Abstract
The nuclear protein high mobility group box protein 1 (HMGB1) promotes inflammation upon extracellular release. HMGB1 induces proinflammatory cytokine production in macrophages via Toll-like receptor (TLR)-4 signaling in a redox-dependent fashion. Independent of its redox state and endogenous cytokine-inducing ability, HMGB1 can form highly immunostimulatory complexes by interaction with certain proinflammatory mediators. Such complexes have the ability to enhance the induced immune response up to 100-fold, compared with induction by the ligand alone. To clarify the mechanisms for these strong synergistic effects, we studied receptor requirements. Interleukin (IL)-6 production was assessed in supernatants from cultured peritoneal macrophages from mice each deficient in one of the HMGB1 receptors (receptor for advanced glycation end products [RAGE], TLR2 or TLR4) or from wild-type controls. The cultures were stimulated with the TLR4 ligand lipopolysaccaride (LPS), the TLR2 ligand Pam3CysSerLys4 (Pam3CSK4), noninflammatory HMGB1 or each TLR ligand in complex with noninflammatory HMGB1. The activity of the HMGB1-TLR ligand complexes relied on engagement of the same receptor as for the noncomplexed TLR ligand, since HMGB1-LPS complexes used TLR4 and HMGB1-Pam3CSK4 complexes used TLR2. Deletion of any of the intracellular adaptor molecules used by TLR2 (myeloid differentiation factor-88 (MyD88), TIR domain-containing adaptor protein (TIRAP)) or TLR4 (MyD88, TIRAP, TIR domain-containing adaptor-inducing interferon-β (TRIF), TRIF-related adaptor molecule (TRAM)) had similar effects on HMGB1 complex activation compared with noncomplexed LPS or Pam3CSK4. This result implies that the enhancing effects of HMGB1-partner molecule complexes are not regulated by the induction of additional signaling cascades. Elucidating HMGB1 receptor usage in processes where HMGB1 acts alone or in complex with other molecules is essential for the understanding of basic HMGB1 biology and for designing HMGB1-targeted therapies.
Similar content being viewed by others
Introduction
High mobility group box protein 1 (HMGB1) is a ubiquitous nuclear protein that binds and bends DNA and facilitates transcription. HMGB1 can also be released to the extracellular environment, where it mediates inflammation and is implicated in the pathogenesis of many inflammatory diseases. Therapeutic HMGB1 blockade has proven to be beneficial in a number of preclinical models of inflammation, including arthritis, sepsis, transplantation and ischemia (rev. in 1). HMGB1 exerts proinflammatory effects by induction of cytokine production, recruitment and activation of dendritic cells and upregulation of adhesion molecules on endothelial cells (2). HMGB1 release occurs via active secretion or via leakage from injured cells. A number of cells including monocytes, macrophages, NK cells, dendritic cells, endothelial cells and platelets secrete HMGB1 on stimulation with exogenous or endogenous stimuli (3–6), whereas dying cells release HMGB1 passively (7). Necrotic cells in particular may release high amounts of HMGB1 (8), which may generate high systemic levels of HMGB1 in patients with sepsis, ischemia, hemorrhagic shock and additional conditions (9–11).
Studies on the cytokine-inducing properties of HMGB1 over the past years have given conflicting results. Although some reports demonstrate potent cytokine production after stimulation with HMGB1 (12,13), others report a lack of this ability (14,15). Emerging evidence now indicates that the cytokine-inducing ability of HMGB1 depends on its redox status, explaining the variable results (16,17). Interestingly, several groups have recently demonstrated that preparations of HMGB1, which do not induce cytokine synthesis, may form complexes with exogenous or endogenous proinflammatory molecules to strongly enhance their cytokine-inducing ability. For example, stimulation with preformed HMGB1-lipopolysaccaride (LPS) complexes induces a 100-fold stronger response than stimulation with comparable levels of LPS alone (14). In addition to the Toll-like receptor (TLR)-4 ligand LPS, HMGB1 binds to Pam3CysSerLys4 (Pam3CSK4) (TLR2 ligand) and CpG oligodeoxynucleotides (TLR9 ligand) and enhances the stimulatory effect of these molecules (14,18–21). Furthermore, HMGB1 can bind to and enhance the activity of endogenous molecules, including interleukin (IL)-1α and IL-1β (22,23), stromal cell-derived factor-1 (SDF-1/CXCL12) (24) and nucleosomes (25). HMGB1-nucleosome complexes can break immunological tolerance to dsDNA and induce cytokine production and dendritic cell activation, which HMGB1-free nucleosomes are unable to do (25). It has recently been reported that the activation of nucleic acid-sensing cytosolic receptors and intracellular TLRs requires the nucleic acids to be presented bound on HMGB molecules (HMGB1, HMGB2 or HMGB3). Nucleic acid-induced type I interferon (IFN) synthesis is substantially diminished in cells with suppressed HMGB expression (26,27). However, the mechanisms, receptor engagement and signaling pathways underlying the strongly enhancing effects of these HMGB1-partner molecule complexes are presently unknown.
Noncomplexed HMGB1 has been reported to interact and signal via several cell surface receptor systems. The most studied interactions involve the receptor for advanced glycation end products (RAGE), TLR2 and TLR4. RAGE is a promiscuous receptor stimulated by HMGB1 and additional different ligands. RAGE mediates chemotactic and proliferative activities induced by HMGB1 (8), whereas it is an unresolved issue whether HMGB1-RAGE signaling may contribute to cytokine synthesis (13,28). HMGB1 binding to TLR2 and TLR4 results in nuclear factor (NF)-κB activation in different cell types (29). Macrophages totally depend on TLR4 for HMGB1-mediated cytokine production, since TLR4-deficient macrophages do not secrete cytokines after HMGB1 exposure (13). Both TLR4 and TLR2 activation involves an intracellular recruitment of adaptor proteins including myeloid differentiation factor-88 (MyD88) and TIR domain-containing adaptor protein (TIRAP; also termed MAL), which activates signaling cascades, ultimately leading to NF-κB and mitogen-activated protein kinase (MAPK) activation and thereby production of cytokines and other inflammatory molecules. Alternatively, TLR4 may induce cytokine and IFN type 1 synthesis via recruitment of the TIR domain-containing adaptor-inducing IFN-β (TRIF; also termed TICAM-1) and TRIF-related adaptor molecule (TRAM; also termed TICAM-2) (30).
Besides its proinflammatory role, HMGB1 also promotes tissue regeneration, for example, by inducing proliferation and migration of stem cells. These functions are mainly reported to be mediated through RAGE. Thus, to better understand the proinflammatory features of HMGB1 and of HMGB1-partner molecule complexes, it is pivotal to understand which receptors and signaling pathways are involved in and activated by HMGB1 and HMGB1-partner molecule complex stimulation, respectively. Such knowledge also has bearing for the design of therapies targeting the proinflammatory function of HMGB1 in different inflammatory conditions. Whereas many studies have addressed the receptor usage of purified or recombinant HMGB1, little is known about the engagement of receptors recognizing HMGB1-partner molecule complexes, the subsequent intracellular signal transduction pathways and the molecular mechanisms underlying the potentiating features of HMGB1-partner molecule complexes. The present study was aimed at resolving this unsolved issue and to test our hypothesis that one of the HMGB1 receptors (RAGE, TLR2 or TLR4) might function as a coreceptor in a heteromeric, signaling receptor complex and contribute to the enhancing effects of HMGB1 complexes.
Materials and Methods
Reagents
Tag-free recombinant rat HMGB1 free of endotoxin was purchased from HMGBiotech (Milan, Italy). Pam3CSK4 was obtained from InvivoGen (San Diego, CA, USA). Dulbecco’s modified Eagle’s medium (DMEM), Opti-MEM®, phosphate-buffered saline (PBS), penicillin, streptomycin, HEPES, sodium pyruvate and β-mercaptoethanol were from Invitrogen (Carlsbad, CA, USA). LPS extracted from Escherichia coli O55:B5 and purified by ion exchange, as well as fetal calf serum, was purchased from Sigma-Aldrich (St. Louis, MO, USA). Heparin (5,000 international units (IU/mL) was from LEO Pharma A/S (Ballerup, Denmark).
Preparation of Recombinant HMGB1
Recombinant rat HMGB1 with a calmodulin-binding protein tag was expressed in E. coli strain BL21 (for sequence, see Wang et al. [3]). The preparation was first run through a calmodulin affinity column (Calmodulin Sepharose 4B; GE Healthcare, Chalfont St. Giles, U.K.), and the eluate was then further purified with ion exchange chromatography (MonoS 5/50 GL column; GE Healthcare). Protein was stored at −80°C in a buffer containing 50 mmol/L Tris, 4 mmol/L EGTA, 200 mmol/L NaCl and 2.5 mmol/L DTT at pH 8.0. A Toxin Eraser Kit (GenScript, Piscataway, NJ, USA) was used for removal of endotoxins, resulting in endotoxin levels below 6 endotoxin units (EU)/mg protein, as measured by Limulus assay.
Mice
All animal procedures were approved by the Stockholm North Ethical Committee in Sweden. RAGE−/− mice were provided by Bernd Arnold and Angelika Bierhaus (German Cancer Research Center, Heidelberg, Germany) and MyD88−/−, TIRAP−/−, TRAM−/−, TRIF−/−, TLR2−/− and TLR4−/− mice by S Akira (Osaka University, Osaka, Japan). All knockout mice were on a C57Bl/6 background. Wild-type C57Bl/6 mice were purchased from Charles River Laboratories (Wilmington, MA, USA). Animals were kept and bred at the Karolinska Institutet animal facility in a specific pathogen-free environment with standard temperature and a 12-h light/dark cycle. Animals had free access to water and standard rodent chow. Female mice were used in all experiments.
Harvest and Culture of Mouse Peritoneal Macrophages
Wild-type or knockout mice were euthanized by CO2 asphyxiation, and peritoneal macrophages were harvested by lavage of the peritoneal cavity with 5 mL DMEM supplemented with 10% fetal calf serum, 100 units/mL penicillin, 100 µg/mL streptomycin, 10 mmol/L HEPES, 1 mmol/L sodium pyruvate, 50 µmol/L β-mercaptoethanol and 10 IU/mL heparin.
Cells were pelleted by centrifugation at 200g at 4°C for 7 min and resuspended in new medium without heparin. Cells were plated at 100,000 cells per well in a 96-well plate (Sarstedt, Nümbrecht, Germany) and allowed to rest overnight. The next day, the cells were washed to remove nonadherent cells and the medium was changed to OPTIMEM supplemented with penicillin, streptomycin, HEPES, sodium pyruvate and β-mercaptoethanol. Cells were then stimulated for 17–20 h with HMGB1, LPS, Pam3CSK4 or HMGB1 in a preformed complex with LPS or Pam3CSK4. Supernatants were thereafter collected and centrifuged at 600g for 10 min. Cytokine levels in supernatants were either analyzed directly or supernatants were kept at −20°C until analysis. Cell viability was recorded by Trypan blue exclusion (Merck, Darmstadt, Germany), showing no difference in cell viability between different genotypes.
Preparation of HMGB1 Complexes
HMGB1 in PBS, either purchased from HMGBiotech or produced in our laboratory, was mixed together with LPS or Pam3CSK4 in polypropylen tubes in different ratios to give final concentrations in cell cultures, as indicated in the figures. HMGB1 from both sources showed the same stimulatory patterns, both alone and in complexes, and we therefore chose not to distinguish them in the text and figures. The stimulatory ability of the complexes depends on both the temperature and the length of time in which they are formed (14; personal communication, P. Lundback). For these studies, the mixtures were standardly incubated at 4°C for 16 h before addition to cell cultures. Verification of complex formation between HMGB1 and LPS has been demonstrated previously by us and others (14,18).
Cytokine Analysis
IL-6 levels were measured in the supernatants by enzyme-linked immunosorbent assay using hIL-6 DuoSet kits from R&D Systems (Minneapolis, MN, USA) and following the manufacturer’s instructions (detection limit: 15.6 pg/mL).
Statistical Analysis
Cytokine production in peritoneal macrophage culture after stimulation with HMGB1 complexes was assumed to follow Gaussian distribution. A two-tailed Student t test was used to calculate statistical significance between different genotypes. One sample t test was used to calculate statistical significance within groups with the hypothetical value set to 1, corresponding to the LPS/Pam3CSK4 10 ng/mL bar (Figures 1C, D). A P value <0.05 was considered to be statistically significant. Statistical analyses were performed using GraphPad Prism version 5.01 for Windows (GraphPad Software, San Diego, CA, USA).
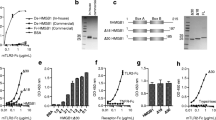
HMGB1 in complex with LPS or Pam3CSK4 induces cytokines via TLR4 and TLR2, respectively, but not via RAGE signaling. Peritoneal macrophages from wild-type, RAGE−/−, TLR2−/− or TLR4−/− mice were stimulated with preformed HMGB1-LPS or HMGB1-Pam3CSK4 complexes, or with individual components separately. Stimulations with 10 µg/mL LPS or Pam3CSK4 were used as positive controls. HMGB1-LPS (A) and HMGB1-Pam3CSK4 (B) complexes both gave enhanced IL-6 production compared with stimulation with the same dose of LPS or Pam3CSK4 alone. (A, B): Results of one representative experiment. Four to five individual experiments were performed with each stimulation. The stimulatory capacity of the complexes (fold change) were calculated by normalizing the raw data in each experiment against 10 ng/mL LPS (n = 5) (C) or Pam3CSK4 (n = 4) (D). All values are expressed as mean ± SD. *P < 0.05, **P < 0.01. ND, not detected; ns, not significant; (A, C)  , unstimulated; □, LPS (10 ng/mL);
, unstimulated; □, LPS (10 ng/mL);  , HMGB1 (1 µg/mL); ■, HMGB1 (1 µg/mL) + LPS (10 ng/mL);
, HMGB1 (1 µg/mL); ■, HMGB1 (1 µg/mL) + LPS (10 ng/mL);  , LPS (1 µg/mL); (B, D)
, LPS (1 µg/mL); (B, D)  , unstimulated; □, Pam3CSK4 (10 ng/mL);
, unstimulated; □, Pam3CSK4 (10 ng/mL);  , HMGB1 (20 µg/mL); ■, HMGB1 (20 µg/mL) + Pam3CSK4 (10 ng/mL);
, HMGB1 (20 µg/mL); ■, HMGB1 (20 µg/mL) + Pam3CSK4 (10 ng/mL);  , Pam3CSK4(10 µg/mL).
, Pam3CSK4(10 µg/mL).
Results
HMGB1 Complexes with TLR Ligands Signal via the Reciprocal TLR of the Partner Ligand
We have previously shown that batches of HMGB1 that induce cell migration but not cytokine production may form complexes with exogenous or endogenous proinflammatory molecules, resulting in strongly enhanced cytokine production compared with the partner ligand alone (14). To elucidate mechanisms mediating this unexplained synergy, we used cultured peritoneal macrophages from mice deficient of RAGE, TLR2 or TLR4 receptors, respectively, and stimulated the cells with HMGB1 complexes or with nonstimulatory HMGB1 alone.
HMGB1 in complex with low amounts of the TLR4 ligand LPS induced significantly enhanced IL-6 production in wildtype cells, compared with stimulation with the individual components alone (Figures 1A, C). These results are in line with our previously published data (14). RAGE−/− and TLR2−/− cells responded equally as well as the wild-type cells to stimulation (Figures 1A, C). In contrast, a lack of TLR4 expression completely abolished the IL-6 production induced by HMGB1-LPS complexes or by LPS alone (Figure 1A).
Stimulation with HMGB1 in complex with a low dose of the synthetic TLR2 ligand Pam3CSK4 generated synergistically increased IL-6 production compared with stimulation with HMGB1 or Pam3CSK4 alone (Figures 1B, D). Cellular depletion of RAGE or TLR4 did not reduce the IL-6 production induced by the HMGB1 complex or by Pam3CSK4 alone compared with that in wild-type macrophages (Figures 1B, D). However, the HMGB1-Pam3CSK4 complex or Pam3CSK4 stimulations were TLR2 dependent, since TLR2−/− cells were unresponsive to these modes of activation.
Taken together, these results demonstrate that HMGB1 in complex with two different TLR ligands signals through the reciprocal receptor of the bound ligand, LPS or PAM3CSK4, to induce cytokines. The results also indicate that the HMGB1-binding receptor RAGE is not required as a coreceptor for the synergy.
Synergistic Effect by HMGB1 Is a Result of Quantitative Rather than Qualitative Changes in Intracellular Signal Pathways
We next asked whether the synergistic enhancement of cytokine production induced by HMGB1 complexes could be a result of changes in the recruitment of intracellular adaptor proteins engaged for intracellular signaling. Peritoneal macrophages from mice lacking individual TLR adaptor molecules including MyD88, TIRAP, TRIF or TRAM were stimulated with HMGB1-LPS or HMGB1-Pam3CSK4 complexes, as well as with the single components. High doses of LPS (1 µg/mL) or Pam3CSK4 (10 µg/mL) were used as positive controls. As demonstrated in Figure 2A, HMGB1-LPS complex stimulation of each of the studied deficient cells led to markedly reduced IL-6 production. TRIF−/−and TRAM−/− cells produced considerably lower levels of IL-6 in response to stimulation with HMGB1-LPS complexes or high-dose LPS compared with wild-type cells. MyD88−/− and TIRAP−/− cells did not produce any or minimal amounts of IL-6 in response to these stimulations, indicating the importance of these adaptor molecules for the signaling of LPS and HMGB1-LPS complexes. When data were normalized to 1 µg/mL LPS, no significant differences were recorded between the different genotypes regarding the relative amounts of IL-6 produced in response to HMGB1-LPS complexes or LPS alone (Figure 3A). This result demonstrates that deficiency of any of the studied adaptor molecules had similar effects on HMGB1-LPS complex stimulation compared with activation with LPS only. MyD88−/− and TIRAP−/− cells expressed equivalent viability and produced comparable amounts of nitric oxide in response to IFN-γ and tumor necrosis factor stimulation as the other genotypes (data not shown), demonstrating that the cells were functional and had the capacity to produce inflammatory mediators in response to stimulation.
Effect of TIRAP MyD88, TRIF and TRAM deficiency on HMGB1 complex stimulation. Peritoneal macrophages from wild-type, MyD88−/−, TIRAP−/−, TRIF−/− or TRAM−/− mice were stimulated with preformed HMGB1-LPS (A) or HMGB1-Pam3CSK4 (B) complexes or individual components separately. Stimulations with 1 µg/mL LPS or 10 µg/mL Pam3CSK4 were used as positive controls. The results of one representative experiment out of three to seven individual experiments performed for each stimulation are shown: wild-type (n = 7), TRAM−/− (n = 3), TIRAP−/− (n = 4), TRIF−/− (n = 3) and MyD88−/− (n = 3). All values are expressed as mean ± SD. ND, not detected; (A)  , unstimulated; □, LPS (1 ng/mL);, HMGB1 (1 µg/mL); ■, HMGB1 (1 µg/mL) + LPS (1 ng/mL);
, unstimulated; □, LPS (1 ng/mL);, HMGB1 (1 µg/mL); ■, HMGB1 (1 µg/mL) + LPS (1 ng/mL);  , LPS (1 µg/mL); (B)
, LPS (1 µg/mL); (B)  , unstimulated; □, Pam3CSK4 (10 ng/mL);
, unstimulated; □, Pam3CSK4 (10 ng/mL);  , HMGB1 (20 µg/mL); ■ HMGB1 (20 µg/mL) + Pam3CSK4 (10 ng/mL);
, HMGB1 (20 µg/mL); ■ HMGB1 (20 µg/mL) + Pam3CSK4 (10 ng/mL);  , Pam3CSK4 (10 µg/mL).
, Pam3CSK4 (10 µg/mL).
Deficiency of TIRAP, MyD88, TRAM or TRIF does not influence the enhancing effect of HMGB1 complexes on IL-6 production. Peritoneal macrophages from wild-type, TRIF−/−, TRAM−/−, TIRAP−/− or MyD88−/− mice were stimulated with preformed HMGB1-LPS or HMGB1-Pam3CSK4 complexes or individual components separately. Stimulations with 1 µg/mL LPS or 10 µg/mL Pam3CSK4 were used as positive controls. Three to seven individual experiments were performed for each stimulation: wild-type (n = 7), TRAM−/− (n = 3), TIRAP−/− (n = 4), TRIF−/− (n = 3) and MyD88−/− (n = 3). Raw data in each experiment were normalized to 1 µg/mL LPS (A) or 10 µg/mL Pam3CSK4 (B) stimulation to generate fold change. All values are expressed as mean ± SD. ND, not detected; ns, not significant. (A) □, LPS (1 ng/mL); ■, HMGB1 (1 µg/mL) + LPS (1 ng/mL); (B) □, Pam3CSK4 (10 ng/mL); ■, HMGB1 (20 µg/mL) + Pam3CSK4 (10 ng/mL).
MyD88 was demonstrated to be an absolute requirement for IL-6 production in response to Pam3CSK4 or HMGB1-Pam3CSK4 complex stimulation, since MyD88 gene deletion totally prevented macrophage activation (Figure 2B). Gene deficiencies for TIRAP, TRIF or TRAM did not influence the relative production of IL-6 in response to Pam3CSK4 versus HMGB1-Pam3CSK4 complexes in comparison to that in wild-type macrophages (Figures 2B and 3B). The observed results in TRIF−/− and TRAM−/− macrophages compared with wild-type cells were to some extent anticipated, since these adaptor proteins do not interact with TLR2 molecules. The results do not support a role for HMGB1 interacting with TLR4 via TRIF/TRAM to enhance the IL-6 synthesis. The results observed in TIRAP−/− cultures demonstrating substantial TLR2-mediated IL-6 production might be explained by the previous demonstration that TIRAP is dispensable in TLR2 signaling at high TLR2 ligand concentrations in macrophage cultures. TLR4 requires the presence of TIRAP to recruit MyD88, whereas TLR2 may dimerize with TLR1 or TLR6 to allow TIRAP-independent recruitment of MyD88 (31).
In summary, although deficiency of the TLR adaptor molecules had quantitative effects in IL-6 production after stimulation with HMGB1 complexes, there were no qualitative effects on the synergistic ability of the HMGB1 complexes when compared with ligand stimulation alone.
Discussion
The common denominator of the experiments with HMGB1-LPS complexes and HMGB1-Pam3CSK4 complexes is that the activity of the complexes relies on engaging the identical cell surface receptors and intracellular adaptor proteins as required for the function of the noncomplexed TLR ligand. No evidence could be demonstrated that the mechanism behind the synergistic effect of HMGB1-TLR ligand complexes is through usage of RAGE, TLR2 or TLR4 as coreceptors. The TLR4 ligand LPS and the HMGB1-LPS complexes signaled via TLR4 without any requirement for TLR2 or RAGE involvement. In comparison, the TLR2 ligand Pam3CSK4 and HMGB1-Pam3CSK4 complexes used TLR2 but not the TLR4 or RAGE pathways. Thus, our working hypothesis that HMGB1-partner molecule complexes used a heteromeric receptor complex appears unlikely. These results agree with our recently published findings that HMGB1 in complex with either IL-1α or IL-1β uses the IL-1 receptor type I, supporting the idea that HMGB1-partner molecule complexes signal through the receptor of the partner molecule (23). We then tested the hypothesis that HMGB1-partner molecule complexes induced an altered signaling pathway compared with the partner molecule alone, and that this could be the mechanism underlying the enhancing effects of HMGB1-partner molecule complexes. When stimulating cells deficient of any of the adaptor molecules used by TLR2 and TLR4 for cytokine induction, similar effects on HMGB1 complex stimulation as on stimulation with noncomplexed LPS or Pam3CSK4 were recorded. We interpret these results of HMGB1-mediated synergy when in complex with TLR ligands to reflect enhanced intracellular signaling without qualitative changes in the signaling pathways. An alternative hypothesis to the initially tested hypothesis, and more likely on the basis of the results of this study, is that the role of HMGB1 is to bind and accumulate multiple TLR ligands, thereby engaging a larger number of TLRs to signal. Alternatively, molecules presented on the surface of the HMGB1 molecule might be more accessible for receptors on target cells. It was recently demonstrated that there are two LPS-binding domains within HMGB1: one in the A box and one in the B box domain (32). Surface plasmon resonance analysis demonstrated that LPS binds to HMGB1 in a concentration-dependent manner and that the binding is stronger to lipid A moiety than to the polysaccharide moiety of LPS (18). The results obtained in TIRAP knockout macrophages using HMGB1-Pam3CSK4 complexes or high-dose Pam3CSK4 stimulation are in line with the idea of a role for HMGB1 to increase the local concentration of the bound partner molecule to activate cells. Low-dose TLR2 ligand requires TIRAP-TLR2 interaction, whereas high TLR2 ligand concentrations are less TIRAP dependent and can be compensated by TIRAP-independent MyD88 recruitment (31). TIRAP−/− macrophages produced substantial amounts of IL-6 after stimulation with a high dose (10 µg/mL) of Pam3CSK4 but minimal quantities in response to a low dose of 10 ng/mL. The cytokine production was strongly increased when HMGB1 was in complex with the low-dose Pam3CSK4 (Figure 2B).
Conclusion
We and other groups have previously established that cytokine production in macrophages stimulated by highly purified, noncomplexed HMGB1 is a totally TLR4-dependent process (13). HMGB1 binds to the TLR4 signaling complex through a mechanism that requires cysteine in position 106 of the HMGB1 molecule to be in the reduced form, expressing a thiol group (13,33). Oxidation of this critical cysteine or exposure of the HMGB1 molecule to dithiothreitol (DTT) denatures the HMGB1-TLR4 interactions and prevents cytokine induction (13,17,33). The present HMGB1-TLR ligand complex experiments were performed with HMGB1 batches exposed to DTT, and they did not bind to TLR4, as revealed by surface plasmon resonance analysis (data not shown). We deliberately designed the experiments to be based on studies of batches of HMGB1 unable to induce cytokines to mimic situations in vivo with non-TLR4-binding forms of HMGB1. Oxidation represents another modification by which HMGB1 is prevented from being able to bind to TLR4, illustrated by the fact that HMGB1 is converted from a proinflammatory to a tolerogenic molecule by oxidation during the apoptotic program of a cell (17). Exposure to oxidative bursts during inflammation will also block HMGB1-TLR4 interactions. The results of our in vitro experiments support the hypothesis that, in an in vivo situation, a pacified version of HMGB1 unable to interact with TLR4 is still capable of promoting and enhancing inflammation by complex formation when present in an environment with microbial TLR ligands. One question not addressed in this study is whether proinflammatory HMGB1 that induces cytokine production on its own would act in a different way in HMGB1-TLR ligand complexes than noninflammatory HMGB1. We know from previous experiments that proinflammatory HMGB1 when in complex with LPS or Pam3CSK4 will also potentiate cytokine production in macrophages, but we have no parallel experiments to comment on quantitative aspects compared with complexes with noninflammatory HMGB1.
The biologically important message in the present study is that endogenously noninflammatory HMGB1 molecules may function as alarm signals and promote cytokine formation in a milieu with low amounts of TLR ligands present. The ability of HMGB1 in complex with low amounts of TLR ligands to induce or strongly enhance a response to low concentrations of TLR ligands might be an important sensory mechanism in the induction and prolongation of an inflammatory response. Complexes on the basis of HMGB1 and TLR ligands and “free” TLR ligands use identical receptor systems to induce cytokine production.
Disclosure
The authors declare that they have no competing interests as defined by Molecular Medicine, or other interests that might be perceived to influence the results and discussion reported in this paper.
References
Andersson U, Tracey KJ. (2011) HMGB1 is a therapeutic target for sterile inflammation and infection. Annu. Rev. Immunol. 29:139–62.
Tang D, Kang R, Zeh HJ 3rd, Lotze MT. (2011) High-mobility group box 1, oxidative stress, and disease. Antioxid. Redox. Signal. 14:1315–35.
Wang H, et al. (1999) HMG-1 as a late mediator of endotoxin lethality in mice. Science. 285:248–51.
Semino C, Angelini G, Poggi A, Rubartelli A. (2005) NK/iDC interaction results in IL-18 secretion by DCs at the synaptic cleft followed by NK cell activation and release of the DC maturation factor HMGB1. Blood. 106:609–16.
Dumitriu IE, et al. (2005) Release of high mobility group box 1 by dendritic cells controls T cell activation via the receptor for advanced glycation end products. J. Immunol. 174:7506–15.
Kawahara K, et al. (2007) HMGB1 release in cocultures of porcine endothelial and human T cells. Xenotransplantation. 14:636–41.
Scaffidi P, Misteli T, Bianchi ME. (2002) Release of chromatin protein HMGB1 by necrotic cells triggers inflammation. Nature. 418:191–5.
Degryse B, et al. (2001) The high mobility group (HMG) boxes of the nuclear protein HMG1 induce chemotaxis and cytoskeleton reorganization in rat smooth muscle cells. J. Cell. Biol. 152:1197–206.
Goldstein RS, et al. (2006) Elevated high-mobility group box 1 levels in patients with cerebral and myocardial ischemia. Shock. 25:571–4.
van Zoelen MA, et al. (2007) Systemic and local high mobility group box 1 concentrations during severe infection. Crit. Care Med. 35:2799–804.
Yang R, et al. (2006) Anti-HMGB1 neutralizing antibody ameliorates gut barrier dysfunction and improves survival after hemorrhagic shock. Mol. Med. 12:105–14.
Andersson U, et al. (2000) High mobility group 1 protein (HMG-1) stimulates proinflammatory cytokine synthesis in human monocytes. J. Exp. Med. 192:565–70.
Yang H, et al. (2010) A critical cysteine is required for HMGB1 binding to Toll-like receptor 4 and activation of macrophage cytokine release. Proc. Natl. Acad. Sci. U. S. A. 107:11942–7.
Hreggvidsdottir HS, et al. (2009) The alarmin HMGB1 acts in synergy with endogenous and exogenous danger signals to promote inflammation. J. Leukoc. Biol. 86:655–62.
Rouhiainen A, Tumova S, Valmu L, Kalkkinen N, Rauvala H. (2007) Pivotal advance: analysis of proinflammatory activity of highly purified eukaryotic recombinant HMGB1 (amphoterin). J. Leukoc. Biol. 81:49–58.
Antoine DJ, Williams DP, Kipar A, Laverty H, Park BK. (2010) Diet restriction inhibits apoptosis and HMGB1 oxidation and promotes inflammatory cell recruitment during acetaminophen hepatotoxicity. Mol. Med. 16:479–90.
Kazama H, et al. (2008) Induction of immunological tolerance by apoptotic cells requires caspase-dependent oxidation of high-mobility group box-1 protein. Immunity. 29:21–32.
Youn JH, Oh YJ, Kim ES, Choi JE, Shin JS. (2008) High mobility group box 1 protein binding to lipopolysaccharide facilitates transfer of lipopolysaccharide to CD14 and enhances lipopolysaccharide-mediated TNF-alpha production in human monocytes. J. Immunol. 180:5067–74.
Ivanov S, et al. (2007) A novel role for HMGB1 in TLR9-mediated inflammatory responses to CpG-DNA. Blood. 110:1970–81.
Tian J, et al. (2007) Toll-like receptor 9-dependent activation by DNA-containing immune complexes is mediated by HMGB1 and RAGE. Nat. Immunol. 8:487–96.
Qin YH, et al. (2009) HMGB1 enhances the proinflammatory activity of lipopolysaccharide by promoting the phosphorylation of MAPK p38 through receptor for advanced glycation end products. J. Immunol. 183:6244–50.
Sha Y, Zmijewski J, Xu Z, Abraham E. (2008) HMGB1 develops enhanced proinflammatory activity by binding to cytokines. J. Immunol. 180:2531–7.
Wahamaa H, et al. (2011) High mobility group box protein 1 in complex with lipopolysaccharide or IL-1 promotes an increased inflammatory phenotype in synovial fibroblasts. Arthritis Res. Ther. 13:R136.
Campana L, Bosurgi L, Bianchi ME, Manfredi AA, Rovere-Querini P. (2009) Requirement of HMGB1 for stromal cell-derived factor-1/CXCL12-dependent migration of macrophages and dendritic cells. J. Leukoc. Biol. 86:609–15.
Urbonaviciute V, et al. (2008) Induction of inflammatory and immune responses by HMGB1-nucleosome complexes: implications for the pathogenesis of SLE. J. Exp. Med. 205:3007–18.
Yanai H, et al. (2009) HMGB proteins function as universal sentinels for nucleic-acid-mediated innate immune responses. Nature. 462:99–103.
Yanai H, et al. (2011) Suppression of immune responses by nonimmunogenic oligodeoxynucleotides with high affinity for high-mobility group box proteins (HMGBs). Proc. Natl. Acad. Sci. U. S. A. 108:11542–7.
Kokkola R, et al. (2005) RAGE is the major receptor for the proinflammatory activity of HMGB1 in rodent macrophages. Scand. J. Immunol. 61:1–9.
Park JS, et al. (2006) High mobility group box 1 protein interacts with multiple Toll-like receptors. Am. J. Physiol. Cell. Physiol. 290:C917–24.
Kawai T, Akira S. (2010) The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat. Immunol. 11:373–84.
Kenny EF, et al. (2009) MyD88 adaptor-like is not essential for TLR2 signaling and inhibits signaling by TLR3. J. Immunol. 183:3642–51.
Youn JH, et al. (2011) Identification of lipopolysaccharide-binding peptide regions within HMGB1 and their effects on subclinical endotoxemia in a mouse model. Eur. J. Immunol. 41:2753–62.
Antoine DJ, Williams DP, Kipar A, Laverty H, Park BK. (2010) Diet restriction inhibits apoptosis and HMGB1 oxidation and promotes inflammatory cell recruitment during acetaminophen hepatotoxicity. Mol. Med. 16:479–90.
Acknowledgments
This study was supported by the Swedish Research Council, the Swedish Rheumatism Association, the King Gustaf V foundation, the Swedish Cancer Society, the Von Kantzow Foundation and the European Commission (AtheroRemo collaborative project).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it.
The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this license, visit (https://doi.org/creativecommons.org/licenses/by-nc-nd/4.0/)
About this article
Cite this article
Hreggvidsdóttir, H.S., Lundberg, A.M., Aveberger, AC. et al. High Mobility Group Box Protein 1 (HMGB1)-Partner Molecule Complexes Enhance Cytokine Production by Signaling Through the Partner Molecule Receptor. Mol Med 18, 224–230 (2012). https://doi.org/10.2119/molmed.2011.00327
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2119/molmed.2011.00327