Abstract
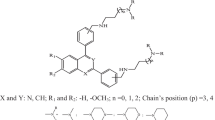
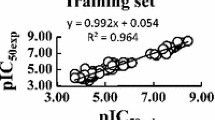
A data set of amidino bis-benzimidazoles, in particular 2′-arylsubstituted-1H,1′H-[2,5′]bisbenzimidazolyl-5-carboximidine derivatives with anti-malarial activity against Plasmodium falciparum was employed in investigating the quantitative structure-activity relationship (QSAR). Quantum chemical and molecular descriptors were obtained from B3LYP/6-31g(d) calculations and Dragon software, respectively. Significant variables, which included total energy (E T), highest occupied molecular orbital (HOMO), Moran autocorrelation-lag3/weighted by atomic masses (MATS3m), Geary autocorrelation-lag8/weighted by atomic masses (GATS8m), and 3D-MoRSEsignal 11/weighted by atomic Sanderson electronegativities (Mor11e), were used in the construction of QSAR models using multiple linear regression (MLR) and artificial neural network (ANN). The results indicated that the predictive models for both the MLR and ANN approaches using leave-one-out cross-validation afforded a good performance in modelling the anti-malarial activity against P. falciparum as observed by correlation coefficients of leave-one-out cross-validation (R LOO-CV) of 0.9760 and 0.9821, respectively, root mean squared error of leave-one-out cross-validation (RMSELOO-CV) of 0.1301 and 0.1102, respectively, and predictivity of leave-one-out cross-validation (Q 2LOO-CV ) of 0.9526 and 0.9645, respectively. Model validation was performed using an external testing set and the results suggested that the model provided good predictivity for both MLR and ANN models with correlation coefficient of the external set (R Ext) values of 0.9978 and 0.9844, respectively, root mean squared error of the external set (RMSEExt) of 0.0764 and 0.1302 respectively, and predictivity of the external set (Q 2Ext ) of 0.9956 and 0.9690, respectively. Furthermore, the robustness of the QSAR models is corroborated by a number of statistical parameters, comprising adjusted correlation coefficient (R 2Adj ), standard deviation (s), predicted residual sum of squares (PRESS), standard error of prediction (SDEP), total sum of squares deviation (SSY), and quality factor (Q). The QSAR models so constructed provide pertinent insights for the future design of anti-malarial agents.
Similar content being viewed by others
References
Afantitis, A., Melagraki, G., Sarimveis, H., Koutentis, P. A., Markopoulos, J., & Igglessi-Markopoulou, O. (2006). A novel QSAR model for predicting induction of apoptosis by 4-aryl-4H-chromenes. Bioorganic & Medicinal Chemistry, 14, 6686–6694. DOI: 10.1016/j.bmc.2006.05.061.
Alp, M., Göker, H., Brun, R., & Yıldız, S. (2009). Synthesis and antiparasitic and antifungal evaluation of 2′-arylsubstituted-1H,1′H-[2,5′]bisbenzimidazolyl-5-carboxamidines. European Journal of Medicinal Chemistry, 44, 2002–2008. DOI: 10.1016/j.ejmech.2008.10.003.
Alves, C. N., Pinheiro, J. C., Camargo, A. J., Ferreira, M. M. C., Romero, R. A. F., & da Silva, A. B. F. (2001). A multiple linear regression and partial least squares study of flavonoid compounds with anti-HIV activity. Journal of Molecular Structure: THEOCHEM, 541, 81–88. DOI: 10.1016/s0166-1280(00)00755-7.
Bloland, P. B. (2001). Drug resistance in malaria. Retrieved November 25, 2011, from http://www.who.int/csr/resources/publications/drugresist/malaria.pdf
Caballero, J., & Fernández, M. (2006). Linear and nonlinear modeling of antifungal activity of some heterocyclic ring derivatives using multiple linear regression and Bayesian-regularized neural networks. Journal of Molecular Modeling, 12, 168–181. DOI: 10.1007/s00894-005-0014-x.
Cao, D. S., Liang, Y. Z., Xu, Q. S., Li, H. D., & Chen, X. (2010). A new strategy of outlier detection for QSAR/QSPR. Journal of Computational Chemistry, 31, 592–602. DOI: 10.1002/jcc.21351.
Castillo-Garit, J. A., Marrero-Ponce, Y., Escobar, J., Torrens, F., & Rotondo, R. (2008). A novel approach to predict aquatic toxicity from molecular structure. Chemosphere, 73, 415–427. DOI: 10.1016/j.chemosphere.2008.05.024.
Del Poeta, M., Schell, W. A., Dykstra, C. C., Jones, S. K., Tidwell, R. R., Kumar, A., Boykin, D. W., & Perfect, J. R. (1998). In vitro antifungal activities of a series of dicationsubstituted carbazoles, furans, and benzimidazoles. Antimicrobial Agents and Chemotherapy, 42, 2503–2510.
Dennington, R., II, Keith, T., Millam, J., Eppinnett, K., Hovell, W. L., & Gilliland, R. (2003). GaussView, Version 3.09 [computer software]. Shawnee Mission, KS, USA: Semichem.
Durand, A. C., Farce, A., Carato, P., Dilly, S., Yous, S., Berthelot, P., & Chavatte, P. (2007). Quantitative structureactivity relationships studies of antioxidant hexahydropyridoindoles and flavonoid derivatives. Journal of Enzyme Inhibition and Medicinal Chemistry, 22, 556–562. DOI: 10.1080/14756360701425238.
Farooq, U., & Mahajan, R. C. (2004). Drug resistance in malaria. Journal of Vector Borne Diseases, 41, 45–53.
Fernández, M., Caballero, J., Helguera, A. M., Castro, E. A., & González, M. P. (2005). Quantitative structure-activity relationship to predict differential inhibition of aldose reductase by flavonoid compounds. Bioorganic & Medicinal Chemistry, 13, 3269–3277. DOI: 10.1016/j.bmc.2005.02.038.
Frisch, M. J., Trucks, G. W., Schlegel, H. B., Scuseria, G. E., Robb, M. A., Cheeseman, J. R., Montgomery, J. A., Jr., Vreven, T., Kudin, K. N., Burant, J. C., Millam, J. M., Iyengar, S. S., Tomasi, J., Barone, V., Mennucci, B., Cossi, M., Scalmani, G., Rega, N., Petersson, G. A., Nakatsuji, H., Hada, M., Ehara, M., Toyota, K., Fukuda, R., Hasegawa, J., Ishida, M., Nakajima, T., Honda, Y., Kitao, O., Nakai, H., Klene, M., Li, X., Knox, J. E., Hratchian, H. P., Cross, J. B., Bakken, V., Adamo, C., Jaramillo, J., Gomperts, R., Stratmann, R. E., Yazyev, O., Austin, A. J., Cammi, R., Pomelli, C., Ochterski, J. W., Ayala, P. Y., Morokuma, K., Voth, G. A., Salvador, P., Dannenberg, J. J., Zakrzewski, V. G., Dapprich, S., Daniels, A. D., Strain, M. C., Farkas, O., Malick, D. K., Rabuck, A. D., Raghavachari, K., Foresman, J. B., Ortiz, J. V., Cui, Q., Baboul, A. G., Clifford, S., Cioslowski, J., Stefanov, B. B., Liu, G., Liashenko, A., Piskorz, P., Komaromi, I., Martin, R. L., Fox, D. J., Keith, T., Al-Laham, M. A., Peng, C. Y., Nanayakkara, A., Challacombe, M., Gill, P. M. W., Johnson, B., Chen, W., Wong, M. W., Gonzalez, C., & Pople, J. A. (2004). Gaussian 03, Revision C.02 [computer software]. Wallingford, CT, USA: Gaussian.
García, J., Duchowicz, P. R., Rozas, M. F., Caram, J. A., Mirífico, M. V., Fernández, F. M., & Castro, E. A. (2011). A comparative QSAR on 1,2,5-thiadiazolidin-3-one 1,1-dioxide compounds as selective inhibitors of human serine proteinases. Journal of Molecular Graphics and Modelling, 31, 10–19. DOI: 10.1016/j.jmgm.2011.07.007.
Gogtay, N. J., Kshirsagar, N. A., & Vaidya, A. B. (2006). Current challenges in drug-resistant malaria. Journal of Postgraduate Medicine, 52, 241–242.
Gramatica, P. (2007). Principles of QSAR models validation: internal and external. QSAR & Combinatorial Science, 26, 694–701. DOI: 10.1002/qsar.200610151.
Greenwood, B. M., Bojang, K., Whitty, C. J., & Targett, G. A. (2005). Malaria. Lancet, 365, 1487–1498. DOI: 10.1016/s0140-6736(05)66420-3.
Gupta, L., Patel, A., Karthikeyan, C., & Trivedi, P. (2010). QSAR studies on dihydro-alkoxy-benzyl-oxopyrimidines (DABOs) derivatives, a new series of potent, broad-spectrum non-nucleoside reverse transcriptase inhibitors. Journal of Current Pharmaceutical Research, 1, 19–25.
Idro, R., Jenkins, N. E., & Newton, C. R. J. C. (2005). Pathogenesis, clinical features, and neurological outcome of cerebral malaria. Lancet Neurology, 4, 827–840. DOI: 10.1016/s1474-4422(05)70247-7.
Idro, R., Marsh, K., John, C. C., & Newton, C. R. J. (2010). Cerebral malaria: Mechanisms of brain injury and strategies for improved neurocognitive outcome. Pediatric Research, 68, 267–274. DOI: 10.1203/pdr.0b013e3181eee738.
Jain, A. K., Veerasamy, R., Vaidya, A., Kashaw, S., Mourya, V. K., & Agrawal, R. K. (2012). QSAR analysis of B-ring-modified diaryl ether derivatives as a InhA inhibitors. Medicinal Chemistry Research, 21, 145–151. DOI: 10.1007/s00044-010-9518-8.
Jalali-Heravi, M., & Parastar, F. (2000). Use of artificial neural networks in a QSAR study of anti-HIV activity for a large group of HEPT derivatives. Journal of Chemical Information and Modeling, 40, 147–154. DOI: 10.1021/ci990314+.
Karelson, M., Lobanov, V. S., & Katritzky, A. R. (1996). Quantum-chemical descriptors in QSAR/QSPR studies. Chemical Reviews, 96, 1027–1044. DOI: 10.1021/cr950202r.
Khosrokhavar, R., Ghasemi, J. B., & Shiri, F. (2010). 2D quantitative structure-property relationship study of mycotoxins by multiple linear regression and support vector machine. International Journal of Molecular Sciences, 11, 3052–3068. DOI: 10.3390/ijms11093052.
Kshirsagar, N. A. (2006). Malaria: Antimalarial resistance and policy ramifications and challenges. Journal of Postgraduate Medicine, 52, 291–293.
Nantasenamat, C., Naenna, T., Isarankura Na Ayudhya, C., & Prachayasittikul, V. (2005). Quantitative prediction of imprinting factor of molecularly imprinted polymers by artificial neural network. Journal of Computer-Aided Molecular Design, 19, 509–524. DOI: 10.1007/s10822-005-9004-4.
Nantasenamat, C., Isarankura-Na-Ayudhya, C., Tansila, N., Naenna, T., & Prachayasittikul, V. (2007). Prediction of GFP spectral properties using artificial neural network. Journal of Computational Chemistry, 28, 1275–1289. DOI: 10.1002/jcc.20656.
Nantasenamat, C., Isarankura-Na-Ayudhya, C., Naenna, T., & Prachayasittikul, V. (2008). Prediction of bond dissociation enthalpy of antioxidant phenols by support vector machine. Journal of Molecular Graphics and Modelling, 27, 188–196. DOI: 10.1016/j.jmgm.2008.04.005.
Nantasenamat, C., Isarankura-Na-Ayudhya, C., Naenna, T., & Prachayasittikul, V. (2009). A practical overview of quantitative structure-activity relationship. EXCLI Journal, 8, 74–88.
Nwaka, S., & Ridley, R. G. (2003). Virtual drug discovery and development for neglected diseases through public-private partnerships. Nature Reviews Drug Discovery, 2, 919–928. DOI: 10.1038/nrd1230.
Nwaka, S., & Hudson, A. (2006). Innovative lead discovery strategies for tropical diseases. Nature Reviews Drug Discovery, 5, 941–955. DOI: 10.1038/nrd2144.
Papa, E., Villa, F., & Gramatica, P. (2005). Statistically validated QSARs, beased on theoretical descriptors, for modeling aquatic toxicity of organic chemicals in Pimephales promelas (fathead minnow). Journal of Chemical Information and Modeling, 45, 1256–1266. DOI: 10.1021/ci050212l.
Parr, R. G., Donnelly, R. A., Levy, M., & Palke, W. E. (1978). Electronegativity: The density functional viewpoint. Journal of Chemical Physics, 68, 3801. DOI: 10.1063/1.436185.
Parr, R. G., & Pearson, R. G. (1983). Absolute hardness: companion parameter to absolute electronegativity. Journal of the American Chemical Society, 105, 7512–7516. DOI: 10.1021/ja00364a005.
Parr, R. G., Szentpály, L., & Liu, S. (1999). Electrophilicity index. Journal of the American Chemical Society, 121, 1922–1924. DOI: 10.1021/ja983494x.
Podunavac-Kuzmanović, S. O., & Cvetković, D. D. (2011). QSAR modeling of antibacterial activity of some benzimidazole derivatives. Chemical Industry & Chemical Engineering Quarterly, 17, 33–38. DOI: 10.2298/ciceq100405050p.
Prachayasittikul, S., Wongsawatkul, O., Worachartcheewan, A., Nantasenamat, C., Ruchirawat, S., & Prachayasittikul, V. (2010). Elucidating the structure-activity relationships of the vasorelaxation and antioxidation properties of thionicotinic acid derivatives. Molecules, 15, 198–214. DOI: 10.3390/molecules15010198.
Prasad, Y. R., Kumar, P. R., Smiles, D. J., & Babu, P. A. (2008). QSAR studies on chalcone derivatives as antibacterial agents against Bacillus pumilis. ARKIVOC, 11, 266–276.
Rastija, V., & Medić-Šarić, M. (2009). QSAR modeling of anthocyanins, anthocyanidins and catechins as inhibitors of lipid peroxidation using three-dimensional descriptors. Medicinal Chemistry Research, 18, 579–588. DOI: 10.1007/s00044-008-9151-y.
Saghaie, L., Sakhi, H., Sabzyan, H., Shahlaei, M., & Shamshirian, D. (2013). Stepwise MLR and PCR QSAR study of the pharmaceutical activities of antimalarial 3-hydroxypyridinone agents using B3LYP/6-311++G** descriptors. Medicinal Chemistry Research, 22, 1679–1688. DOI: 10.1007/s00044-012-0152-5.
Sawant, R. L., Bansode, C. A., & Wadekar, J. B. (2013). In vitro anti-inflammatory potential and QSAR analysis of oxazolo/thiazolo pyrimidine derivatives. Medicinal Chemistry Research, 22, 1884–1892. DOI: 10.1007/s00044-012-0189-5.
Shahlaei, M., Fassihi, A., & Nezami, A. (2009). QSAR study of some 5-methyl/trifluoromethoxy-1H-indole-2,3-dione-3-thiosemicarbazone derivatives as anti-tubercular agents. Research in Pharmaceutical Sciences, 4, 123–131.
Suksrichavalit, T., Prachayasittikul, S., Nantasenamat, C., Isarankura-Na-Ayudhya, C., & Prachayasittikul, V. (2009). Copper complexes of pyridine derivatives with superoxide scavenging and antimicrobial activities. European Journal of Medicinal Chemistry, 44, 3259–3265. DOI: 10.1016/j.ejmech.2009.03.033.
Sun, M., Zheng, Y. G., Wei, H. T., Chen, J. Q., & Ji, M. (2009). QSAR studies on 4-anilino-3-quinolinecarbonitriles as Src kinase inhibitors using robust PCA and both linear and nonlinear models. Journal of Enzyme Inhibition and Medicinal Chemistry, 24, 1109–1116. DOI: 10.1080/14756360802632906.
Suvannang, N., Nantasenamat, C., Isarankura-Na-Ayudhya, C., & Prachayasittikul, V. (2011). Molecular docking of aromatase inhibitors. Molecules, 16, 3597–3617. DOI: 10.3390/molecules16053597.
Talete (2007). Dragon for windows (software for molecular descriptor calculations), version 5.5 [computer software]. Milano, Italy: Talete.
Tanious, F. A., Hamelberg, D., Bailly, C., Czarny, A., Boykin, D. W., & Wilson, W. D. (2004). DNA sequence dependent monomer-dimer binding modulation of asymmetric benzimidazole derivatives. Journal of the American Chemical Society, 126, 143–153. DOI: 10.1021/ja030403+.
Thanikaivelan, P., Subramanian, V., Raghava Rao, J., & Unni Nair, B. (2000). Application of quantum chemical descriptor in quantitative structure activity and structure property relationship. Chemical Physics Letters, 323, 59–70. DOI: 10.1016/s0009-2614(00)00488-7.
Thippakorn, C., Suksrichavalit, T., Nantasenamat, C., Tantimongcolwat, T., Isarankura-Na-Ayudhya, C., Naenna, T., & Prachayasittikul, V. (2009). Modeling the LPS neutralization activity of anti-endotoxins. Molecules, 14, 1869–1888. DOI: 10.3390/molecules14051869.
Vahdani, S., & Bayat, Z. (2011). A quantitative structure-activity relationship (QSAR) study of anti-cancer drugs. Der Chemica Sinica, 2, 235–243.
Veerasamy, R., Ravichandran, S., Jain, A., Rajak, H., & Agrawal, R. K. (2009). QSAR studies on novel anti-HIV agents using FA-MLR, FA-PLS and PCRA techniques. Digest Journal of Nanomaterials and Biostructures, 4, 823–834.
Verma, R. P. (2006). Anti-cancer activities of 1,4-naphthoquinones: a QSAR study. Anti-Cancer Agents in Medicinal Chemistry, 6, 489–499. DOI: 10.2174/187152006778226512.
Verma, R. P., & Hansch, C. (2010). QSAR modeling of taxane analogues against colon cancer. European Journal of Medicinal Chemistry, 45, 1470–1477. DOI: 10.1016/j.ejmech.2009.12.054.
Vlaia, V., Olariu, T., Vlaia, L., Butur, M., Ciubotariu, C., Medeleanu, M., & Ciubotariu, D. (2009). Quantitative structure-activity relationship (QSAR). V. Analysis of the toxicity of aliphatic esters by means of molecular compressibility descriptors. Farmacia, 57, 549–561.
Whitley, D. C., Ford, M. G., & Livingstone, D. J. (2000). Unsupervised forward selection: A method for eliminating redundant variables. Journal of Chemical Information and Modeling, 40, 1160–1168. DOI: 10.1021/ci000384c.
White, N. J. (2004). Antimalarial drug resistance. Journal of Clinical Investigation, 113, 1084–1092. DOI: 10.1172/jci200421682.
Witten, I. H., Frank, E., & Hall, M. A. (2011). Data mining: practical machine learning tools and techniques (3rd ed.). San Francisco, CA, USA: Morgan Kaufmann.
Worachartcheewan, A., Nantasenamat, C., Naenna, T., Isarankura-Na-Ayudhya, C., & Prachayasittikul, V. (2009). Modeling the activity of furin inhibitors using artificial neural network. European Journal of Medicinal Chemistry, 44, 1664–1673. DOI: 10.1016/j.ejmech.2008.09.028.
Worachartcheewan, A., Nantasenamat, C., Isarankura-Na-Ayudhya, C., Prachayasittikul, S., & Prachayasittikul, V. (2011). Predicting the free radical scavenging activity of curcumin derivatives. Chemometrics and Intelligent Laboratory Systems, 109, 207–216. DOI: 10.1016/j.chemolab.2011.09.010.
Worachartcheewan, A., Prachayasittikul, S., Pingaew, R., Nantasenamat, C., Tantimongcolwat, T., Ruchirawat, S., & Prachayasittikul, V. (2012). Antioxidant, cytotoxicity, and QSAR study of 1-adamantylthio derivatives of 3-picoline and phenylpyridines. Medicinal Chemistry Research, 21, 3514–3522. DOI: 10.1007/s00044-011-9903-y.
Yan, X. F. (2011). Multivariate outlier detection based on selforganizing map and adaptive nonlinear map and its application. Chemometrics and Intelligent Laboratory Systems, 107, 251–257. DOI: 10.1016/j.chemolab.2011.04.007.
Zahouily, M., Lazar, M., Elmakssoudi, A., Rakik, J., Elaychi, S., & Rayadh, A. (2006). QSAR for anti-malarial activity of 2-aziridinyl and 2,3-bis(aziridinyl)-1,4-naphthoquinonyl sulfonate and acylate derivatives. Journal of Molecular Modeling, 12, 398–405. DOI: 10.1007/s00894-005-0059-x.
Zhang, L., Zhu, H., Oprea, T. I., Golbraikh, A., & Tropsha, A. (2008). QSAR modeling of the blood-brain barrier permeability for diverse organic compounds. Pharmaceutical Research, 25, 1902–1914. DOI: 10.1007/s11095-008-9609-0.
Zou, C., & Zhou, L. (2007). QSAR study of oxazolidinone antibacterial agents using artificial neural networks. Molecular Simulation, 33, 517–530. DOI: 10.1080/08927020601188528.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Worachartcheewan, A., Nantasenamat, C., Isarankura-Na-Ayudhya, C. et al. QSAR study of amidino bis-benzimidazole derivatives as potent anti-malarial agents against Plasmodium falciparum . Chem. Pap. 67, 1462–1473 (2013). https://doi.org/10.2478/s11696-013-0398-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11696-013-0398-5