Abstract
Multiple lines of evidence suggest that inhibition of Type I Interferons, including IFN-α, may provide a therapeutic benefit for autoimmune diseases. Using a chemical genomics approach integrated with cellular and in vivo assays, we screened a small compound library to identify modulators of IFN-α biological effects. A genomic fingerprint was developed from both ex vivo patient genomic information and in vitro gene modulation from IFN-α cell-based stimulation. A high throughput genomic-based screen then was applied to prioritize 268 small molecule inhibitors targeting 41 different intracellular signaling pathways. Active compounds were profiled further for their ability to inhibit the activation and differentiation of human monocytes using disease-related stimuli. Inhibitors targeting NF-κB or Janus Kinase/Signal Transducer and Activator of Transcription (JAK/STAT) signaling emerged as “dissociated inhibitors” because they did not modulate IFN-α anti-viral effects against HSV-1 but potently inhibited other immune-related functions. This work describes a novel strategy to identify small molecule inhibitors for the treatment of autoimmune disorders.
Similar content being viewed by others
Introduction
Systemic Lupus Erythematosus (SLE) is a prototypic systemic autoimmune disorder that is characterized by anti-nuclear autoantibodies and the presence of inflammatory lesions targeting a variety of tissues including the skin, joints, brain, heart, lung, and kidney (1). Development of the disease can lead to deposition of immune complexes in the kidney, renal failure, and death. SLE is diagnosed mostly in woman during childbearing years and affects approximately five million people worldwide. Therapies have remained essentially unchanged for over 20 years and still rely largely on undesirable long term use of corticosteroids and immunosuppressive drugs to slow disease progression. The need for safe, new, effective therapies is urgently required. Recently, it has emerged that type I Interferons (IFN) play a prominent role in the pathogenesis of lupus, however, type I IFNs also play an important role in host defense against viral infection (2). Therefore, we are faced with the challenge of developing a screening strategy that identifies drugs that inhibit the pro-inflammatory response of type I IFNs while retaining protection from viral infection.
Type I IFNs are a family of pleiotropic cytokines that play an important role in modulating nearly all phases of immune and inflammatory responses (2,3). Type I IFNs include 13 functional IFN-α genes, and single IFN-β, IFN-ε, IFN-κ, and IFN-ω genes (3). Binding of type I IFNs to a common receptor (IFNAR) composed of a unique IFNAR1 subunit and a functionally active IFNAR2c subunit, results in the activation of JAK1 and TYK2 kinases that subsequently activate the signal transducer and activator of transcription (STAT) proteins 1, 2, 3, 4, and 5, and regulate the expression of hundreds of interferon-stimulated genes (ISGs) (4–6).
The connection between type I IFNs (particularly IFN-α) and SLE is compelling (7). Type I IFN-regulated genes are overexpressed significantly in PBMCs from SLE patients (8–12), and elevated levels of IFN-α activity correlate with both disease activity and severity (10,13). Moreover, the observation that patients with non-autoimmune disorders who are treated with recombinant IFN-α can develop antinuclear antibodies, anti-dsDNA antibodies, and, occasionally, SLE, indicates that IFN-α plays a direct role in the pathogenesis of SLE (14). In vivo models of autoimmune disease also show that the administration of exogenous IFN-α induces glomerulonephritis in normal mice and accelerates the onset of the spontaneous autoimmune disease of NZB/W mice (15). Furthermore, autoimmune-predisposed mice deficient in the IFN-α/β receptor exhibit significantly reduced anti-erythrocyte auto-antibodies, hemolytic anemia, anti-DNA autoantibody, kidney disease, and mortality (16). Altogether, these data strongly indicate that targeting the IFN-α pathway may provide an effective approach for the treatment of SLE. Moreover, this approach may also be applicable to other autoimmune disorders associated with dysregulation of type I IFN-signaling pathways such as psoriasis, type I diabetes, Sjögren’s disease, and inflammatory myopathies (17).
Therapeutic modulation across the spectrum of type I IFN-pathways represents a novel and promising approach which represents a challenge to the conventional single target drug discovery. Recent advances in molecular biology, robotics, and assay detection technologies make it feasible to explore gene, protein, and signaling pathways in an integrated cellular context (18,19). Molecular profiling by these approaches has several potential advantages, both as a primary anchor to drug discovery and as a complement to more conventional target-based discovery efforts. The use of large complex sets of genomic bio-markers already has found its way into standard use in the identification and validation of drug targets (18,19). Profiling the expression of large gene sets in normal, compared with disease, states can provide critical clues to the activities of cellular control pathways as well as identifying specific gene signatures as the surrogate markers in disease processes. An exciting use of such molecular surrogate markers that has the potential to revolutionize drug discovery is its utility in defining cellular states as the primary driver for the identification of drug candidates (20–22).
Here, we illustrate a robust and novel gene expression platform based on high-throughput integrated transcriptional screening (HITS) followed by secondary biological assays to identify small molecular compounds that normalize the perturbed PBMC gene signatures of SLE patients. A library of 268 well-annotated small molecule inhibitors spanning 41 mechanism of actions (MOAs) that inhibit or modulate well-defined signaling pathways were screened. We found that inhibitors targeting either NF-κb or JAK/STAT signaling were able to block IFN-α-mediated biological activities that contribute to the pathogenesis of SLE without modulating the IFN-α-dependent anti-viral response to Herpes simplex virus type 1(HSV-1). Our results indicate that small molecules targeting JAK/STAT and NF-κb pathways are potential drug candidates for SLE or IFN-α-associated autoimmune disease.
Materials and Methods
Genome-Wide Gene Expression Analysis
Human U133A microarrays (Affymetrix Inc. Santa Clara, CA, USA) were used to profile transcriptional changes in THP-1 cells stimulated with cytokines. THP-1 cells were treated with 100 IU/mL IFN-α, 10 ng/mL IFN-α, 10 ng/mL TNF-α, or vehicle-only control for 4 h. Total RNA was isolated in Trizol reagent (Invitrogen Inc, Carlsbad, CA, USA) and purified on RNeasy plate (Qiagen Inc, Valencia, CA, USA). The purified total RNA from eight biological replicates for IFN-α and IFN-γ treatments, six replicates of TNF-α treatments, and 14 replicates of vehicle-only controls were processed and hybridized on HT_HG-U133A high-throughput 96 well array plates according to Affymetrix high-throughput array platform protocols provided by the microarray supplier. All the stimulated gene expression data sets were normalized to the vehicle control treatments for the pathway gene marker set analysis. Raw data are available upon request.
High-Throughput Integrated Transcriptional Screening
THP-1 cells were treated with either compound (10 µM final concentration) or vehicle control (0.1% DMSO) for 30 min prior to a 4 h stimulation with 100 IU/mL IFN-α or PBS. Plates were incubated at 37° C in a humidified incubator. Cell lysis and RNA isolation were carried out according to the manufacturer’s instructions (Qiagen Inc.). Real time polymerase chain reactions (PCRs) were carried out in SYBR green master mix (Quanta Biosciences, Gaithersburg, MD, USA) on ABI9700 thermocyclers.
Oligo Pairs Used for RT-PCR
IFI35 (Fwd: 5′ ggagtggctcagcgtctgt 3′ and Rev: 5′ actggctgcgacctgatct 3′), OAS3 (Fwd: 5′ tgaaggctgctgtgtgaagt 3′ and Rev: 5′ cacacacacatgtacacaatctctc 3′), G1P2 (Fwd: 5′ cgaactcatctttgccagtaca 3′ and Rev: 5′ gagctgctcagggacacc 3′), RSAD2 (Fwd: 5′ tggatagcaaatcctgagacaat 3′ and Rev: 5′ cctgtgtgtattccttctttttagc 3′), HNRPA0 (Fwd: 5′ Gggtgggttcagagtacctttt 3′ and Rev: 5′ gcttcttagtatagctttgagccttc 3′), DDX58 (Fwd: 5′ Tgaactgtaagggttagtggagagt 3′ and Rev: 5′ aaataatccatttgtattgggtct 3′), MX1 (Fwd: 5′ Ggacatcactgctctcatgc 3′ and Rev: 5′ tttatggccttcttgaaaattg 3′).
HITS Scoring Model
We used the housekeeping gene, GAPDH, in the HITS as the normalization control for the IFN-α gene signature set to correct the overall variability in the qPCR-based HITS process. The corrected profiles then were normalized to the basal gene expression levels determined by using the vehicle-only treatments. We used the SNR statistics as the weight function to adjust the contribution of each signature gene to the HITS score based on the reliability of the gene expression measurements (20). Hits were the tested compounds where HITS score > cut-off HITS score as established by the vehicle controls to give false discovery rate < 0.05.
Human PBMC Stimulation with IFN-α or Patient Serum
SLE patient and control serum were purchased from Bioreclamation (Hicksville, NY, USA). IFN-α 2a was from PBL Biomedical Laboratories (Piscataway, NJ, USA). Fresh PBMCs from healthy donors were prepared by Ficollhypaque fraction according to the manufacturer’s instructions. Cells were cultured at 2 × 105 cells/0.1 ml in 96-well flat-bottomed plates in a culture medium. To study the effect of compounds on gene expression, compounds and vehicle controls were pre-incubated with cells for 30 min at 37° C before stimulation with 50% lupus serum, or 100 IU/mL IFN-α 2a were incubated with PBMC. After 6 h stimulation, cells were lysed in Qiagen 2XTCL lysis buffer followed by HITS analysis.
In Vitro Anti-Viral Assay
Hep-2 cells were propagated in MEM (Invitrogen) with 10% fetal bovine serum. HSV-1 recombination virus with both firefly and Renilla luciferase genes in a divergent orientation from a single multiple cloning site were used. Hep-2 cells were seeded at 2 × 104 cells/well in a 96-well plate. Twenty-four h post-seeding, cells were infected with virus at a multiplicity of infection (MOI) of 5. After absorption for 1 h at 37° C, free viral particles were removed by aspiration, cells were washed, and 200 µl of medium containing IFN-α with or without test compounds. After 48 h incubation, cells were washed and lysed in 20 µl of passive lysis buffer (Promega Inc, Madison, WI, USA) frozen, thawed, and assayed for firefly and Renilla luciferase activity using the Dual Luciferase Assay kit (Promega).
IFN-α-Induced Chemokine Release In Vivo
Female NZBW/F1 mice (Jackson Laboratories, Bar Harbor, ME, USA) were housed in pathogen-free conditions with access to food and water. At 11 wks of age, mice were randomized and placed into control (n = 4 to 5) and treatment groups (n = 6). IKK2 inhibitor IV was administered in 0.9% DMSO, 7% dimethylacetoacetamide (Aldrich Chemical Co, Milwaukee, WI, USA) and 10% Cremophor EL (Sigma-Aldrich, St. Louis, MO, USA) 2 h prior to stimulation. Antimouse IFN-α receptor antibodies (5A3) or mouse IgG1 isotype control antibodies were administered 16 h prior to stimulation as a positive control. Mice were stimulated by adenovirus delivery of IFN-α5, 1 × 1010 viral particles were injected intravenously. Six h post-stimulation, the mice were killed by CO2 exposure and blood samples were collected by cardiac puncture. Serum levels of IP10 were assessed by Enzyme-Linked ImmunoSorbent Assay (ELISA) (R&D Systems, Minneapolis, MN, USA).
Statistical Analysis
Statistical comparison between two groups was conducted by student t-tests.
Results
Establishing a Chemical Genomics-Based Platform to Identify Specific Inhibitors of the IFN-α Pathway
To develop a robust and reproducible genomic-based high throughput screen, we carried out genome-wide differential gene expression analysis using a robust and reproducible in vitro platform. A human monocytic cell line, THP1 cells, was used to characterize the genomic signature of relevant pro-inflammatory cytokines. A total of 302 genes were identified that either were activated or repressed more than 1.4-fold by the IFN-α stimulation relative to the unstimulated cellular state at false discovery rate (FDR) < 0.01. Because IFN-α, IFN-γ, and TNF-α share a large set of overlapping signaling pathways, we next counterselected against genes that were significantly modulated by IFN-γ or TNF-α stimulation. That left a total of 76 genes uniquely modulated by IFN-α. The 25 genes displaying the highest degree of modulation were subsequently selected for qPCR-based HITS assay evaluation and development (Figure 1A). Seven genes showing the best correlation and most robust activation (MX1, OAS3, DDX58, RSAD2, G1P2, IFI35) or repression (HNRP0) upon IFN-α stimulation were selected. These genes were used in our HITS assay for screening modulators of the IFN-α signaling pathway. A modified weighted voting model based on the SNR method was established to score the active compounds (20) (Figure 1B).
Establishing a chemical genomics-based platform to identify specific inhibitors of the IFN-α pathway. The unique gene expression signature of the IFN-α signaling pathway was established in THP-1 cells by comparing the gene profiles of the fully activated IFN-α pathway to the basal gene expression. THP-1 cells were stimulated either with 100 IU/mL IFN-α, 100 IU/mL IFN-γ, or 10 ng/mL TNF-α to generate the activated pathway gene signatures. (A) Heat-map representation of the 25 genes selected for HITS (high-throughput integrated transcriptional screening). Columns represent cytokine stimulation treatments and rows represent genes (red: higher expression; green: lower expression). The significance values were computed across the replicates of each IFN-α stimulation compared to the vehicle-only treatments using student t-tests. (B) The six most reproducibly upregulated genes, one downregulated gene, and a normalization control gene (GAPDH) were validated and run in the qPCR-based HITS assay. This set of genes was used to screen for inhibitors of IFN-α signaling pathway. Shown is a typical expression profile of the gene set following IFN-α treatment. The HITS scoring weight and halfmax threshold are indicated.
Stimulation of healthy donor PBMCs with serum isolated from SLE patients induces the upregulation of IFN-α pathway-associated genes (Interferoninduced genes, IFIGs), such as MX1 (23). Furthermore, expression of IFIG correlates with disease severity and organ involvement (23,24). We also confirmed that the IFN-α pathway gene signature set was relevant to the disease state of SLE patients. Healthy donor PBMCs were stimulated with either SLE serum or healthy donor serum. Significant induction of all six selected upregulated IFN-α pathway signature genes was observed (Figure 2). The induction ranged from five-fold (G1P2) to a 40-fold median increase (OAS3) depending on the gene. Our results are consistent with the marked induction pattern of type I IFN-inducible genes observed ex vivo with SLE samples (8–12). These data further support the therapeutic relevance of our genomic screening platform and demonstrate that IFN-α is an important contributor of the SLE serum-induced gene signature.
HITS platform identifies SLE disease state. Human healthy donor PBMCs were stimulated with 50% serum isolated from either SLE patients or healthy donors. Following a 4 h treatment incubation, cells were lysed, and RNA was prepared for qPCR analysis using Avalonrx-HITS assay. Relative expression of six IFN-α upregulated genes (MX1, OAS3, DDX58, RSAD2, G1P2, and IFI35) is shown in A to F respectively. Each dot represents an individual sample (n= 38 for SLE samples, n= 11 for healthy donor samples). Horizontal lines show median gene expression values normalized to GAPDH. Statistic comparison between the two groups was conducted by simple student t-tests.
High-Throughput Integrated Transcriptional Screening (HITS)
HITS assays then were carried out for screening of 268 target-specific compounds. The screen consists of THP1 cells stimulated with 100 IU/mL IFN-α for 4 h. A desirable hit would reverse the seven-gene signature back toward basal gene expression levels. Vehicle-only treatments were used to establish baseline gene expression, and treatment with 100 IU/mL IFN-α was used to establish the maximal gene expression levels. Genes whose expression was neutralized to at least 50% of maximal levels were used in a modified weighted voting model based on the SNR statistics to score the compound treatments. We used the HITS scores from both vehicle-only and vehicle with 100 IU/mL IFN-α treated THP-1 cells to establish the confidence interval of the HITS calling model. Any compound consistently scoring positive at FDR < 0.05 across the multiple runs was classified as an active compound. The HITS screen identified 30 compounds from eight mechanisms of action (MOA) groups (data not shown). Compounds with undesirable MOA and cytotoxicity were excluded. Representative compounds from three groups, Apicidin 1a from the HDAC inhibitor group, IKK2 inhibitor IV from the NF-κb inhibitor group, and JAK inhibitor I, a direct inhibitor of the JAK/STATs pathway, were selected for further validation. Dose titrations then were performed in the same HITS assays. We observed a dose-dependent inhibition of the IFN-α pathway signatures. The TI50 values, defined as the dosage that inhibited 50% of the IFN-α stimulation gene expression profile, were determined for all three compounds. TI50 of JAK inhibitor I is 0.3 µM, TI50 of IKK2 inhibitor IV is 0.6 µM, and TI50 of Apicidin 1a is 0.2 µM. It is important to note that there was no general cellular toxicity observed in the THP-1 cells when treated with up to 1 µM of these compounds.
Selected Compounds Inhibit SLE-Associated Gene Signatures
To further evaluate the role of small molecular inhibitors on the type I IFN-gene signature, freshly isolated PBMC stimulated with 50% lupus serum were used in HITS assays. As shown in Figure 3, Apicidin 1a, IKK2 inhibitor IV, and JAK inhibitors I significantly blocked the upregulation of the six most robustly induced IFN-α signature gene set in a dose-dependent manner. Apicidin 1a, IKK-2 inhibitor IV, and JAK inhibitor I showed 80%, 77%, and 60% inhibition, respectively. No cytotoxity was apparent at the test compound concentration of 1 µM. Importantly, these experiments were consistent with SLE serums from patients with different level of IFN-α activity and autoantibody profile (data not shown). These data suggest that JAK inhibitor I, IKK-2 inhibitor IV, and Apicidin 1a are effective inhibitors of the IFN-α gene signature induced by SLE serum. Because the biological activity of SLE serum has been associated with pathogenesis, our results suggest that small molecule inhibitors targeting HDAC, NF-κb, and JAK/STAT signaling pathways could modulate SLE disease activity.
Effect of small molecule inhibitors on SLE-associated gene signature. JAK inhibitor I (A), IKK2 inhibitor IV (B), or Apicidin 1a (C) were preincubated with freshly isolated monocytes from healthy donors for 30 min at the concentration indicated before the cells were stimulated with 50% SLE patient serum for 4 h. RNA from lysed cells was examined by qPCR analysis in HITS assay. Percentage inhibition represents the mean inhibition of the six most robustly induced IFN-α signature genes compared with the vehicle control. Each bar represents the average ± SEM of three independent experiments using serum from different SLE patients. *P < 0.05, **P < 0.01 when compared with the vehicle control.
Effect of Apicidin 1a, IKK2 Inhibitor IV, and JAK Inhibitors in IP-10 and MCP-1 Expression Induced by IFN-α
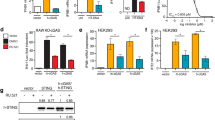
Multiple chemokines, including monocyte chemo-attractant protein-1 (MCP-1) and activated T cell chemokine interferon-inducible protein 10 (IP-10) regulate leukocytes migration and infiltration into inflamed organs (25). Expression of MCP-1 and IP-10 are elevated in the serum of SLE patients (25,26), and in the monocytes of healthy donors stimulated in vitro by IFN-α (27). Consequently, the effect of Apicidin 1a, IKK2 inhibitor IV, and JAK inhibitors in IP-10 and MCP-1 expression induced by IFN-α from human monocytes was examined. As indicated in Figures 4A and 4B, doses as low as 0.03 µM of JAK inhibitor I and IKK-2 inhibitor IV blocked the expression of IP-10 induced by IFN-α significantly (P < 0.05). MCP-1 expression required higher doses of JAK inhibitor I and of IKK-2 inhibitor IV. In contrast, 1 µM Apicidin 1a treatment neutralized MCP-1 expression induction totally, whereas IP-10 expression was unaffected. Furthermore, treatment of JAK inhibitor I and of IKK-2 inhibitor IV resulted in a dose-dependent inhibition of monocyte differentiation marker, such as CD38, CD80, and CD123 (data not shown).
Effect of inhibitors on in vitro and in vivo chemokine release. Freshly isolated monocytes were treated with 500 IU/mL IFN-α with or without indicated compounds for 48 h. Supernatants were collected and MCP-1 and IP-10 levels were analyzed by Searchlight Inc. Percentage inhibition represents inhibition of the upregulation of chemokine induced by IFN-α compared with the vehicle control. (A) Effect of compounds on IP-10 secretion. (B) Effect of compounds on MCP-1 secretion. Each bar represents the average ± SEM of triplicates from a single experiment. Data representative of three independent experiments. *P < 0.05, when compared with the vehicle control. (C) IKK2 inhibitor IV blocked IFN-α-induced IP-10 protein level in vivo. Female NZBW/F1 mice were pre-treated with either IgG control or anti-INFR antibody (intraperitoneal injection 30 mg/kg) or IKK2 inhibitor or vehicle control (intraperitoneal injection 45 mg/kg BID) as described in Materials and Methods. Adenovirus-IFN-a (1 × 1010 viral particles) was then administrated intravenously at time 0. Samples were collected 6 h post-stimulation, IP-10 protein level was measured by enzyme-linked immunosorbent assay (ELISA). Each bar indicated mean ± SEM from one experiment. **P < 0.01, ***P < 0.001. Data representative of two experiments.
Based on the results from in vitro assays, the IKK-2 inhibitor IV was examined in vivo for its capacity to inhibit IP-10 expression induced by IFN-α. The serum level of IP-10 was elevated after mice were infected with adenovirusencoding IFN-α5. Treatment of IKK-2 inhibitor IV and a surrogate mouse antiinterferon receptor antibody inhibited serum level of IP-10 by 98% relative to control (P < 0.001, P < 0.01, respectively, Figure 4C). These observations illustrate the robustness of our strategy for identifying small molecule inhibitors with desirable immunosuppressive effect.
Impact of Apicidin 1a, IKK2 Inhibitor IV, and JAK Inhibitors in IFN-α-Regulated HSV-1 Replication
Herpes simplex virus-1 (HSV-1) represents one of the major recurrent virus infections observed in SLE patients (28). Type-I and Type-II IFN signals are known to block HSV-1 dissemination in mice (29), and, as a consequence, a therapeutic approach that neutralizes their combined activity may constitute an important safety concern. Therefore, the impact of Apicidin 1a, IKK2 inhibitor IV, and JAK inhibitors on HSV-1 replication regulated by IFN-α in Hep-2 cells was examined in vitro. HSV-1/luciferase was used to infect Hep-2 cells, and viral replication was monitored by luciferase expression. We first confirmed that reporter gene activity rose concomitantly and proportionally with the detection of viral progeny (data not shown).
As shown in Figure 5, in absence of IFN-α, luciferase expression indicates high levels of HSV-1 replication. IFN-α treatment significantly reduced viral replication. Both the JAK1 and IKK2 inhibitors retained the majority of IFN-α-dependent anti-viral activity (Figure 5A and 5B) even at doses as high as 1 µM. At this concentration, the JAK1 and IKK2 inhibitors significantly inhibited IFN-a gene signatures (Figure 3), monocyte activation (data not shown), and chemokine production (Figure 4). In contrast, Apicidin 1a inhibited the anti-viral effects of IFN-α at a low dose (0.1 µM), but at a higher dose, where this drug effectively inhibited MCP-1 production, IFN-α-dependent anti-viral activity was abolished or viral growth actually was promoted (Figure 5C). Based on these results, both JAK I and IKK-2 IV inhibitors emerge as possible lead candidates with desirable immunosuppressive and dissociated anti-viral effects.
Dissociated anti-viral properties of lead candidates. Hep-2 cells were seeded at 2 × 104 cells/well in 96-well plates. Cells were infected with HVS1 at a multiplicity of infection (MOI) of five for 1 h at 37° C 24 h post-seeding. Virus was aspirated, cells were washed, and 200 µl of medium with or without IFN-α or compound were added as indicated. After 48 h, cells were washed, lysed, and luciferase activity was measured according to the manufacturer’s instructions. Three independent experiments were performed with similar results. Each bar represents the mean ± SEM of triplicates from one experiment. *P < 0.05, **P < .01, ***P < 0.001 when compared with virus alone.
Discussion
The protective role of type I IFNs in viral infections has been well established. More recently, type I IFNs have been clearly indicated in the pathogenesis of lupus and other autoimmune diseases (7,14). One key goal is to identify a small molecule inhibitor that blocks type I IFN-mediated biological activity effectively, and that is responsible for the pathogenesis of autoimmune disease without compromising IFN-dependent anti-viral activity. We describe here an integrated chemical genomics-based drug discovery approach and its application in the screening for compounds that reverse a gene signature associated with the activation of the IFN-α pathway. Using these chemical probes, we identified the signaling nodes or the cross-talk pathways that can modulate the IFN-α responses. Compounds targeting HDAC, JAK/STAT, and NF-κb pathways inhibited IFN-α responses. However only compounds targeting JAK/STAT and NF-κb inhibited IFN-α without markedly compromising anti-viral responses. Potentially, compounds targeting these pathways could be useful therapeutically for patients with SLE and other autoimmune conditions with INF-α involvement.
The major signaling pathway activated by type I IFNs involves sequential phosphorylation of the tyrosine residues of the JAK and STAT proteins, however, more and more evidence demonstrates that JAK-STAT signaling alone is not sufficient to explain all the biological effects of type I IFNs. The PI3k and p38 kinase pathways have emerged as critical additional components of IFN-induced signal transduction (6). There also is emerging evidence that modulation of the function of a distinct STAT protein might account for a specific response. For example, a recent report showed that IFN-α and IFN-β-mediated activation of STAT 4 is required for IFN-γ production during viral infection (30). However, STAT1 negatively regulates IFN-α dependent induction of IFN-γ (31). It is not surprising that JAK inhibitors were identified in our assay and demonstrate strong inhibitory potency toward the IFN gene signature. However, despite the complexity of the IFN system, we are able to identify a concentration that effectively inhibits IFN-α related biological activity that contributes to the pathogenesis of the disease and yet retains IFN-α-dependent anti-viral activity. These data suggest that developing drugs that target JAK/STAT signaling is an attractive direction for the treatment of autoimmune disease.
Besides STAT proteins, type I IFNs also activate other transcription factors. Among them, NF-κb is the most important transcription factor activated by IFNs. The key regulator of NF-κb is the signalsome, which comprises the scaffold protein NEMO and the two kinase inhibitors of NF-κb, Iκb kinase (IKK1) and IKK2. IKK2 is particularly important because it phosphorylates the NF-κb inhibitor Iκb, which is subsequently ubiquitinated by the SCFβtrcp ligase system, leading to the degradation of the kinase and activation of p50–p65 dimer (32). In addition to this major pathway for the p50–p65 activation, there is an alternative NF-κb pathway, again involving the IKKs, but leading to the activation of two other NF-κb proteins, p100 and RELB (32). NF-κb positively and negatively regulates IFN-induced gene expression as well as antiviral activity. A recent report showed that NF-κb positively induced antiviral activity toward VSV (33), whereas another report suggests that NF-κb suppressed both antiviral and immunomodulatory actions of IFN against the influenza virus (34). The data presented here indicate that IKK2 inhibitors exhibit only small effects on IFN-dependent anti-HSV-1 activity, which is consistent with a previous observation that efficient replication of HSV-1 involves activation of the NF-κb pathway (35). Interestingly, the IKK2 inhibitor that was identified in our assay has been shown to block inflammation in human airway smooth muscle and in a rat model of asthma (36). Taken together, inhibitors of the NF-κb signaling pathway may present attractive approaches for the treatment of autoimmune disease.
The requirement for HDAC as positive regulators of IFN-α- and cytokine-induced gene expression has been well established. The deacetylase protein HDAC1 can interact with both the STAT1 and STAT 2 subunits of ISGF3. Whilst the inhibition of deacetylase activity has no effect on IFN-α signaling that leads to STAT phosphorylation, nuclear translocation, the assembly of the ISGF3 or the ISGF3 DNA binding, inhibition of HDAC does target downstream events required for IFN-stimulated gene expression (37). All these data support a model that deacetylase enzyme may serve as a transcriptional coactivator for ISGF3. In addition, the IFN anti-viral response also requires HDAC activity. The anti-viral response against HCV, EMCV, and VSV were impaired in the presence of HDAC inhibitors (38). In fact, treatment with HDAC inhibitors increased the viral cytopathic activity, most likely through inhibition of autocrine IFNs. Consistent with previous findings, the HDAC inhibitor Apicilin 1a also was identified in our primary screen and showed strong inhibition of the IFN-α gene signature. However, in our in vitro HSV-1 assay, it also blocked IFN-α dependent anti-viral activity significantly. These results not only validate our screening approach, but also highlight the importance of HDAC pathway on viral replication. Among the ISG blocked by HDAC inhibitors will be genes crucial for antiviral response. Because the Apicilin 1a significantly impaired innate anti-viral immunity, this HDAC inhibitor is not considered suitable for therapy.
The approach that we developed here (Figure 6) can be adapted readily to screen a large library of small molecular compounds that modulate other cytokine signaling pathways. The unique gene signature sets of multiple cytokine pathways and the ones of the chemical probes identified in this study can be used to provide the necessary landmarks for screening, characterization, and optimization of the resulted active compounds generated in the gene expression profiling process. The use of molecular profiling throughout the drug discovery and development process is likely to increase dramatically over the next few years. This will be based on the clear advantages of multi-variant biomarker approaches including the ability to provide a broad view of the biological state of a cell or tissue, the increased predictive power of monitoring multiple parameters simultaneously, and the power of correlating specific molecular phenotypes to clinical, histopathological, or disease model endpoints. It is clear that the increased use of molecular profiling will continue to make an important contribution to drug discovery and development efforts worldwide and, hopefully, will lead to lower failure rates, faster progression through the development process, and increasingly precise tests to match the right medicine with the right patient.
In summary, this is the first time that a large collection of well-annotated small molecule inhibitors targeting multiple intracellular signaling pathways has been evaluated using a combined chemical genomic approach. Our data suggest that targeting NF-κb and JAK/STAT signaling pathways may provide potential therapeutic benefit to type I interferonrelated diseases such as the SLE, Sjögren’s syndrome among others. In this regard, our finding provides an important proof of principle that demonstrates that small molecule inhibitors that target these two signaling pathways represent potential drug candidates for IFN-α-associated autoimmune diseases.
References
Boumpas DT et al. (1995) Systemic lupus erythematosus: emerging concepts. Part 2: Dermatologic and joint disease, the antiphospholipid antibody syndrome, pregnancy and hormonal therapy, morbidity and mortality, and pathogenesis. Ann. Intern. Med. 123:42–53.
Pestka S, Langer JA, Zoon KC, and Samuel CE. (1987) Interferons and their actions. Annu. Rev. Biochem. 56:727–77.
Pestka S, Krause CD, and Walter MR. (2004) Interferons, interferon-like cytokines, and their receptors. Immunol. Rev. 202:8–32.
Stark GR, Kerr IM, Williams BR, Silverman RH, and Schreiber RD. (1998) How cells respond to interferons. Annu. Rev. Biochem. 67:227–64.
Darnell JE Jr, Kerr IM, and Stark GR. (1994) Jak-STAT pathways and transcriptional activation in response to IFNs and other extracellular signaling proteins. Science. 264:1415–21.
Platanias LC. (2005) Mechanisms of type-I- and type-II-interferon-mediated signalling. Nat. Rev. Immunol. 5:375–86.
Pascual V, Farkas L, and Banchereau J. (2006) Systemic lupus erythematosus: all roads lead to type I interferons. Curr. Opin. Immunol. 18:676–82.
Kirou KA et al. (2004) Coordinate overexpression of interferon-alpha-induced genes in systemic lupus erythematosus. Arthritis. Rheum. 50:3958–67.
Baechler EC, Gregersen PK, and Behrens TW. (2004) The emerging role of interferon in human systemic lupus erythematosus. Curr. Opin. Immunol. 16:801–7
Crow MK, Kirou KA, and Wohlgemuth J. (2003) Microarray analysis of interferon-regulated genes in SLE. Autoimmunity. 36:481–90
Bennett L et al. (2003) Interferon and granulopoiesis signatures in systemic lupus erythematosus blood. J. Exp. Med. 197:711–23.
Baechler EC et al. (2003) Interferon-inducible gene expression signature in peripheral blood cells of patients with severe lupus. Proc. Natl. Acad. Sci. U. S. A. 100:2610–5.
Ytterberg SR and Schnitzer TJ. (1982) Serum interferon levels in patients with systemic lupus erythematosus. Arthritis. Rheum. 25:401–6.
Ronnblom L and Alm GV. (2003) Systemic lupus erythematosus and the type I interferon system. Arthritis. Res. Ther. 5:68–75.
Mathian A, Weinberg A, Gallegos M, Banchereau J, and Koutouzov S. (2005) IFN-alpha induces early lethal lupus in preautoimmune (New Zealand Black x New Zealand White) F1 but not in BALB/c mice. J. Immunol. 174:2499–506.
Santiago-Raber ML et al. (2003) Type-I interferon receptor deficiency reduces lupus-like disease in NZB mice. J. Exp. Med. 197:777–88.
Banchereau J and Pascual V. (2006) Type I interferon in systemic lupus erythematosus and other autoimmune diseases. Immunity. 25:383–392
Bol D and Ebner R. (2006) Gene expression profiling in the discovery, optimization and development of novel drugs: one universal screening platform. Pharmacogenomics. 7:227–35.
Stoughton RB and Friend SH. (2005) How molecular profiling could revolutionize drug discovery. Nat. Rev. Drug. Discov. 4:345–50.
Hieronymus H et al. (2006) Gene expression signature-based chemical genomic prediction identifies a novel class of HSP90 pathway modulators. Cancer Cell. 10:321–30.
Lamb J et al. (2006) The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science. 313:1929–35.
Wei G et al. (2006) Gene expression-based chemical genomics identifies rapamycin as a modulator of MCL1 and glucocorticoid resistance. Cancer Cell. 10:331–42.
Feng X et al. (2006) Association of increased interferon-inducible gene expression with disease activity and lupus nephritis in patients with systemic lupus erythematosus. Arthritis. Rheum. 54:2951–62.
Hua J, Kirou K, Lee C, and Crow MK. (2006) Functional assay of type I interferon in systemic lupus erythematosus plasma and association with anti-RNA binding protein autoantibody. Arthritis. Rheum. 54:1906–16.
Lit LC, Wong CK, Tam LS, Li EK, and Lam CW. (2006) Raised plasma concentration and ex vivo production of inflammatory chemokines in patients with systemic lupus erythematosus. Ann. Rheum. Dis. 65:209–15.
Narumi S, Takeuchi T, Kobayashi Y, and Konishi K. (2000) Serum levels of IFN-inducible PROTEIN-10 relating to the activity of systemic lupus erythematosus. Cytokine. 12:1561–5.
Svane IM et al. (2006) Characterization of monocyte-derived dendritic cells maturated with IFN-alpha. Scand. J. Immunol. 63:217–22.
Rider JR, Ollier WE, Lock RJ, Brookes ST, and Pamphilon DH. (1997) Human cytomegalovirus infection and systemic lupus erythematosus. Clin. Exp. Rheumatol. 15:405–9.
Harle P et al. (2001) Expression of human and macaque type I IFN-transgenes interferes with HSV-1 replication at the transcriptional and translational levels: IFN-beta is more potent than IFN-alpha 2. Virology. 290:237–48.
Nguyen KB et al. (2002) Critical role for STAT4 activation by type 1 interferons in the interferongamma response to viral infection. Science. 297: 2063–6.
Rothfuchs AG et al. (2006) STAT1 regulates IFN-alpha beta- and IFN-gamma-dependent control of infection with Chlamydia pneumoniae by nonhemopoietic cells. J. Immunol. 176:6982–90.
O’Neill LA. (2006) Targeting signal transduction as a strategy to treat inflammatory diseases. Nat. Rev. Drug. Discov. 5:549–63.
Boulares AH, Ferran MC, and Lucas-Lenard J. (1996) NF-kappaB activation Is delayed in mouse L929 cells infected with interferon suppressing, but not inducing, vesicular stomatitis virus strains. Virology. 218:71–80.
Wei L et al. (2006) NFkappaB negatively regulates interferon-induced gene expression and antiinfluenza activity. J. Biol. Chem. 281:11678–84.
Gregory D, Hargett D, Holmes D, Money E, and Bachenheimer SL. (2004) Efficient replication by herpes simplex virus type 1 involves activation of the IkappaB kinase-IkappaB-p65 pathway. J. Virol. 78:13582–90.
Birrell MA et al. (2005) Ikappa-B kinase-2 inhibitor blocks inflammation in human airway smooth muscle and a rat model of asthma. Am. J. Respir. Crit. Care. Med. 172:962–71.
Chang HM et al. (2004) Induction of interferonstimulated gene expression and antiviral responses require protein deacetylase activity. Proc. Natl. Acad. Sci. U. S. A. 101:9578–83.
Nusinzon I and Horvath CM. (2005) Unexpected roles for deacetylation in interferon- and cytokine-induced transcription. J. Interferon. Cytokine. Res. 25:745–8.
Acknowledgments
HSV-1 recombination virus with both firefly and Renilla luciferase genes in a divergent orientation from a single multiple cloning site were a gift from David A Leib (Leib Laboratory, Washington University in St. Louis, School of Medicine, St. Louis, MO, USA).
Author information
Authors and Affiliations
Corresponding author
Additional information
*
BC, QZ, and RC contributed equally to this work.
Rights and permissions
Open Access This article is published under license to BioMed Central Ltd. This is an Open Access article is distributed under the terms of the Creative Commons Attribution License ( https://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Chen, B., Zong, Q., Cibotti, R. et al. Genomic-Based High Throughput Screening Identifies Small Molecules That Differentially Inhibit the Antiviral and Immunomodulatory Effects of IFN-α. Mol Med 14, 374–382 (2008). https://doi.org/10.2119/2008-00028.Chen
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2119/2008-00028.Chen