Abstract
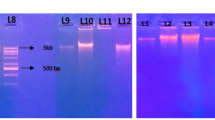
An important prerequisite for successful construction of a metagenome library is an efficient procedure for extracting DNA from environmental samples. We compared three indirect and four direct extraction methods, including a commercial kit, in terms of DNA yield, purity, and time requirement. A special focus was on methods that are appropriate for the extraction of environmental DNA (eDNA) from very limited sample sizes (0.1 g) to enable a highly parallel approach. Direct extraction procedures yielded on average 100-fold higher DNA amounts than indirect ones. A drawback of direct extraction was the small fragment sizeof approx 12 kb. The quality of the extracted DNA was evaluated by the ability of different restriction enzymes to digest the eDNA. Only the commercial kit and a direct extraction method using freeze-thaw cell lysis in combination with an in-gel patch electrophoresis with hydroxyapatite to remove humic acid substances yielded DNA, which was completely digested by all restriction enzymes. Moreover, only DNA extracted by these two procedures could be used as template for the amplification of fragments of several 16S rDNA, 18SrDNA groups under standard polymerase chain reaction conditions.
Similar content being viewed by others
References
Schleper, C., Jurgens, G., and Jonuscheit, M. (2005), Nat. Rev. Microbiol. 3, 479–488.
Streit, W. R., Daniel, R., and Jaeger, K. E. (2004), Curr. Opin. Biotechnol. 15, 1–6.
Daniel, R. (2004), Curr. Opin. Biotechnol. 15, 199–204.
Galbraith, E. A., Antonopoulos, D. A., and White, B. A. (2004), Environ. Microbiol. 6(9), 928–937.
Gabor, E. M., Alkema, W. B. L., and Janssen, D. B. (2004), Environ. Microbiol. 6(9), 879–886.
Voget, S., Leggewie, C., Uesbeck, A., Raasch, C., Jaeger, K.-E., and Streit, W. R. (2003), Appl. Environ. Microbiol. 69, 6235–6242.
Torsvik, V. and Ovreas, L. (2002), Curr. Opin. Biotechnol. 5, 240–245.
Rondon, M. R., Goodman, R. M., and Handelsman, J. (1999), Science 17, 403–409.
Amann, R. I., Ludwig, W., and Schleifer, K. H. (1995), Microbiol. Rev. 59, 143–169.
Zengler, K., Toledo, G., Rappé, M., Elkins, J., Mathur, E. J., Short, J. M., and Keller, M. (2002), Proc. Natl. Acad. Sci. USA 99, 15,681–15,686.
Kaeberlein, T., Lewis, K., and Epstein, S. S. (2002), Science 296, 1127–1129.
Kozdrój, J. and van Elsas, J. D. (2000), Biol. Fertil. Soils 31, 372–378.
Kauffmann, I. M., Schmitt, J., and Schmid, R. D. (2004), Appl. Microbiol. Biotechnol. 64, 665–670.
Robe, P., Nalin, R., Capellano, C., Vogel, T. M., and Simonet, P. (2003), Eur. J. Soil Biol. 39, 183–190.
Gabor, E. M., de Vries, E. J., and Janssen, D. B. (2003), FEMS Microbiol. Ecol. 44, 153–163.
Lloyd-Jones, G. and Hunter, D. W. F. (2001), Soil Biol. Biochem. 33, 2053–2059.
Quaiser, A., Ochsenreiter, T., Klenk, H.-P., et al. (2002) Environ. Microbiol. 4 (10), 603–611.
Orsini, M. and Romano-Spica, V. (2001), Lett. Appl. Microbiol. 33, 17–20.
Courtois, S., Frostegård, Å, Göransson, P., Depret, G., Jeannin, P., and Simonet, P. (2001), Environ. Microbiol. 3(7), 431–439.
Tien, C. C., Chao, C. C., and Chao, W. L. (1999), J. Appl. Microbiol. 86, 937–943.
Moreira, D. (1998), Nucleic Acids Res. 26, 3309, 3310.
Yeates, C., Gillings, M. R., Davidson, A. D., Altavilla, N., and Veal, D. A. (1997), Lett. Appl. Microbiol. 25, 303–307.
Zhou, J., Bruns, M. A., and Tiedje, J. M. (1996), Appl. Environ. Microbiol. 62, 316–322.
Young, C. C., Burghoff, R. L., Keim, L. G., Minak-Bernero, V., Lute, J. R., and Hinton, S. M. (1993), Appl. Environ. Microbiol. 59, 1972–1974.
Jacobsen, C. S. and Rasmussen, O. F. (1992), Appl. Environ. Microbiol. 58, 2458–2462.
Tsai, Y.-L. and Olson, B. H. (1991), Appl. Environ. Microbiol. 57, 1070–1074.
Holben, W. E. (1994), in Methods of Soil Analysis, Part 2. Microbiological and Biochemical Properties: Isolation and Purification of Bacterial DNA from Soil, SSSA Book Series no 5, Weaver, R. W., Angle, S., and Bottomley, P. D. eds, Soil Science Society of America, Madison, WI, pp. 727–752.
Santos, D. A. (2001), Mol. Biochnol. 17, 59–64.
Roh, C. H., Villatte, F., Kim, B. G., and Schmid, R.D. (2005), Electrophoresis, 26, 3055–3061.
Bertrand, H., Poly, F., Van, V. T., Lombard, N., Nalin, R., Vogel, T. M., and Simonet, P. (2005), J. Microbiol. Methods 62, 1–11.
Rhee, J. K., Ahn, D. G., Kim, Y. G., and Oh, J. W. (2005), Appl. Environ. Microbiol. 71(2), 817–825.
Ginolhac A., Jarrin C., Gillet B., et al. (2004), Appl. Environ. Microbiol. 70, 5522–5527.
Miller, D. N., Bryant, J. E., Madsen, E. L., and Ghiorse, W. C. (1999), Appl. Environ. Microbiol. 65, 4715–4724.
Miller, D. N. (2001), J. Microbiol. Methods 44, 49–58.
Yun, J., Kang, S., Park, S., Yoon, H., Kim, M. J., Heu, S., and Ryu, S. (2004), Appl. Environ. Microbiol., 70, 7229–7235.
Knietsch, A., Bowien, S., Whited, G., Gottschalk, G., and Daniel, R. (2003), Appl. Environ. Microbiol. 69, 3048–3060.
Henne, A., Schmitz, R.A., Bomeke, M., Gottschalk, G., and Daniel, R. (2000), Appl. Environ. Microbiol. 66, 3113–3116.
Henne, A., Daniel, R., Schmitz, R. A., and Gottschalk, G. (1999), Appl. Environ. Microbiol. 65, 3901–3907.
Edwards, U., Rogall, T.L., Blocker, H., Emde, M., and Bottger, E. C. (1989), Nucleic Acids Res. 17, 7843–7853.
Kuske, C. R., Banton, K. L., Adorada, D. L., Stark, P. C., Hill, K. K., and Jackson, P.J. (1998), Appl. Environ. Microbiol. 64, 2463–2472.
Stackebrandt, E., Liesack, W., and Goebel, B. M. (1993), FASEB J. 7, 232–236.
Yeates, C., Gillings, M. R., Davison, A. D., Altavilla, N., and Veal, D. A. (1998) Biol. Procedures 14, 40–47.
White, T. J., Bruns, T., Lee, S., and Taylor, J. (1990), in PCR Protocols: A Guide to Methods and Applications: Amplification and Direct Sequencing of Ribosomal RNA Genes for Phylogenetics, Innis, M. A., Gelfand, D. H., Sninsky, J. J., and White, T. J., eds., Academic, New York, pp. 315–322.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Roh, C., Villatte, F., Kim, BG. et al. Comparative study of methods for extraction and purification of environmental DNA from soil and sludge samples. Appl Biochem Biotechnol 134, 97–112 (2006). https://doi.org/10.1385/ABAB:134:2:97
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1385/ABAB:134:2:97