Abstract
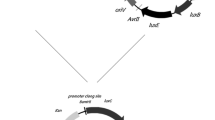
The NagR protein is a response regulatory protein found in the bacterium Ralstonia sp. U2 that is involved in sensing for salicylic acid and the subsequent induction of the operaon just upstream of its gene. The genes encoded for in this operon are involved in the degradation of salicylic acid. Escherichia coli strain RFM443 carrying a fusion of the Photorhabdus luminesscens luxCDABE operon with the nagR gene and upstream region of the nagAa gene was constructed and characterized with respect to its optimum temperature, its response time and kinetics, and its ability to deterctnumerous benzoic acid derivatives. Although capable of detecting 0.5 mM salicylic acid at any temperature between 28 and 40°C, this E. coli strain, labeled DNT5, showed its greatest relative activity at 30°C, i.e., the temperature at which the largest induction was seen. Furthermore, experiments done with numerous benzoic acid derivatives found the NagR protein to be responsive to only a few of the compounds tested, including salicylic acid and 3-methyl salicylic acid and 3-methyl saliyclic acid, and acetyl salicylic acid was the strongest inducer. The lower limits of detection for these compounds with E. coli strain DNT5 were also established, wit the native inducer, salicylic acid, giving the most sensitive response and detectable down to a concentration of about 2 μM. A second lux fusion plasmid was also constructed and transformed into an NahR background, Pseudomonas putida KCTC1768. Within this strain, NAGK-1768, the supplemental activity of the NahR protein on the nagAn promoter, was shown to extend both the range of chemicals detected and the sensitivity.
Similar content being viewed by others
References
Garcia-Valdes, E., Cozar, E., Rotger, R., Lalucat, J., and Ursing, J. (1988), Appl. Environ. Microbiol. 54, 2478–2485.
Grund, A. D. and Gunsalus, I. C. (1983), J. Bacteriol. 156, 89–94.
Kuhm, A. E., Stolz, A., and Knackmuss, H. J. (1991) Biodegradation 2, 115–120.
Mihelcic, J. R. and Luthy, R. G. (1988), Appl. Environ. Microbiol. 54, 1188–1198.
Kiyohara, H., Nagao, K., Kouno, K., and Yano K. (1982), Appl. Environ. Microbiol. 43, 458–461.
Weissenfels, W. D., Beyer, M., and Klein, J. (1990), Appl. Microbiol. Biotechnol. 32, 479–484.
Grifoll, M., Selifonov, S. A., Gatlin, C. V., and Chapman, P. J. (1995), Appl. Environ. Microbiol. 61, 3711–3723.
Heitkamp, M. A., Franklin, W., and Cerniglia, C. E. (1988) Appl. Environ. Microbiol. 54, 2549–2555.
Schneider, J., Grosser, R., Jayasimhulu, K., Xue, W., and Warshawsky, D. (1996), Appl. Environ. Microbiol. 62, 13–19.
Stringfellow, W. T. and Aitken, M. D. (1995), Appl. Environ. Microbiol. 61, 357–362.
Saneseverino, J., Applegate, B. M., King, J. M., and Sayler, G. S. (1993), Appl. Environ. Microbiol. 59, 1931–1937.
Tsuda, M. and Iino, T. (1990), Mol. Gen. Genet. 223, 33–39.
Yen, K. M. and Gunsalus, I. C. (1985), J. Bacteriol. 162, 1008–1013.
Zhou, N. Y., Fuenmayor, S. L., and Williams, P. A. (2001), J. Bacteriol. 183, 700–708.
Yen, K. M. and Serdar, C. M. (1988), Crit. Rev. Microbiol. 15, 247–268.
Huang, J. Z. and Schell, M. A. (1991), J. Biol. Chem. 266, 10830–8.
Schell, M. A. (1985), Gene 36, 301–309.
Schell, M. A. (1986), Proc. Natl. Acad. Sci. USA 83, 369–373.
Schell, M. A. and Poser, E. F. (1989), J. Bacteriol. 171, 837–846.
Cebolla, A., Sousa, C., and de Lorenzo, V. (1997), J. Biol. Chem. 272, 3986–3892.
Schell, M. A. and Wender, P. E. (1986), J. Bacteriol. 166, 9–14.
You, I. S., Ghosal, D., and Gunsalus, I. C. (1988), J. Bacteriol. 170, 5409–5415.
Schell, M. A., Brown, P. H., and Raju, S. (1990), J. Biol. Chem. 265, 3844–3850.
Jones, R. M., Britt-Compton, B., and Williams, P. A. (2003), J. Bacteriol. 185, 5847–5853.
Van Dyk, T. K. and Rosson, R. A. (1998), in Methods in Molecular Biology: Bioluminecence Methods and Protocols, vol. 102, LaRossa, R. A., ed., Humana, Totowa, NJ, pp. 85–95.
Rogowsky, P. M., Close, T. J., Chimera, J. A., Shaw, J. J., and Kado, C. I. (1987), J. Bacteriol. 169, 5101–5112.
Burlage, R. S., Sayler, G. S., and Larimer F. (1990), J. Bacteriol. 172, 4749–4757.
King, J. M. H., DiGrazia, P. M., Applegate, B., Burlage, R., Sanseverino, J., Dunbar, P., Larimer, F., and Sayler, G. S. (1991), Science 249, 778–781.
Heitzer, A., Webb, O. F., Thonnard, J. E., and Sayler, G. S. (1992), Appl. Environ. Microbiol. 58, 1839–1846.
Fuemayor, S. L., Wild, M., Boyes, A. L., and Williams, P. A. (1998), J. Bacteriol. 180, 2522–2530.
Lee, H. Y., Choi, S. H., and Gu, M. B. (2003), Biotechnol. Bioprocess. Eng. 8, 101–105.
Mitchell, R. J., Ahn, J. M., and Gu, M. B. (2005), J. Microbiol. Biotech., in press.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mitchell, R.J., Gu, M.B. Construction and evaluation of nagR-nagAa::lux fusion strains in biosensing for salicylic acid derivatives. Appl Biochem Biotechnol 120, 183–197 (2005). https://doi.org/10.1385/ABAB:120:3:183
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1385/ABAB:120:3:183