Abstract
Based on several collections from Italy, a detailed description (including also macrophotographs, microphotographs, and drawings) of the morphological characters of the poorly known Pluteus variabilicolor, originally described from Hungary, is provided. The analysis of the ITS sequences placed this species within the P leoninus clade of sect. Hispidoderma, in spite of the presence of clavate elements in the pileipellis. According to the molecular comparison of the type collections, Pluteus castri, a species recently described on the basis of material collected in Japan and Central Russia, is reduced to a synonym of P. variabilicolor.
Similar content being viewed by others
Introduction
The genus Pluteus, typified by P. cervinus, includes saprobic agaricoid fungi characterized by free lamellae, no universal veil (a partial veil may be present in a few species), pinkish spore-print, inverse hymenophoral trama, and inamyloid, cyanophilic spores (Vellinga & Schreurs 1985, Orton 1986, Singer 1986, Vellinga 1990, Heilmann-Clausen 2012). It belongs to the pluteoid clade as circumscribed by several authors on the basis of molecular data (Moncalvo et al. 2000, 2002, Bodensteiner et al. 2004, Binder et al. 2006, Matheny et al. 2006, Justo et al. 2011a, b, Vizzini et al. 2011). The infrageneric taxonomy is based primarily on the characteristics of the hymenial cystidia and pileipellis. Historically, three sections based on morphological features are widely accepted (Homola 1975, Horak & Heinemann 1978, Orton 1986, Singer 1986, Citérin & Eyssartier 1998, Malysheva et al. 2009, Minnis & Sundberg 2009, 2010): namely sect. Pluteus, with thick-walled (metuloid) pleurocystidia, usually with apical hooks, and pileipellis as a cutis; sect. Hispidoderma with non-metuloid pleurocystidia, and a pileipellis consisting of elongated, filamentous elements organized either as a cutis, a hymeniderm or a trichoderm; and sect. Celluloderma with non-metuloid pleurocystidia and a pileipellis composed of swollen clavate or spheropedunculate elements organized in a hymeniderm, with transitions to an epithelium. Singer (1956, 1958, 1986) sometimes also recognized two subsections in section Celluloderma: Mixtini and Eucellulodermini, differing respectively in the presence or absence of elongated cystidiumlike elements in the pileipellis. Finally, Vellinga & Schreurs (1985) proposed an infrageneric classification for Pluteus differing from Singer’s classificatory scheme only with respect to the placement of species with non-metuloid cystidia and a filamentous pileipellis: they reduced sect. Hispidoderma to a subsection of Celluloderma, including only species with a hymenodermal or trichodermal pileipellis, and established sect. Villosi for species with a pileipellis which was a differentiated cutis. This system has not been widely accepted by later authors (e.g. Banerjee 1992, Banerjee & Sundberg 1993, Citérin & Eyssartier 1998, Rodríguez & Guzmán-Dávalos 2001).
Justo et al. (2011a, b) showed a general agreement between the morphological subdivision of Pluteus and the molecular phylogeny they obtained, even though: (1) some species with non-metuloid pleurocystidia and a pileipellis which was a differentiated cutis (formerly sect. Villosi) were placed either in sect. Celluloderma, together with the species characterized by a hymenidermal pileipellis, or in sect. Pluteus with metuloid bearing species; and (2) the division of sect. Celluloderma into subsections Mixtini and Eucellulodermini was not supported by the phylogenetic analyses of the molecular data; species with a mixture of vesiculose-clavate and elongate elements in the pileipellis were not monophyletic and were distributed over sections Celluloderma and Hispidoderma.
Here we fully describe, based on recent Italian collections, the rarely reported Pluteus variabilicolor, which was originally assigned to subsect. Mixtini of sect. Celluloderma by Babos (1978). Aphylogenetic analysis of ITS sequences was performed to: (1) provide molecular insights on its status and taxonomic placement within Pluteus; and (2) assess whether this species, with a mixture of elongated and short elements in the pileipellis, was phylogenetically closer to species in sect. Hispidoderma or in sect. Celluloderma.
The ITS sequence of the isotype collection was also obtained to confirm our identifications of the recent collections.
Material and Methods
Morphology
Microscopic descriptions are based on fresh and dried specimens. The optical microscope used was a Nikon Eclipse E400 trinocular with plan-achromatic objectives. Micrographs were taken with a Moticam 580 camera. The primary mounting media used were: distilled water for the observation of the pigments, and ammoniacal Congo red for the other structures. Spore measurements were based on 90 elements in ammoniacal Congo red and randomly selected from three collections. The following abbreviations are used: I = number of lamellulae between each pair of entire lamellae; and Q = the length divided by width of the spores in side view, Qm being the average quotient. Descriptive terms used in the descriptions follow Vellinga (1988), and collection acronyms follow Index Herbariorum except that “TL” and “VM” refer to the personal reference collections of Tomaso Lezzi and Vincenzo Migliozzi.
DNA extraction, amplification and sequencing
Genomic DNA was isolated from 1 mg per specimen of four dried specimens (TM20130521-01, VM20111105-01, VM20130504-01 and BP-FN 56936) using the DNeasy Plant Mini Kit (Qiagen, Milan, Italy). Universal primers ITS1F/ITS4 were used for the ITS region amplification (White et al. 1990, Gardes & Bruns 1993). Amplification reactions were performed in a PE9700 thermal cycler (Perkin-Elmer, Applied Biosystems) in a 25 µL reaction mixture using the following final concentrations or total amounts: 10 ng DNA, 1×PCR buffer (20 mM Tris/HCl pH 8.4, 50 mM KCl), 0.4 µM ofeach primer, 2.5 mM MgCl2, 0.25 mM of each dNTP, 0.5 units of Taq polymerase (Promega). The PCR program was as follows: 3 min at 95 °C for one cycle; 30 s at 94 °C, 45 s at 50 °C, 2 min at 72 °C for 35 cycles, and 10 min at 72 °C for one cycle. PCR products were resolved ona1.0% agarose gel and visualized after staining with ethidium bromide. The PCR products were purified with the AMPure XP kit (Beckman) and sequenced by MACROGEN (Seoul, Republic of Korea). Sequence assembly and editing were performed using Geneious v. 5.3 (Drummond et al. 2010). The sequences are deposited in GenBank (www.ncbi.nlm.nih.gov/) under the accession numbers given in Fig. 1.
Sequence alignment and phylogenetic analysis
The sequences obtained in this study were combined with published Pluteus ITS rDNA sequences selected from GenBank and UNITE (http://unite.ut.ee) databases on the basis of the greatest similarity based on BLASTsearch, outcomes of recent phylogenetic studies focused on Pluteus (Justo et al. 2011a, b, Pradeep et al. 2012), and subsequent phylogenetic analysis (preliminary trees not shown). Alignments were generated using MAFFT (Katoh et al. 2002) with default conditions for gap openings and gap extension penalties. The sequence alignments were then imported into MEGA v. 5.10 (Tamura et al. 2011) for manual adjustment. Pluteus diettrichii (HM562143) and Pluteus seticeps (HM562199) were used as outgroup taxa following Justo et al. (2011a) and Pradeep et al. (2012). Best-fit models were estimated by both the Akaike information criterion (AIC) and the Bayesian information criterion (BIC) with jModelTest v. 0.1.1 (Posada 2008) to provide a substitution model for the alignment. Phylogenetic analyses were performed using the Bayesian Inference (BI) and Maximum likelihood (ML) approaches. The BI was performed with MrBayes v. 3.1.2 (Huelsenbeck & Ronquist 2001) with four incrementally heated simultaneous Monte Carlo Markov Chains (MCMC) run over 10 million generations, under the GTR+Γ evolutionary model. Trees were sampled every 1 000 generations resulting in an overall sampling of 10 001 trees; the first 2 500 trees were discarded as “burn-in” (25%). Forthe remaining trees, a majority rule consensus tree showing all compatible partitions was computed to obtain estimates for Bayesian Posterior Probabilities (BPP). ML estimation was performed through RAxML v. 7.3.2 (Stamatakis 2006) with 1 000 bootstrap replicates (Felsenstein 1985) using the GTRGAMMAalgorithm to perform a tree inference and search for a good topology. Support values from bootstrapping runs (MLB) were mapped on the globally best tree using the “-f a” option of RAxML and “-x 12345” as a random seed to invoke the novel rapid bootstrapping algorithm. Only BPP values over 0.70 and MLB over 50% are reported in the resulting tree (Fig. 1). Branch lengths were estimated as mean values over the sampled trees. Clade names follow Justo et al. (2011a) and Pradeep et al. (2012); pairwise % identity values of ITS sequences were calculated using MEGA 5.10 (Tamura et al. 2011).
Results
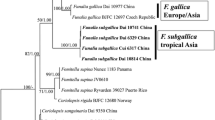
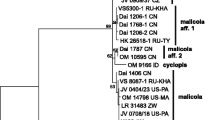
Both Bayesian and Maximum likelihood analyses produced the same topology; therefore, only the Bayesian tree with both BPP and MLB values is shown (Fig. 1). The ITS data matrix comprised 46 sequences (including 38 from GenBank and 4 from UNITE). The alignment comprised 715 characters, and contains 312 variable sites. The three Italian collections of P. variabilicolor and the isotype collection cluster with the three collections of P. castri (holotype included) forming a well-supported clade (/variabilicolor; BPP = 1, MLB = 100%) within the Pluteus leoninus clade of sect. Hispidoderma. The pairwise % identity value among the sequences of the /variabilicolor is 98.8. The clade is in sister position to /chrysaegis (BPP = 1, MLB = 100%). The clade consisting of /variabilicolor and /chrysaegis is sister (BPP = 1, MLB = 100%) to /leoninus (= the P. leoninus complex).
The P. leoninus complex includes collections identified as P. leoninus, P. roseipes, P. flavofuligineus, and two provisionally undescribed taxa (Pluteus aff. leoninus I and Pluteus aff. leoninus II).
The three phylogenetic major clades recognized in sect. Hispidoderma by Justo et al. (2011a, b) and Pradeep et al. (2012) are also recovered in our nrITS sequence analyses (Fig. 1), with high statistical support.
Taxonomy
Pluteus variabilicolor Babos, Annls hist.-nat. Mus. natn. hung. 70: 93 (1978).
Pluteus variabilicolor. Macroscopic features. A, C, E. Basidiomata (TL20130521-01, TL20130530-01, VM20130504-01). B. Pileus smooth and lined up to the middle (TL20130521-01). D. Pileus with a rugulose-veined centre (VM20111105-01). F. Stipe and hymenophore (free lamellae with lamellulae) (TL20130521-01). Photos A-B, F by Tomaso Lezzi; C by Massimo Biraghi; and D-E by Vincenzo Migliozzi. Bars = 10 mm.
Pluteus variabilicolor. Microscopic features. A. Portion of pileipellis with both short and elongated elements (in H2O) (TL2030521-01). B, C. Portions of pileipellis with only short elements (in H2O) (TL2030521-01). D. Pleurocystidia (in ammoniacal Congo red) (TL20130530-01). E, F. Cheilocystidia (in ammoniacal Congo red) (TL2030521-01). G. Caulocystidia (in ammoniacal Congo red) (TL20130726-01). H. Basidiospores (in ammoniacal Congo red) (TL2030521-01). Photos A–C, E–H by Tomaso Lezzi; and D by Massimo Biraghi. Bars = 20 µm.
Synonym: Pluteus castri Justo & E.F. Malysheva, Mycol. Progr. 10: 470 (2011).
Selected descriptions: Babos (1978:193–195); Lohmeyer et al. (1994: 96–98); Migliozzi (2011: 4–7); Justo et al. (2011a: 470–471,as P. castri).
Selected illustrations: Lohmeyer et al. (1994: fig. 5, p. 101); Lanconelli et al. (1998: 96, 127); Albert (2011, two unnumbered figs); Migliozzi (2011: 4, 8); Justo et al. (2011a: fg. 5, p. 471, as P. castri).
Description: Basidiomata pluteoid. Pileus medium-sized, 30–80 mm broad, smooth, lined up to middle, colour distinctly yellow-orange, with central darkerumbo, often radially rugulose-veined, especially in mature specimens, hygrophanous, clearly striate or not at margin. Lamellae free, quite crowded, ventricose, first white then pink, with the presence of lamellulae (I = 0–2), with concolorous or whitish, flocculose edge. Stipe 30–70 × 4–12 mm, cylindrical, slightly enlarged at the base, streaked-fibrillose over the entire length, yellow, with reddish tinges at the base in mature specimens. Context white-yellowish, yellowish orange under the pileus surface, without distinctive smell and taste. Spore print pinkish. Basidiospores 5.5−7.0 × 4.5−5.5(−6.0) µm, on average 6.0 × 4.9 µm, Q = 1.10−1.35; Qm = 1.22, broadly ellipsoid to subglobose, thin-walled. Basidia 25−32 × 6−8 µm, clavate, tetrasporic. Cheilocystidia fusiform to lageniform, 50−90 × 25−30 µm, hyaline, thin-walled, frequently with short, wide appendix at the apex (mucronate). Pleurocystidia not frequent, fusiform, lageniform or utriform, 60−160 × 20−40 µm. Pileipellis a hymeniderm consisting of clavate, rounded terminal elements and cylindrical, elongated cells 40−200 × 22−40 µm; with yellow intracellular pigment. In some parts, the hymeniderm with short cells is strongly predominant; in other parts the elongated cells are strongly predominant. Often the two types of elements are mixed independently from whether they are found: in the centre or in the margin of the pileus. Stipitipellis a cutis made of cylindrical, 2–3 µm wide hyphae. Caulocystidia present over the whole length of the stipe, 13−70 × 3−15 µm, cylindrical to claviform, fusiform, sometimes mucronate, usually grouped in clusters. Clamp-connections not observed.
Habitat: On decaying wood of Quercus cerris and branches of Castanea sativa on the ground.
Specimens examined (of P. variabilicolor): Italy: Lazio: Manziana (RM), on decaying wood of Quercus cerris, 21 May 2013, T. Lezzi (TL20130521-01); loc. cit., 26 July 2013, L. Perrone (TL20130726-01); Lombardia: Parco di Monza (MI), on decaying wood of Castanea sativa, 30 May 2013, M. Biraghi (TL20130530-01); Lazio: Manziana (RM) on decaying wood of Quercus cerris, 5 Nov. 2011, V. Migliozzi (VM20111105-01); loc. cit, 4 May 2013, V. Migliozzi (VM20130504-01). —Hungary: nearSzárliget, on a pile of sawdust, 06 July 1977, M. Babos & A. Friesz (BP-FN 56936 — isotype); Budakeszi, on a pile of sawdust, 27 July 1980, L. Babos (BP-FN 65137); Vallis Lepence, near Visegrád, on a pile of sawdust, 31 July 1981, M. Babos (BP-FN 72875); Com. Heves, Mátra-hegység, Gyöngyössolymos, on sawdust, 3 Aug. 2008, G. Vasas & C. Locsmándi (BP-FN 100005).
Specimens examined (of P. castri): Japan: Fukuoka Prefecture: Kyushu, on piled wood chips, 5 May 2007, M. Shintani & S. Takehashi (TNSF 17602 — holotype); Ibaraki Prefecture: Honshu, Tsukuba, 25 Oct. 2007, K. Osaku (TNSF 17081). — Russia: Central Russia, Moscow Region, Prioksko-Terrasny State Reserve, on decaying wood of Populus tremula, 15 Aug. 1991, G. E. Levitskaya (LE 216873); Samara Region, near Pribrezhny, on decaying wood of deciduous tree, 1 July 2007, E. F. Malysheva (LE 212090).
Discussion
According to our molecular analysis (Fig. 1) the four sequenced collections of Pluteus variabilicolor (isotype included) and the three of P. castri retrieved from GenBank (holotype included) are conspecific. Consequently, we have placed P. castri as a later synonym of P. variabilicolor.
Pluteus variabilicolor was originally described from Hungary and primarily characterized by a pileus which was at first yellowish orange and then becoming chrome-yellow (hence the epithet “variabilicolor”), rugose-veined at the centre, and a pileipellis consisting of dimorphic elements, viz. a mixture of spheropeduncolate-vesiculose and elongated, more or less cystidioid elements, the presence of caulocystidia, and growth on decaying sawdust (Babos 1978). Because of the pileipellis structure, it was included in subsect. Mixtini of sect. Celluloderma. Apart from Hungary, where it was found several times after the first report (e.g. Babos 1981, 1989, 1991, Rimóczi & Vetter 1990, Lukács 2010, Albert 2011), P. variabilicolor is reported only from Austria (Lohmeyer et al. 1994), Italy (Lanconelli et al. 1998, Migliozzi 2011), Romania (Béres 2012), Slovenia (Jogan et al. 2012), and Moldova (E. Musumeci, pers. comm.).
Pluteus variabilicolor is mainly reported from sawdust deposits, even though our finds and those of Migliozzi (2011) and Musumeci (pers. comm.) are on branches and rotten wood of Fagaceae (mainly Quercus species).
As suggested by Babos (1978, 1981) and Courtecuisse (2006), the species may not be native to Europe and could have been introduced through timber import. In support of this hypothesis, all the nations in which the species was found are geographically related. On the other hand, the species is widely distributed from Europe to Japan in relatively “natural” habitats, so it may very well be a native Eurasian species that is just not common.
Our description, based on Italian specimens, fits quite well with the protologue, with the morphologic features reported by later authors (Lohmeyer et al. 1994, Lanconelli et al. 1998, Migliozzi 2011), and the Hungarian collections we have checked (isotype included).
Pluteus castri was recently described (Justo et al. 2011a) based on collections from Japan (on piled wood chips, holotype) and Central Russia (on decaying wood of deciduous trees). The main difference with respect to the original description of P. variabilicolor and the Italian collections found was in the structure of the pileipellis. In P. castri we did not observe the elongated elements (to 200 µm long) found in other collections of P. variabilicolor. However, molecular data indicate that all collections of P. castri should be considered to represent P. variabilicolor.
In the phylogenetic analysis (Fig. 1) P. variabilicolor, in spite of the presence of both vesiculose-clavate and elongate elements in the pileipellis, clustered into the leoninus clade of sect. Hispidoderma as delimited by Justo et al. (2011a, b) and Pradeep et al. (2012), where it is sister to P. chrysaegis and very close to the P. leoninus complex. Most species of the leoninus clade are characterized by yellowish tinges on the pileus surface.
Pluteus chrysaegis (syn. P. conizatus var. africanus) resembles P. variabilicolor in the general structure of the pileipellis and the yellow colouring of the basidiome, the veined pileus surface, and the presence of caulocystidia (Pradeep & Vrinda 1996, Pradeep et al. 2012). However, it differs in the even shorter elements of the pileipellis (to 40 µm long), and the non-mucronate, slightly thick-walled cheilocystidia (Horak & Heinemann 1978, Pradeep et al. 2012).
The species in the P. leoninus complex (i.e. P. leoninus, P. roseipes, P. flavofuligineus, P. aff. leoninus I and P. aff. leoninus II) all have a pileipellis made up exclusively of narrowly fusiform elements to 230 µm long (Justo et al. 2011a) and, with the exception of P. aff. leoninus II, a stipitipellis without caulocystidia. The separation of the species inside this complex still requires further study.
References
Albert L (ed.) (2011) Szines oldalak. Clusiana 50: 119–134.
Babos M (1978) Pluteus studies, I. (Basidiomycetes, Pluteaceae). Annales Historico-Naturales Musei Nationalis Hungarici, Budapest 70: 93–97.
Babos M (1981) Mycological examination of sawdust depots in Hungary. Studia Botanica Hungarica 15: 31–44.
Babos M (1989) Magyarország kalaposgombáinak (Agaricales s.l.) jegyzéke. (The Agaricales s.l. taxa of Hungary). Mikologiai Közlemények, Clusiana 28: 3–234.
Babos M (1991) Bazídiumos nagygombák. In: Baktérium-, alga, gomba-, zuzmó-és mohahatározó (Simon T, ed.): 403–574. Budapest: Tankönyvkiadö.
Banerjee P (1992) A systematic and phylogenetic study of the genus Pluteus with special reference to Section Pluteus. Doctoral dissertation, Southern Illinois University.
Banerjee P, Sundberg J (1993) Reexamination of Pluteus type specimens: types housed at the New York Botanical Garden. Mycotaxon 49: 413–435.
Béres M (2012) Macromycetes species included in Bern Convention Appendix in the Red List for Romania, and rare presence in historical Maramures area (Romania). Acta Oecologica Carpatica 5: 19–38.
Binder M, Hibbett DS, Wang Z, Farnham W (2006) Evolutionary relationships of Mycaureola dilseae (Agaricales), a basidiomycete pathogen of a subtidal rhodophyte. American Journal of Botany 93: 547–556.
Bodensteiner P, Binder M, Moncalvo JM, Agerer R, Hibbett DS (2004) Phylogenetic relationships of cyphelloid homobasidiomycetes. Molecular Phylogenetics and Evolution 33: 501–515.
Citérin M, Eyssartier G (1998) Clé analytique du genre Pluteus Fr. Documents Mycologiques 28: 47–67.
Courtecuisse R (2006) La notion d’espèce invasive appliquée aux champignons — Etat des lieux, en particulier en France. La Lettre de la Société Mycologique de France 8: 5–8.
Drummond AJ, Ashton B, Cheung M, Heled J, Kearse M, et al. (2010) Geneious. Version 5.3. http://www.geneious.com/
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39: 783–791.
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes — application to the identification of mycorrhizae and rusts. Molecular Ecology 2: 113–118.
Heilmann-Clausen J (2012) Pluteus Fr. In: Funga Nordica:agaricoid, boletoid, clavarioid, cyphelloid and gastroid genera (Knudsen H, Vesterholt J, eds) 1: 386–395. 2nd edn. Copenhagen: Nordsvamp.
Homola RL (1975) Phylogenetic relationships within the genus Pluteus. Beihefte zurNova Hedwigia 51: 139–144.
Horak E, Heinemann P (1978) Flore Illustrée des Champignons d’Afrique Centrale. Vol. 6. Pluteus & Volvariella (compléments). Meise: National Botanical Garden of Belgium.
Huelsenbeck JP, Ronquist F (2001) MrBayes: Bayesian inference of phylogeny. Bioinformatics 17: 754–755.
Jogan N, Bačič T, Strgulc-Krajšek S (2012) Neobiota Slovenije: Invazivne tujerodne vrste v Sloveniji ter vpliv na ohranjanje biotske raznovrstnosti in trajnostno rabo virov. [Končno poročilo projekta CRP, Konkurenčnost Slovenije 2006–2013]. Ljubljana: Univerza v Ljubljani, Biotehniška fakulteta.
Justo A, Minnis AM, Ghignone S, Menolli N, Capelari M, et al. (2011a) Species recognition in Pluteus and Volvopluteus (Pluteaceae, Agaricales): morphology, geography and phylogeny. Mycological Progress 10: 453–479.
Justo A, Vizzini A, Minnis AM, Menolli N, Capelari M, et al. (2011b) Phylogeny of the Pluteaceae (Agaricales, Basidiomycota): taxonomy and characterevolution. Fungal Biology 115: 1–20.
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Research 30: 3059–3066.
Lanconelli L, Ballanti F, Rava M (1998) Funghi del Lughese. Faenza: Edit. Faenza.
Lohmeyer TR, Christan J, Gruber O (1994) Ein Nachweis von Pluteus variabilicolor in Oberösterreich. Österreichische Zeitschrift für Pilzkunde 3: 95–100.
Lukács Z (2010) Újabb adatok Magyarország gombavilágához IV. Mikológiai közlemények, Clusiana 49: 79–119.
Malysheva EF, Malysheva VF, Krasilnikova AA (2009) Morphological and molecular approaches to study the genus Pluteus Fr. Mikologiya i Fitopatologiya 43: 216–231.
Matheny PB, Curtis JC, Hofstetter V, Aime MC, Moncalvo JM, et al. (2006) Major clades of Agaricales: a multi-locus phylogenetic overview. Mycologia 98: 982–995.
Migliozzi V (2011) Pluteus variabilicolor, specie frequente nella cerreta di Macchiagrande di Manziana (RM). Parliamo di funghi 19(1): 3–9.
Minnis AM, Sundberg WJ (2009) Preliminary notes on Pluteus phylogeny. Nova Hedwigia 89: 303–319.
Minnis AM, Sundberg WJ (2010) Pluteus section Celluloderma in the USA. North American Fungi 5: 1–107.
Moncalvo JM, Lutzoni FM, Rehner SA, Johnson J, Vilgalys R (2000) Phylogenetic relationships of agaric fungi based on nuclear large subunit ribosomal DNA sequences. Systematic Biology 49: 278–305.
Moncalvo JM, Vilgalys R, Redhead SA, Johnson JE, James TY, et al. (2002) One hundred and seventeen clades of euagarics. Molecular Phylogenetics and Evolution 23: 357–400.
Orton PD (1986) British Fungus Flora Agarics and Boleti. Vol. 4. Pluteaceae: Pluteus & Volvariella. Edinburgh: Royal Botanic Garden Edinburgh.
Posada D (2008) jModeltest: phylogenetic model averaging. Molecular Biology and Evolution 25: 1253–1256.
Pradeep CK, Vrinda KB (2006) New and noteworthy species of Pluteus (Pluteaceae, Agaricales) from Kerala state, India. Persoonia 19: 95–99.
Pradeep CK, Justo A, Vrinda KB, Shibu VP (2012) Two new species of Pluteus (Pluteaceae, Agaricales) from India and additional observations on Pluteus chrysaegis. Mycological Progress 11: 869–878.
Rimóczi I, Vetter J (eds) (1990) Gombahatározó 1–2. (Polyporales, Boletales, Agaricales, Russulales). Budapest: Országos Erdészeti Egyesület Mikológiai Társasága.
Rodriguez O, Guzmán-Dávalos L (2001) Clave dicotômica de las especies del género Pluteus Fr. (Pluteaceae) conocidas de la region de Nueva Galicia y algunas areas aledanas, México. Acta Botanica Mexicana 57: 23–36.
Singer R (1956) Contributions towards a monograph of the genus Pluteus. Transanctions of the British Mycological Society 39: 145–232.
Singer R (1958) Monographs of South American Basidiomycetes, especially those of the east slope of the Andes and Brazil 1. The genus Pluteus in South America. Lloydia 21: 195–299.
Singer R (1986) The Agaricales in Modern Taxonomy. 4th edn. Koenigstein: Koeltz Scientific Books.
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22: 2688–2690.
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, et al. (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution 28: 2731–2739.
Vellinga EC (1988) Glossary. In: Flora Agaricina Neerlandica (Bas C, Kuyper TW, Noordeloos ME, Vellinga EC, eds) 1: 54–64. Rotterdam: AA Balkema.
Vellinga EC (1990) Pluteus Fr. In: Flora Agaricina Neerlandica (Bas C, Kuyper TW, Noordeloos ME, Vellinga EC, eds): 31–55. Rotterdam: AA Balkema.
Vellinga EC, Schreurs J (1985) Pluteus Fr. in Western Europe. Persoonia 12: 337–373.
Vizzini A, Para R, Fontenla R, Ghignone S, Ercole E (2011) A preliminary ITS phylogeny of Melanoleuca (Agaricales) with special reference to European taxa. Mycotaxon 118: 361–381.
White TJ, Bruns TD, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: PCR Protocols: a guide to methods and applications (Innis MA, Gelfand D, Sninsky J, White T, eds): 315–322. San Diego: Academic Press.
Acknowledgements
We are grateful to Massimo Biraghi (Arcene, Bergamo) and Luigi Perrone (Rome) who put some collections to compare at our disposal. We also thank Gizella Vasas (BP, Budapest) for the loan of the isotype material of P. variabilicolor and othercollections.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by-nc/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Lezzi, T., Vizzini, A., Ercole, E. et al. Phylogenetic and morphological comparison of Pluteus variabilicolor and P. castri (Basidiomycota, Agaricales). IMA Fungus 5, 415–423 (2014). https://doi.org/10.5598/imafungus.2014.05.02.06
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.5598/imafungus.2014.05.02.06