid residues of CDR-H2 in YES8c and the IGHV1–69/IGKV3–20-derived RFs. (e) Interactions between CDR-H2 of YES8c and Fc. The Fc surface is shown with the hydrophobicity scale (Eisenberg et al. 1984). (f) Interaction footprints in the light chain of YES8c with Fc (left), and the 90° rotated view (right). The contact area of the light chain with Fc is shown in green. The other surface of the light chain is shown in pale yellow. Fc is shown as a magenta cartoon model
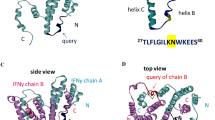
5.5 The Hydrophobic Region in CDR-H2 Common to IGHV1-69-Derived Antibodies
The neutralizing antibodies anti-influenza hemagglutinin antibody CR6261 (Lingwood et al. 2012), the anti-HCV E2 envelope antibody AR3C (Kong et al. 2013), and the anti-HIV-1 gp41 antibody D5A (Luftig et al. 2006) are all derived from the IGHV1-69 germline gene. Crystal structures of these antibody−antigen complexes revealed that the hydrophobic residues in the CDR-H2 commonly recognize the hydrophobic patch of the antigen. Therefore, the naive antibody derived from IGHV1-69 is considered to be a “pattern-recognizing” antibody capable of hydrophobic interactions with CDR-H2 residues. CDR-H2 hydrophobicity is highly conserved in IGHV1-69-derived RFs (Fig. 3d). In YES8c, the hydrophobic residues (H-Leu53 and H-Phe54) at the tip of the CDR-H2 loop are located at the CH2-CH3 cleft and form hydrophobic interactions with Ile253 and Leu314 across a large interfacial area (Fig. 3e). Because the H-L53A and H-F54A mutants show significantly decreased affinity, recognition by H-Leu53 and H-Phe54 might significantly contribute to RF activity (Shiroishi et al. 2018).
5.6 The Importance of the Light Chain Paired with IGHV1-69 Heavy Chain
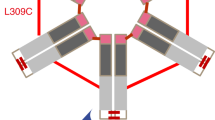
In IGHV1-69-derived RFs, the paired light chain is derived from IGKV3-20 in most cases, and the amino acid sequence of the light chain is highly conserved throughout. In the YES8c−Fc complex, 10–11 residues of the light chain are involved in Fc interaction, and the interfacial area between the light chain and Fc is 430–450 Å2, which is considerably larger than the light chains of RF-AN (2 residues and 196 Å2) and RF61 (5 residues and 255 Å2). CDR-L1 and CDR-L3 in YES8c form a flat binding surface and recognize the flat β-sheet region of the CH3 domain (Fig. 3f). Although the area of the interaction interface between the light chain and Fc is large, the calculated shape complementarity between the light chain and Fc is low, indicating that the light chain is loosely bound with the flat binding interface. Surprisingly, however, substitution of various residues in the binding site in the light-chain region with an alanine (L-S29A, L-S31A, L-Y32A, and L-S94A) resulted in a significant decrease in Fc binding, suggesting that highly conserved light-chain residues might contribute important interactions with Fc via hydrophilic residues (Shiroishi et al. 2018).
5.7 Effects of SHMs in RFs
Many identified RFs harbor SHMs; however, some antibodies exhibit RF activity in the absence of SHMs (Charles et al. 2013; Randen et al. 1992b; Carayannopoulos et al. 2000). A naive antibody of RF-TS1, an RF derived from IGHV1-69, displays higher affinity for Fc than RF-TS1 (Carayannopoulos et al. 2000), whereas in mixed macroglobulinemia related to HCV, IgM derived from IGHV1-69 reportedly acquires RF activity through SHMs (Charles et al. 2013). However, the degree to which SHMs affect the acquisition of RF activity and by what mechanism it imparts higher RF activity remain unknown. The structure of RF-AN and RF61 suggested that SHMs might be important for RF activity, but there previously had been no mutants created or analyzed to investigate the effects of SHMs. Therefore, we generated mutants reverting to the germline amino acid residues and analyzed their interactions with Fc (Shiroishi et al. 2018). Three SHMs (L-V28I, L-K39R, and L-S53T) exist in the VL domain, and seven SHMs (H-A33P, H-M48V, H-I53L, H-K62R, H-S76R, H-L82V, H-S82BV, and H-D100EF) exist in the VH domain. Among these, we selected three sites (L28, H33, and H53) close to the Fc-binding site and prepared mutants (L-I28V, H-P33A, and H-L53I) by reverting to the respective germline amino acid. Because the affinity of the H-P33A mutant for Fc was greatly reduced, SHM at this site might be important for acquisition of RF activity. On the other hand, the affinity for Fc was increased in the H-L53I mutant, whereas that in the L-I28V mutant was unchanged. These findings suggest that during the process of B cell proliferation, mutations are introduced that result in a spectrum of contributions to Fc affinity, thereby dictating changes in RF activity. In the case of YES8c, it is considered that higher RF activity is gradually acquired as a result of SHMs.
Poly-reactive RFs with low affinity are found in healthy individuals as naturally occurring IgM. On the other hand, mono-reactive RFs with high affinity are observed in various autoimmune diseases, implying that increased affinity might be related to disease progression. Based on the structure of the YES8c−Fc complex, we designed mutants potentially capable of forming interactions resulting in improved affinity. Our analyses using Fc showed Kd values for H-T56K, H-N58K, L-Q27E, and L-S27AN of 42 μM, 94 μM, 78 μM, and 69 μM, respectively, revealing increased affinities relative to wild type (160 μM). Similar to results observed with the H-L53I mutant, these are amino acid mutations derived from a single-base substitution, suggesting that the antibody derived from IGHV1-69 can easily increase its affinity via various SHMs to become a disease-aggravation factor.
6 Concluding Remarks
RFs derived from the germline gene combination of IGHV1-69 have been studied for many years as a paraprotein IgM-RF in MC and WM. Additionally, IGHV1-69-derived RFs are found at high levels as stereotype RFs in patients with B cell lymphoma caused by HCV infection (Charles et al. 2008; Bende et al. 2005; De Re et al. 2002). The crystal structure and mutant analyses of YES8c revealed the following possible antigen-recognition mechanisms common to IGHV1-69-derived stereotypic RFs: (1) interaction with the Leu432−His435 region of Fc allows orientation of the CDR-H3 within the space formed by the CH2-CH3 elbow, which allows amino acid diversity but limits the number of residues in the CDR-H3; (2) hydrophobic residues in the CDR-H2 typical of IGHV1-69-derived antibodies interact with a hydrophobic region in Fc; and (3) paired light chains are highly conserved and form a loose but significant interaction with Fc. Confirmation of these models require clarification of the complex structures of other IGHV1-69-derived RFs in complex with Fc. Furthermore, the structures of other germline-derived RFs will allow the elucidation of the whole picture of autoreactivity of RFs.
References
Agnello V, Chung RT, Kaplan LM (1992) A role for hepatitis C virus infection in type II cryoglobulinemia. N Engl J Med 327(21):1490–1495. https://doi.org/10.1056/NEJM199211193272104
Artandi SE, Canfield SM, Tao MH, Calame KL, Morrison SL, Bonagura VR (1991) Molecular analysis of IgM rheumatoid factor binding to chimeric IgG. J Immunol 146(2):603–610
Artandi SE, Calame KL, Morrison SL, Bonagura VR (1992) Monoclonal IgM rheumatoid factors bind IgG at a discontinuous epitope comprised of amino acid loops from heavy-chain constant-region domains 2 and 3. Proc Natl Acad Sci U S A 89(1):94–98
Bende RJ, Aarts WM, Riedl RG, de Jong D, Pals ST, van Noesel CJ (2005) Among B cell non-Hodgkin’s lymphomas, MALT lymphomas express a unique antibody repertoire with frequent rheumatoid factor reactivity. J Exp Med 201(8):1229–1241. https://doi.org/10.1084/jem.20050068
Bende RJ, Slot LM, Hoogeboom R, Wormhoudt TA, Adeoye AO, Guikema JE, van Noesel CJ (2015) Stereotypic rheumatoid factors that are frequently expressed in mucosa-associated lymphoid tissue-type lymphomas are rare in the labial salivary glands of patients with Sjogren’s syndrome. Arthritis Rheumatol 67(4):1074–1083. https://doi.org/10.1002/art.39002
Bende RJ, Janssen J, Wormhoudt TA, Wagner K, Guikema JE, van Noesel CJ (2016) Identification of a novel stereotypic IGHV4-59/IGHJ5-encoded B-cell receptor subset expressed by various B-cell lymphomas with high affinity rheumatoid factor activity. Haematologica 101(5):e200–e203. https://doi.org/10.3324/haematol.2015.139626
Bonagura VR, Artandi SE, Davidson A, Randen I, Agostino N, Thompson K, Natvig JB, Morrison SL (1993) Mapping studies reveal unique epitopes on IgG recognized by rheumatoid arthritis-derived monoclonal rheumatoid factors. J Immunol 151(7):3840–3852
Borretzen M, Randen I, Natvig JB, Thompson KM (1995) Structural restriction in the heavy chain CDR3 of human rheumatoid factors. J Immunol 155(7):3630–3637
Carayannopoulos MO, Potter KN, Li Y, Natvig JB, Capra JD (2000) Evidence that human immunoglobulin M rheumatoid factors can Be derived from the natural autoantibody pool and undergo an antigen driven immune response in which somatically mutated rheumatoid factors have lower affinities for immunoglobulin G Fc than their germline counterparts. Scand J Immunol 51(4):327–336
Chan CH, Hadlock KG, Foung SK, Levy S (2001) V(H)1-69 gene is preferentially used by hepatitis C virus-associated B cell lymphomas and by normal B cells responding to the E2 viral antigen. Blood 97(4):1023–1026
Charles ED, Green RM, Marukian S, Talal AH, Lake-Bakaar GV, Jacobson IM, Rice CM, Dustin LB (2008) Clonal expansion of immunoglobulin M+CD27+ B cells in HCV-associated mixed cryoglobulinemia. Blood 111(3):1344–1356. https://doi.org/10.1182/blood-2007-07-101717
Charles ED, Orloff MI, Nishiuchi E, Marukian S, Rice CM, Dustin LB (2013) Somatic hypermutations confer rheumatoid factor activity in hepatitis C virus-associated mixed cryoglobulinemia. Arthritis Rheum 65(9):2430–2440. https://doi.org/10.1002/art.38041
Corper AL, Sohi MK, Bonagura VR, Steinitz M, Jefferis R, Feinstein A, Beale D, Taussig MJ, Sutton BJ (1997) Structure of human IgM rheumatoid factor Fab bound to its autoantigen IgG Fc reveals a novel topology of antibody-antigen interaction. Nat Struct Biol 4(5):374–381
De Re V, De Vita S, Gasparotto D, Marzotto A, Carbone A, Ferraccioli G, Boiocchi M (2002) Salivary gland B cell lymphoproliferative disorders in Sjogren’s syndrome present a restricted use of antigen receptor gene segments similar to those used by hepatitis C virus-associated non-Hodgkins’s lymphomas. Eur J Immunol 32(3):903–910. https://doi.org/10.1002/1521-4141(200203)32:3<903::AID-IMMU903>3.0.CO;2-D
Deisenhofer J (1981) Crystallographic refinement and atomic models of a human Fc fragment and its complex with fragment B of protein A from Staphylococcus aureus at 2.9- and 2.8-A resolution. Biochemistry 20(9):2361–2370
DeLano WL, Ultsch MH, de Vos AM, Wells JA (2000) Convergent solutions to binding at a protein-protein interface. Science 287(5456):1279–1283. https://doi.org/10.1126/science.287.5456.1279
Duquerroy S, Stura EA, Bressanelli S, Fabiane SM, Vaney MC, Beale D, Hamon M, Casali P, Rey FA, Sutton BJ, Taussig MJ (2007) Crystal structure of a human autoimmune complex between IgM rheumatoid factor RF61 and IgG1 Fc reveals a novel epitope and evidence for affinity maturation. J Mol Biol 368(5):1321–1331. https://doi.org/10.1016/j.jmb.2007.02.085
Eisenberg D, Schwarz E, Komaromy M, Wall R (1984) Analysis of membrane and surface protein sequences with the hydrophobic moment plot. J Mol Biol 179(1):125–142
Ekiert DC, Bhabha G, Elsliger MA, Friesen RH, Jongeneelen M, Throsby M, Goudsmit J, Wilson IA (2009) Antibody recognition of a highly conserved influenza virus epitope. Science 324(5924):246–251. https://doi.org/10.1126/science.1171491
Ermel RW, Kenny TP, Chen PP, Robbins DL (1993) Molecular analysis of rheumatoid factors derived from rheumatoid synovium suggests an antigen-driven response in inflamed joints. Arthritis Rheum 36(3):380–388
Ezaki I, Kanda H, Sakai K, Fukui N, Shingu M, Nobunaga M, Watanabe T (1991) Restricted diversity of the variable region nucleotide sequences of the heavy and light chains of a human rheumatoid factor. Arthritis Rheum 34(3):343–350
Hoogeboom R, Wormhoudt TA, Schipperus MR, Langerak AW, Dunn-Walters DK, Guikema JE, Bende RJ, van Noesel CJ (2013) A novel chronic lymphocytic leukemia subset expressing mutated IGHV3-7-encoded rheumatoid factor B-cell receptors that are functionally proficient. Leukemia 27(3):738–740. https://doi.org/10.1038/leu.2012.238
Ivanovski M, Silvestri F, Pozzato G, Anand S, Mazzaro C, Burrone OR, Efremov DG (1998) Somatic hypermutation, clonal diversity, and preferential expression of the VH 51p1/VL kv325 immunoglobulin gene combination in hepatitis C virus-associated immunocytomas. Blood 91(7):2433–2442
James LC, Keeble AH, Khan Z, Rhodes DA, Trowsdale J (2007) Structural basis for PRYSPRY-mediated tripartite motif (TRIM) protein function. Proc Natl Acad Sci U S A 104(15):6200–6205. https://doi.org/10.1073/pnas.0609174104
Kong L, Giang E, Nieusma T, Kadam RU, Cogburn KE, Hua Y, Dai X, Stanfield RL, Burton DR, Ward AB, Wilson IA, Law M (2013) Hepatitis C virus E2 envelope glycoprotein core structure. Science 342(6162):1090–1094. https://doi.org/10.1126/science.1243876
Kunkel HG, Agnello V, Joslin FG, Winchester RJ, Capra JD (1973) Cross-idiotypic specificity among monoclonal IgM proteins with anti- -globulin activity. J Exp Med 137(2):331–342. https://doi.org/10.1084/jem.137.2.331
Lingwood D, McTamney PM, Yassine HM, Whittle JR, Guo X, Boyington JC, Wei CJ, Nabel GJ (2012) Structural and genetic basis for development of broadly neutralizing influenza antibodies. Nature 489(7417):566–570. https://doi.org/10.1038/nature11371
Luftig MA, Mattu M, Di Giovine P, Geleziunas R, Hrin R, Barbato G, Bianchi E, Miller MD, Pessi A, Carfi A (2006) Structural basis for HIV-1 neutralization by a gp41 fusion intermediate-directed antibody. Nat Struct Mol Biol 13(8):740–747. https://doi.org/10.1038/nsmb1127
Mantovani L, Wilder RL, Casali P (1993) Human rheumatoid B-1a (CD5+ B) cells make somatically hypermutated high affinity IgM rheumatoid factors. J Immunol 151(1):473–488
Martin WL, West AP Jr, Gan L, Bjorkman PJ (2001) Crystal structure at 2.8 A of an FcRn/heterodimeric Fc complex: mechanism of pH-dependent binding. Mol Cell 7(4):867–877
Muchtar E, Magen H, Gertz MA (2017) How I treat cryoglobulinemia. Blood 129(3):289–298. https://doi.org/10.1182/blood-2016-09-719773
Pascual V, Randen I, Thompson K, Sioud M, Forre O, Natvig J, Capra JD (1990) The complete nucleotide sequences of the heavy chain variable regions of six monospecific rheumatoid factors derived from Epstein-Barr virus-transformed B cells isolated from the synovial tissue of patients with rheumatoid arthritis. Further evidence that some autoantibodies are unmutated copies of germ line genes. J Clin Invest 86(4):1320–1328. https://doi.org/10.1172/JCI114841
Pascual V, Victor K, Randen I, Thompson K, Steinitz M, Forre O, Fu SM, Natvig JB, Capra JD (1992) Nucleotide sequence analysis of rheumatoid factors and polyreactive antibodies derived from patients with rheumatoid arthritis reveals diverse use of VH and VL gene segments and extensive variability in CDR-3. Scand J Immunol 36(2):349–362
Randen I, Thompson KM, Pascual V, Victor K, Beale D, Coadwell J, Forre O, Capra JD, Natvig JB (1992a) Rheumatoid factor V genes from patients with rheumatoid arthritis are diverse and show evidence of an antigen-driven response. Immunol Rev 128:49–71
Randen I, Brown D, Thompson KM, Hughes-Jones N, Pascual V, Victor K, Capra JD, Forre O, Natvig JB (1992b) Clonally related IgM rheumatoid factors undergo affinity maturation in the rheumatoid synovial tissue. J Immunol 148(10):3296–3301
Sasso EH, Barber CV, Nardella FA, Yount WJ, Mannik M (1988) Antigenic specificities of human monoclonal and polyclonal IgM rheumatoid factors. The C gamma 2-C gamma 3 interface region contains the major determinants. J Immunol 140(9):3098–3107
Sauer-Eriksson AE, Kleywegt GJ, Uhlen M, Jones TA (1995) Crystal structure of the C2 fragment of streptococcal protein G in complex with the Fc domain of human IgG. Structure 3(3):265–278
Shiroishi M, Ito Y, Shimokawa K, Lee JM, Kusakabe T, Ueda T (2018) Structure-function analyses of a stereotypic rheumatoid factor unravel the structural basis for germline-encoded antibody autoreactivity. J Biol Chem 293(18):7008–7016. https://doi.org/10.1074/jbc.M117.814475
Silverman GJ, Goldfien RD, Chen P, Mageed RA, Jefferis R, Goni F, Frangione B, Fong S, Carson DA (1988) Idiotypic and subgroup analysis of human monoclonal rheumatoid factors. Implications for structural and genetic basis of autoantibodies in humans. J Clin Invest 82(2):469–475. https://doi.org/10.1172/JCI113620
Stamatopoulos K, Belessi C, Moreno C, Boudjograh M, Guida G, Smilevska T, Belhoul L, Stella S, Stavroyianni N, Crespo M, Hadzidimitriou A, Sutton L, Bosch F, Laoutaris N, Anagnostopoulos A, Montserrat E, Fassas A, Dighiero G, Caligaris-Cappio F, Merle-Beral H, Ghia P, Davi F (2007) Over 20% of patients with chronic lymphocytic leukemia carry stereotyped receptors: pathogenetic implications and clinical correlations. Blood 109(1):259–270. https://doi.org/10.1182/blood-2006-03-012948
Sui J, Hwang WC, Perez S, Wei G, Aird D, Chen LM, Santelli E, Stec B, Cadwell G, Ali M, Wan H, Murakami A, Yammanuru A, Han T, Cox NJ, Bankston LA, Donis RO, Liddington RC, Marasco WA (2009) Structural and functional bases for broad-spectrum neutralization of avian and human influenza A viruses. Nat Struct Mol Biol 16(3):265–273. https://doi.org/10.1038/nsmb.1566
Sutton B, Corper A, Bonagura V, Taussig M (2000) The structure and origin of rheumatoid factors. Immunol Today 21(4):177–183
Teplyakov A, Obmolova G, Malia TJ, Luo J, Muzammil S, Sweet R, Almagro JC, Gilliland GL (2016) Structural diversity in a human antibody germline library. MAbs 8(6):1045–1063. https://doi.org/10.1080/19420862.2016.1190060
Thompson KM, Randen I, Borretzen M, Forre O, Natvig JB (1994) Variable region gene usage of human monoclonal rheumatoid factors derived from healthy donors following immunization. Eur J Immunol 24(8):1771–1778. https://doi.org/10.1002/eji.1830240808
Tzarum N, Giang E, Kong L, He L, Prentoe J, Augestad E, Hua Y, Castillo S, Lauer GM, Bukh J, Zhu J, Wilson IA, Law M (2019) Genetic and structural insights into broad neutralization of hepatitis C virus by human VH1-69 antibodies. Sci Adv 5(1):eaav1882. https://doi.org/10.1126/sciadv.aav1882
van Delft MAM, Huizinga TWJ (2020) An overview of autoantibodies in rheumatoid arthritis. J Autoimmun 102392. https://doi.org/10.1016/j.jaut.2019.102392
Victor KD, Randen I, Thompson K, Forre O, Natvig JB, Fu SM, Capra JD (1991) Rheumatoid factors isolated from patients with autoimmune disorders are derived from germline genes distinct from those encoding the Wa, Po, and Bla cross-reactive idiotypes. J Clin Invest 87(5):1603–1613. https://doi.org/10.1172/JCI115174
Xu R, Krause JC, McBride R, Paulson JC, Crowe JE Jr, Wilson IA (2013) A recurring motif for antibody recognition of the receptor-binding site of influenza hemagglutinin. Nat Struct Mol Biol 20(3):363–370. https://doi.org/10.1038/nsmb.2500
Yeung YA, Foletti D, Deng X, Abdiche Y, Strop P, Glanville J, Pitts S, Lindquist K, Sundar PD, Sirota M, Hasa-Moreno A, Pham A, Melton Witt J, Ni I, Pons J, Shelton D, Rajpal A, Chaparro-Riggers J (2016) Germline-encoded neutralization of a Staphylococcus aureus virulence factor by the human antibody repertoire. Nat Commun 7:13376. https://doi.org/10.1038/ncomms13376
Ying T, Prabakaran P, Du L, Shi W, Feng Y, Wang Y, Wang L, Li W, Jiang S, Dimitrov DS, Zhou T (2015) Junctional and allele-specific residues are critical for MERS-CoV neutralization by an exceptionally potent germline-like antibody. Nat Commun 6:8223. https://doi.org/10.1038/ncomms9223
Youngblood K, Fruchter L, Ding G, Lopez J, Bonagura V, Davidson A (1994) Rheumatoid factors from the peripheral blood of two patients with rheumatoid arthritis are genetically heterogeneous and somatically mutated. J Clin Invest 93(2):852–861. https://doi.org/10.1172/JCI117040
Acknowledgements
This work was supported by JSPS KAKENHI (19 K06514) to M.S. We would like to thank Editage (www.editage.com) for English language editing.
Conflicts of Interest
The author declares no conflicts of interest associated with this manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Shiroishi, M. (2020). Structural Basis of a Conventional Recognition Mode of IGHV1-69 Rheumatoid Factors. In: Atassi, M.Z. (eds) Protein Reviews . Advances in Experimental Medicine and Biology(), vol 21. Springer, Cham. https://doi.org/10.1007/5584_2020_510
Download citation
DOI: https://doi.org/10.1007/5584_2020_510
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-67813-5
Online ISBN: 978-3-030-67814-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)