Abstract
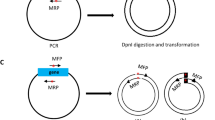
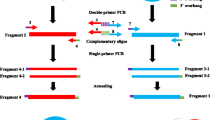
Restriction enzyme mediated integration is a widely used and effective method for insertional mutagenesis in Dictyostelium discoideum. In this method, plasmid rescue is used to clone the genomic deoxyribonucleic acid (DNA) sequences that flank the insertion site. For this to be effective, it is necessary to first find a convenient restriction enzyme site within the genomic DNA. This is a time-consuming process that requires Southern blot analysis of the mutant DNA. In addition, plasmid rescue requires transformation into highly competent Escherichia coli. Problems can arise owing to unstable genomic sequences, damage to the plasmid DNA and exogenous plasmid contamination. We have established a simple and rapid polymerase chain reaction-based technique that works for all mutants and circumvents the need for Southern blot analysis and plasmid rescue.
Similar content being viewed by others
References
Kuspa, A. and Loomis, W. F. (1992) Tagging developmental genes in Dictyostelium by restriction enzyme-mediated integration of plasmid DNA. Proc. Natl. Acad. Sci. USA 89, 8803–8807.
Long, D., Martin, M., Sundberg, E., Swinburne, J., Puangsomlee, P., and Coupland, G. (1993) The maize transposable element system Ac/Ds as a mutagen in Arabidopsis: identification of an albino mutation induced by Ds insertion. Proc. Natl. Acad. Sci. USA 90, 10,370–10,374.
Hartl, D. L. and Ochman, H. (1996) Inverse polymerase chain reaction. In Methods in Molecular Biology, vol. 58: Basic DNA and RNA Protocols, Harwood, A. J., ed. Humana, Totowa, NJ, pp. 293–301.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Keim, M., Williams, R.S.B. & Harwood, A.J. An inverse PCR technique to rapidly isolate the flanking DNA of Dictyostelium insertion mutants. Mol Biotechnol 26, 221–224 (2004). https://doi.org/10.1385/MB:26:3:221
Issue Date:
DOI: https://doi.org/10.1385/MB:26:3:221