Abstract
Neonates manifest a unique host response to sepsis even among other children. Preterm neonates may experience sepsis soon after birth or during often-protracted birth hospitalizations as they attain physiologic maturity. We examined the transcriptome using genome-wide expression profiling on prospectively collected peripheral blood samples from infants evaluated for sepsis within 24 h after clinical presentation. Simultaneous plasma samples were examined for alterations in inflammatory mediators. Group designation (sepsis or uninfected) was determined retrospectively on the basis of clinical exam and laboratory results over the next 72 h from the time of evaluation. Unsupervised analysis showed the major node of separation between groups was timing of sepsis episode relative to birth (early, <3 d, or late, ≥3 d). Principal component analyses revealed significant differences between patients with early or late sepsis despite the presence of similar key immunologic pathway aberrations in both groups. Unique to neonates, the uninfected state and host response to sepsis is significantly affected by timing relative to birth. Future therapeutic approaches may need to be tailored to the timing of the infectious event based on postnatal age.
Similar content being viewed by others
Introduction
Death or major disability occurs in nearly 40% of neonates with sepsis (1). The incidence of severe sepsis in newborns doubled (from 4.5 to 9.7 cases per 1,000 births) between 1995 and 2005 (2). In a large (n = 8,515) multicenter study of preterm infants born after <29 wks completed gestation, 36% developed sepsis during their birth hospitalization (3). Neither the treatment of neonatal sepsis nor the neurodevelopmental outcomes in surviving infants have changed significantly over the last 30 years despite multiple failed attempts to reduce the burden of infection (4,5). Accurate diagnostic tests for sepsis and prognostic tests to determine risk of poor outcome when infected are lacking.
The study of sepsis in neonates is extremely challenging for a number of reasons. Neonates uniquely experience a dramatic physiologic transition from intrauterine to extra-uterine life that begins immediately after birth and continues for several hours. The impact of dynamic transitional physiology is amplified and prolonged in infants born prematurely and is further modified by the presence of early-onset sepsis (<3 d after birth [6]). Size and blood volume limitations, particularly in the extremely preterm infant, limit the depth of diagnostic evaluation and reduce availability of samples for clinical research. Preterm neonates have underdeveloped immune system function (7) and require prolonged invasive support due to physiologic immaturity. These necessary interventions increase the risk of late-onset sepsis (≥3 d after birth and up to 120 d [8]). Lastly, the use of multiple definitions for sepsis in neonates severely limits the ability to build on experimental findings between research groups and hinders progress toward an improved understanding of the pathophysiology (9).
Molecular endotyping can be successfully leveraged to identify diagnostic and prognostic gene signatures, identify novel therapeutic targets, uncover mechanisms behind differential sepsis outcomes, and reveal rapid and dynamic shifts in transcription patterns associated with various phases of sepsis (10). We and others have shown genome-wide expression profiling (GWEP) can be a powerful tool to help uncover sepsis pathophysiology and reveal unique molecular endotypes among pediatric patients (11–15). Specifically, we showed term (completed a minimum of 36 wks gestation) neonates with septic shock manifested a unique host response compared with other pediatric patient groups (infants, toddlers and school-age children) (11). Therefore, the use of GWEP-based molecular phenotyping in septic neonates has great potential to reveal distinct pathways and unique host responses that may reveal novel pathophysiology. These findings are likely to lead to increased diagnostic accuracy and useful prognostic testing and will lay the foundation for interventional studies to improve outcomes in this vulnerable population.
In this study, we examined GWEP in prospectively collected whole blood samples from infants evaluated for early and late-onset sepsis. Infants with sepsis were distinguished from uninfected infants by the persistence of clinical signs in the setting of objective evidence of systemic inflammation with and without pathogen identification. Herein, we show that postnatal age is a critical determinant of the both the baseline and host response to sepsis.
Materials and Methods
Human Neonates and Blood Processing
All studies were approved by the Institutional Review Boards at Vanderbilt University and Duke University before their initiation. Eligible neonates admitted to the neonatal intensive care unit (NICU) and evaluated for sepsis on the basis of the judgment of the supervising clinician were enrolled after giving informed consent. No infant in the analysis had any known or suspected syndrome, abnormal karyotype or congenital malformations of any system at discharge. The decision to evaluate a neonate for sepsis was at the discretion of the attending clinician. Whole blood was collected prospectively within 24 h of the evaluation for sepsis. Infants were retrospectively classified as having sepsis on the basis of the presence of all three criteria: (a) persistently abnormal clinical examination (at least 2 d of clinical signs [Supplementary Table S1]), (b) positive culture results (blood) and (c) presence of abnormal laboratory studies supporting systemic inflammation (C-reactive protein [CRP] within 48 h of evaluation >45 mg/L). Infants with negative cultures but persistently abnormal exams and systemic inflammation were classified as having clinical sepsis. Uninfected patients had serial (at least two values 24 h apart) CRP <10 mg/dL and sterile cultures and had antimicrobial treatment discontinued <48 h after empiric initiation without subsequent clinical impact. Whole blood samples (400–500 µL) were collected in EDTA (ethylenediaminetetraacetic acid) (purple top) microtainer tubes followed by immediate RNA (ribonucleic acid) stabilization in Pax-Gene™ reagent (16). Complete blood counts were obtained to identify differential leukocyte representation with whole blood samples.
Human RNA Isolation and Microarray Hybridization
Whole blood RNA was stabilized immediately after blood draw by using the PaxGene reagent and stored at −80°C until batch processing. Total RNA was isolated from human neonatal whole blood by using the PaxGene Blood RNA System (PreAnalytiX; Qiagen/Becton Dickson). Isolated RNA was quantified by using the Nanodrop 100 (Thermo Scientific), and quality was assessed by using an Agilent 2100 BioAnalyzer (RNA integrity number [RIN], range 6.2–10) with the RNA PicoChip (Agilent). Fifteen nanograms of total RNA was amplified by using the Nugen Ovation Pico WTA System V2 (Nugen Technologies). Ten micrograms cDNA was fragmented and terminally labeled by using the Nugen Biotin Encore Kit. Labeled targets were hybridized to Affymetrix GeneChip® GGh3 Transcriptome Array (17) for 18 h at 45°C and washed according to Affymetrix standard protocols. For gene expression, analysis arrays were normalized by using Robust Multi-array Average (RMA), as implemented in Partek Genomics Suite 6.6 (Partek Incorporated). We only used annotated probe sets in the subsequent analysis, resulting in a reduction from 34,834 to 20,533 probe sets, representing 20,322 unique genes.
Plasma Cytokine Analysis
Plasma was isolated from whole blood immediately and stored at −80°C for batch processing of all samples. Granulocyte colony-stimulating factor (G-CSF), granulocyte-macrophage (GM)-CSF, tumor necrosis factor (TNF)-α, TNF receptor 1 (TNF-R1), interferon (IFN)-γ, IFN-γ RI, growth-regulated protein (GRO)-α, interleukin (IL)-1α, IL-1β, IL-1ra, IL-1sRI, IL-1sRII, IL-6, IL-8, IL-10, IL-12 p70, IL-13, IL-18, IL-18BPa, IL-19, IL-23, IL-33, IP-10, monocyte chemoattractant protein (MCP)-1, monokine induced by gamma interferon (MIG), macrophage inflammatory protein (MIP)-1α, MIP-1β, matrix metalloproteinase (MMP)-8, MMP-9, receptor for advanced glycation endproducts (RAGE), surfactant protein D (SP-D), and soluble FAS (sFAS) were determined by multiplex assay (RND; eBioscience) by using a magnetic bead-based platform (Luminex).
Statistics
A Student t test or a Wilcoxon signed-rank test was used to compare results from two groups. Values were considered significant if p < 0.05. Cytokines were analyzed by using an analysis of variance (ANOVA). Analyses were performed with Prism 6. Microarray data handling was performed as previously described (18). Significant genes were identified at a significance of p < 0.001 by using the class prediction tool implemented in BRB-Array Tools, version 4.3.0, Stable Release, developed by Richard Simon and the BRB-Array Tools Development Team (https://doi.org/linus.nci.nih.gov/BRB-Array-Tools.html). Gene expression data were deposited in GEO.
All supplementary materials are available online at https://doi.org/www.molmed.org .
Results
Patient demographics among groups are shown in Table 1. When compared by sepsis evaluation timing relative to birth, patients with sepsis were similar in gestational age and birth weight to uninfected infants, although a larger number of patients with sepsis were male. Clinical and laboratory characteristics of patients included in the analyses are shown in Supplementary Table S1. Because we examined GWEP on whole blood, we examined the differential representation among circulating white blood cells (WBCs) in each group derived from simultaneously obtained complete blood counts. A comparison of cellular representation among groups revealed no statistically significant differences in absolute number of neutrophils, lymphocytes or monocytes. Of note, we found antenatal steroid exposure (at least one dose before delivery) in 67% of the infants with sepsis and in 65% of uninfected infants. Postnatal steroid exposure (at least one dose before presentation or being actively treated with steroids at the time of blood sampling) occurred in 13% of infants with sepsis and in 10% of uninfected infants.
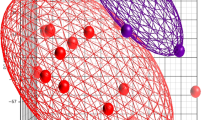
To evaluate the transcriptome in neonates with and without sepsis, we performed an unsupervised analysis that revealed 2,955 probe sets with a coefficient of variation of >0.05 between groups (Figure 1A). As shown by the heat map (Figure 1B) and principle component analysis (PCA) (Figure 1C), examination of these 2,955 probe sets by clinical group revealed timing of the sepsis episode (early versus late) as the major node of separation.
Unsupervised analysis of normalized expression data. (A) Unsupervised heat map of 2,955 probe sets with coefficient of variation >0.5 identified among all arrays. (B) Unsupervised heat map of 2,955 probe sets with coefficient of variation >0.5 identified among all groups. (C) Principle component analysis for each group on probe sets with coefficient of variation >0.5.
A supervised analysis revealed 2,573 unique genes with a p < 0.001 and at least a 1.5-fold change from uninfected infants (early or late timing) (sepsis f test; Figures 2A, B). To derive a global view of the respective gene expression patterns, we uploaded the expression data for the 2,573-gene sepsis f test to the gene expression dynamics inspector (GEDI) platform (11) and generated gene expression mosaics for each group (Figure 2C). Similar to the results of the PCA on the genes identified in the sepsis f test, the GEDI mosaics demonstrate the differences in gene expression among the groups by timing of episode. Importantly, the supervised analysis of the transcriptomic profile of uninfected infants was unique because of the timing of sepsis evaluation (Figure 3). These data indicate that timing relative to birth is a significant factor both in uninfected infants and with regards to the host immune response to sepsis. All groups showed similar trends of transcriptomic dysregulation with a greater number of upregulated genes compared with downregulated genes (Supplementary Table S2).
Supervised analysis of normalized data. Heat map (A) and PCA (B) of the 2,573 unique genes with a p < 0.001 by t test and at least a 1.5-fold change from uninfected infants (early or late timing) (Sepsis f test). (C) GEDI-generated mosaics for the 2,573 unique genes identified in the sepsis f test for each group.
To identify the affected canonical pathways in the neonatal transcriptomic host response to sepsis, the expression among groups for the 2,573 sepsis f test genes was upload to the Ingenuity Pathway Analysis™ platform. The top five canonical pathways represented among those with early and late sepsis timing were identical (role of nuclear factor of activated T cells [NFAT] in regulation of the immune response, T-cell receptor signaling, Fcγ receptor-mediated phagocytosis in macrophages and monocytes, Tec kinase signaling, and CD28 signaling in T-helper cells). However, as an example of the differential specific gene aberrations that were uncovered among those top pathways, the leukocyte extravasation network for early sepsis and late sepsis are shown in Supplementary Figure S1. Upregulated genes present in the top 10 for all groups included resistin (RETN) and IL-1R2 (Supplementary Table S2). RETN is the gene that codes for resistin, which may function as a proinflammatory cytokine (19) and is elevated in infected preterm neonates (20). IL-1R2 is a decoy receptor for IL-1 and highlights the tightly regulated activity of IL-1 in neonates (21). Several innate immunity-related genes with biology significance were represented among the top 10 in three of the four infected groups, including MMP-8 (met alloproteinase-8, a neutrophil-derived collagenase involved in the degradation of type I collagen [22]), olfactomedin 4 (OLFM4; present in neutrophil granules [23] and potentially important in neutrophil extracellular trap [NET] formation [24]), membrane-spanning 4-domains, subfamily A, member 4A (MS4A4A; tetraspanin; important for cell migration [25]) and IL-18R1 (receptor for proinflammatory cytokine IL-18 [21] and also ranked eleventh in the early sepsis group). Top downregulated genes were far more varied among groups compared with those that were upregulated. Among them, MME (membrane met allo-endopeptidase) was the most commonly represented. MME may negatively regulate the inflammatory response and is downregulated in adult nonhuman primate neutrophils after stimulation with endotoxin (26).
Plasma protein analysis revealed significant alterations in sFAS (reduces neutrophil apoptosis [27]), matrix met alloproteinases [MMP-8, MMP-9], chemokines [G-CSF, IL-8, IP-10], proinflammatory cytokines [IFN-γ, IL-1β, IL-18] and soluble receptors/inhibitors [TNF-RI, IL-1sRI, IL-1sRII, IL-18BP]) (Figure 4; all p < 0.05 by ANOVA). Plasma mediators that distinguished infected (sepsis or clinical sepsis) from uninfected infants included G-CSF, IL-8, IL-18, IL-18BP, TNF-RI and IL-1sRII. IFN-γ, sFAS and IL-1β were uniquely elevated among infants with microbiologically confirmed sepsis, whereas MMP-8 was the lone specific identifier of infants with clinical sepsis.
Discussion
We reported that term neonates with septic shock manifest a unique host response among other pediatric age-groups (11). Here, we show that timing after birth is a critical determinant of the transcriptomic baseline and the host immune response to sepsis in preterm neonates. We also used molecular endotyping to support a definition of neonatal sepsis on the basis of the presence and persistence of clinical aberrations in the setting of systemic inflammation with or without confirmed infection that can be used in future studies. These findings underscore the importance of incorporating neonatal sepsis episode timing in future diagnostic, prognostic and interventional clinical investigations.
In the moments after birth, the newborn undergoes dramatic physiologic and microbiologic changes. Thus, the difference in the whole blood transcriptomic response we found between uninfected infants in early and late periods was expected to be associated with this remarkable transition from intrauterine to extrauterine life. Early life exposures, in addition to premature birth, may also have modified the baseline. Exposure to antimicrobials and stress, both of which are nearly ubiquitous for preterm infants (28), are known to modify the subsequent immune response (29) and may represent changes associated with trained innate immunity (30,31) or adaptive immunity (32). More neonate-specific studies are necessary to delineate the impact of preterm birth (their postnatal management) on the developing immune system’s function.
The definition of neonatal sepsis is variable and often does not incorporate the presence and progression of clinical signs in the setting of laboratory evidence of systemic inflammation consistent with sepsis (9). For example, the inclusion of preterm infants with a single blood culture positive for coagulase-negative Staphylococcus (CoNS) may be considered sepsis (33), which is outside of the accepted Centers for Disease Control and Prevention definition for CoNS infection (34). CoNS can be a causative organism of sepsis in neonates but is also considered a frequent contaminant, especially when not supported by documented sustained clinical or laboratory aberrations (33,35). To avoid this important potential limitation in our investigation, we used a mixed (clinical and laboratory) definition for sepsis that incorporated (a) timing of the sepsis event relative to birth, (b) persistence of significant clinical signs for at least 2 d, (c) objective evidence of a systemic inflammatory response and (d) exclusion of all CoNS infections. We compared infants who met those criteria to infants evaluated for sepsis but who were treated with empiric antimicrobials for <48 h and who did not have a positive blood culture or evidence of systemic inflammatory response.
The translational value of genome-wide expression analyses may appear limited at first. However, unbiased exploratory “omic” approaches such as genome-wide expression profiling have the potential to reveal proteins, genes and pathways that have diagnostic, treatment and prognostic implications that were previously unknown and or unsuspected (36,37). Expression measurement of the entire genome would not be required once a suitable set of predictive genes (<100) is identified. The identification of molecular endotypes that identify classes of infants via modern multiplex messenger RNA (mRNA) quantification platforms can be obtained rapidly (8–12 h) (38,39). Integration of these techniques into intensive care environments has great potential to benefit patient populations with diverse clinical presentations of infection such as preterm neonates.
We performed our analyses on whole blood, and thus cell-specific gene expression cannot be accurately determined from these data. While the role that each cell type plays is an important consideration, examination of the host transcriptomic response to sepsis in whole blood integrates the impact of important interactions, both known and unknown, that occur in vivo. Differential WBC presence would be expected to modify the type of RNA recovered, but we did not identify statistically significant alterations in circulating WBCs between groups. This finding further substantiates the documented limitations of WBC and WBC indices to identify infected preterm infants (40,41). Minimal mortality in our cohort prevented meaningful comparisons between survivors and nonsurvivors. The study is ongoing and our intent is to analyze mortality in a separate manuscript. Importantly, whole blood-based expression profiling clearly yields biologically relevant information (11,12,42–44). These findings can be rapidly translated to the bedside to facilitate novel diagnostic and prognostic testing.
Conclusion
By using genome-wide expression profiling, we show that timing after birth is a critical determinant of the host immune response to sepsis in neonates. These findings underscore the importance of incorporating neonatal sepsis episode timing in future diagnostic, prognostic and interventional clinical investigations.
Disclosure
The authors declare that they have no competing interests as defined by Molecular Medicine, or other interests that might be perceived to influence the results and discussion reported in this paper.
References
Brocklehurst P, et al. (2011) Treatment of neonatal sepsis with intravenous immune globulin. N. Engl. J. Med. 365:1201–11.
Hartman ME, Linde-Zwirble WT, Angus DC, Watson RS. (2013) Trends in the epidemiology of pediatric severe sepsis. Pediatr. Criti. Care Med. 14:686–93.
Stoll BJ, et al. (2010) Neonatal outcomes of extremely preterm infants from the NICHD Neonatal Research Network. Pediatrics. 126:443–56.
Wynn JL, Neu J, Moldawer LL, Levy O. (2009) Potential of immunomodulatory agents for prevention and treatment of neonatal sepsis. J. Perinatol. 29:79–88.
Stoll BJ, et al. (2004) Neurodevelopmental and growth impairment among extremely low-birth-weight infants with neonatal infection. JAMA. 292:2357–65.
Stoll BJ, et al. (2002) Changes in pathogens causing early-onset sepsis in very-low-birth-weight infants. N. Engl. J. Med. 347:240–7.
Wynn J, Cornell TT, Wong HR, Shanley TP, Wheeler DS. (2010) The host response to sepsis and developmental impact. Pediatrics. 125:1031–41.
Stoll BJ, et al. (2002) Late-onset sepsis in very low birth weight neonates: the experience of the NICHD Neonatal Research Network. Pediatrics. 110:285–91.
Wynn JL, et al. (2014) Time for a neonatal-specific consensus definition for sepsis. Pediatr. Criti. Care Med. 15:523–8.
Maslove DM, Wong HR. (2014) Gene expression profiling in sepsis: timing, tissue, and translational considerations. Trends Mol. Med. 20:204–13.
Wynn JL, et al. (2011) The influence of developmental age on the early transcriptomic response of children with septic shock. Mol. Med. 17:1146–56.
Wong HR, et al. (2011) Validation of a gene expression-based subclassification strategy for pediatric septic shock. Crit. Care Med. 39:2511–7.
Wong HR, Freishtat RJ, Monaco M, Odoms K, Shanley TP. (2010) Leukocyte subset-derived genomewide expression profiles in pediatric septic shock. Pediatr Crit. Care Med. 11:349–55.
Wong HR, Odoms K, Sakthivel B. (2008) Divergence of canonical danger signals: the genomelevel expression patterns of human mononuclear cells subjected to heat shock or lipopolysaccharide. BMC Immunol. 9:24.
Wong HR, et al. (2007) Genome-level expression profiles in pediatric septic shock indicate a role for altered zinc homeostasis in poor outcome. Physiol. Genomics. 30:146–55.
Carrol ED, et al. (2007) Successful downstream application of the Paxgene Blood RNA system from small blood samples in paediatric patients for quantitative PCR analysis. BMC Immunol. 8:20.
Xu W, et al. (2011) Human transcriptome array for high-throughput clinical studies. Proc. Natl. Acad. Sci. U. S. A. 108:3707–12.
Seok J, et al. (2013) Genomic responses in mouse models poorly mimic human inflammatory diseases. Proc. Natl. Acad. Sci. U. S. A. 110:3507–12.
Sunden-Cullberg J, et al. (2007) Pronounced elevation of resistin correlates with severity of disease in severe sepsis and septic shock. Crit. Care Med. 35:1536–42.
Aliefendioglu D, Gursoy T, Caglayan O, Aktas A, Ovali F. (2014) Can resistin be a new indicator of neonatal sepsis? Pediatr. Neonatol. 55:53–7.
Garlanda C, Dinarello CA, Mantovani A. (2013) The interleukin-1 family: back to the future. Immunity. 39:1003–18.
Solan PD, et al. (2012) A novel role for matrix met alloproteinase-8 in sepsis. Crit. Care Med. 40:379–87.
Clemmensen SN, et al. (2012) Olfactomedin 4 defines a subset of human neutrophils. J. Leukoc. Biol. 91:495–500.
Welin A, et al. (2013) The human neutrophil subsets defined by the presence or absence of OLFM4 both transmigrate into tissue in vivo and give rise to distinct NETs in vitro. PLoS One. 8:e69575.
Ancuta P, et al. (2009) Transcriptional profiling reveals developmental relationship and distinct biological functions of CD16+ and CD16-monocyte subsets. BMC Genomics. 10:403.
Kaneko T, et al. (2003) Reduced neutrophil CD10 expression in nonhuman primates and humans after in vivo challenge with E. coli or lipopolysaccharide. Shock. 20:130–7.
Paunel-Gorgulu A, Flohe S, Scholz M, Windolf J, Logters T. (2011) Increased serum soluble Fas after major trauma is associated with delayed neutrophil apoptosis and development of sepsis. Crit. Care. 15:R20.
Clark RH, Bloom BT, Spitzer AR, Gerstmann DR. (2006) Reported medication use in the neonatal intensive care unit: data from a large national data set. Pediatrics. 117:1979–87.
Schokker D, et al. (2014) Early-life environmental variation affects intestinal microbiota and immune development in new-born piglets. PLoS One. 9:e100040.
Cheng SC, et al. (2014) mTOR- and HIF-1alpha-mediated aerobic glycolysis as metabolic basis for trained immunity. Science. 345:1250684.
Saeed S, et al. (2014) Epigenetic programming of monocyte-to-macrophage differentiation and trained innate immunity. Science. 345:1251086.
Wynn JL, et al. (2013) Blood stream infection is associated with altered heptavalent pneumococcal conjugate vaccine immune responses in very low birth weight infants. J. Perinatol. 33:613–8.
Smith CL, et al. (2014) Identification of a human neonatal immune-metabolic network associated with bacterial infection. Nat. Commun. 5:4649.
Horan TC, Andrus M, Dudeck MA. (2008) CDC/NHSN surveillance definition of health care-associated infection and criteria for specific types of infections in the acute care setting. Am. J. Infect. Control. 36:309–32.
Cernada M, et al. (2014) Genome-wide expression profiles in very low birth weight infants with neonatal sepsis. Pediatrics. 133:e1203–11.
Ng PC, et al. (2010) Host-response biomarkers for diagnosis of late-onset septicemia and necrotizing enterocolitis in preterm infants. J. Clin. Invest. 120:2989–3000.
Wong HR. (2013) Genome-wide expression profiling in pediatric septic shock. Pediatr. Res. 73:564–9.
Cuenca AG, et al. (2013) Development of a genomic metric that can be rapidly used to predict clinical outcome in severely injured trauma patients. Crit. Care Med. 41:1175–85.
Wong HR, et al. (2015) Developing a clinically feasible personalized medicine approach to pediatric septic shock. Am. J. Respir. Crit. Care Med. 191:309–15.
Hornik CP, et al. (2012) Use of the complete blood cell count in late-onset neonatal sepsis. Pediatr. Infect. Dis. J. 31:803–7.
Hornik CP, et al. (2012) Use of the complete blood cell count in early-onset neonatal sepsis. Pediatr Infect. Dis J 31:799–802.
Shanley TP, et al. (2007) Genome-level longitudinal expression of signaling pathways and gene networks in pediatric septic shock. Mol. Med. 13:495–508.
Wong HR, et al. (2009) Genomic expression profiling across the pediatric systemic inflammatory response syndrome, sepsis, and septic shock spectrum. Crit. Care Med. 37:1558–66.
Wong HR, et al. (2009) Identification of pediatric septic shock subclasses based on genome-wide expression profiling. BMC Med. 7:34.
Acknowledgments
Grant support for this work was provided by National Institutes of Health (NIH)/National Institute of General Medical Sciences (NIGMS) (GM106143 to JL Wynn and GM096994 to HR Wong), The Gerber Foundation (to JL Wynn), and Vanderbilt Turner-Hazinski awards (to JL Wynn).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it.
The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this license, visit (https://doi.org/creativecommons.org/licenses/by-nc-nd/4.0/)
About this article
Cite this article
Wynn, J.L., Guthrie, S.O., Wong, H.R. et al. Postnatal Age Is a Critical Determinant of the Neonatal Host Response to Sepsis. Mol Med 21, 496–504 (2015). https://doi.org/10.2119/molmed.2015.00064
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2119/molmed.2015.00064