Abstract
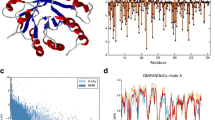
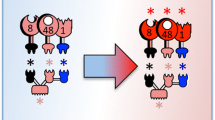
Cellulases are a complex group of enzymes that are fundamental for the degradation of amorphous and crystalline cellulose in lignocellulosic material. Unfortunately, cellulases have a low catalytic efficiency on their substrates when compared to similar enzymes such as amylases, which has led to a strong interest in improving their activities. Thermobifida fusca secretes six cellulose degrading enzymes: two exo- and three endocellulases and an endo/exocellulase Cel9A (formerly called E4). Cel9A shows unique properties because of its endo- and exocellulase characteristics, strong activity on crystalline cellulose, and good synergistic properties. Therefore, it is an excellent target for mutagenesis techniques to improve crystalline cellulose degradation. In this article, we describe research conducted to improve Cel9A catalytic efficiency using a rational design and computer modeling. A computer model of Cel9A was created using the program CHARMM plus its PDB structure and a cellohexose molecule attached to the catalytic site as a starting model. Initially molecular graphics and energy minimization were used to extend the cellulose chain to 18 glucose residues spanning the catalytic domain and cellulose-binding domain (CBD). The interaction between this cellulose chain and conserved CBD residues was determined in the model, and mutations likely to improve the binding properties of the CBD were selected. Site-directed mutations were carried out using the pET vector pET26b, Escherichia coli DH5-α, and the QuickChange mutagenesis method. E. coli BL21-DE3 was used for protein production and expression. The purified proteins were assayed for enzymatic activity on filter paper, swollen cellulose, bacterial microcrystalline cellulose, and carboxymethylcellulose (CMC). Mutation of the conserved residue F476 to Y476 gave a 40% improved activity in assays with soluble and amorphous cellulose such as CMC and swollen cellulose.
Similar content being viewed by others
References
Smith, J. E. (1996), in Biotechnology, Cambridge University Press, Cambridge, MA, pp. 22–23.
Walker, L. P., et al. (1993) Biotech. Bioeng. 42, 1019–1028.
Teeri, T. T. (1997), Trends Biotech. 15, 160.
Walker, L. P., et al. (1992), Biotech. Bioeng. 40, 1019–1026.
Irwin, D. C., Spezio, M., Walker, L. P., and Wilson, D. B. (1993), Biotech. Bioeng. 42, 1002–1013.
Davies, G. J., et al. (1996), Acta Crystallographica D52, 7–17.
Henrissat, B., et al. (1989), Gene (Amst.) 81, 83–95.
Juy, M., et al. (1992), Nature 357, 89–91.
Davies, G. J., et al. (1993), Nature 365, 362–364.
Takashima, S., Nakamura, A., Masaki, H., and Uozumi, T. (1996), Biosci. Biotech. Biochem. 60(1), 77–82.
Kraulis, P. J., et al. (1989), Biochemistry 28, 7241–7257.
Dalbøge, H. and Heldt-Hansen, H. P. (1994), Mol. Gen. Genet. 243, 253–260.
Azevedo, Mde O., et al. (1990), J. Gen. Microbiol. 136, 120–123.
Cui, Z., et al. (1992), Biosci. Biotechnol. Biochem. 56, 1230–1235.
Huang, J., et al. (1992), J. Bacteriol. 174, 1314–1323.
Ong, E., et al. (1989), Trends Biotechnol. 7, 239–243.
Divne, C., et al. (1994), Science 265, 524–528.
Rouvinen, J., et al. (1990), Science 249, 380–386.
Spezio, M., et al. (1993), Biochemistry 32, 9906–9916.
Sakon, J., Irwin, D., Wilson, D. B., and Karplus, P., A. (1997), Nat. Struct. Biol. 4, 810–818
Irwin, D. C., et al. (1998), J. Bacteriol. 180(7), 1709–1714.
Sacco, M., Millet, J., and Aubert, J. P. (1984), Ann. Microbiol. (Inst. Pasteur) 135A, 485–488.
Curry, C., et al. (1988), Appl. Environ. Microbiol. 54, 476–484.
Konstantinidis, A. K., et al. (1993), Biochem. J. 291, 883.
Stemmer, W. P. C. (1994), Nature 370, 389–391.
Stemmer, W. P. C. (1996), Nat. Biotechnol. 14, 315–319.
Stemmer, W. P. C. (1995), Biotechnology 13, 549–553.
Matsumura, I. and Ellington, A. D. (1996), Nat. Biotechnol. 14, 366.
Zhang, J. H., Dawes, G., and Stemmer, W. P. C. (1997), Proc. Natl. Acad. Sci. USA 94, 4504–4509.
Cleland, J. L. and Craik, C. (1996), in Protein Engineering Principles and Practice, Wyley-Liss, New York, NY, pp. 22.
Taylor, J. S., Teo, B., Wilson, D. B., and Brady, J. W. (1988), Prot. Eng. 8(11), 1145–1152.
Brooks, C. L., Karplus, M. and Pettit, B. M. (1988), in Proteins: A Theoretical Perspective of Dynamics, Structure an Thermodynamics, Adv. Chem. Phys., Vol. 71, Wiley-Interscience, New York, NY.
MacKerell, A. D., Bashford, D., et al (1998), J. Physi. Chem. 102, 3586–3616.
Palma, R., Zuccato, P., et al. (2000), in Glycosyl Hydrolases in Biomass Conversion. Himmel, M. E., ed., American Chemical Society, Washington, DC.
Van Gunsteren, W. F., and Berendsen, H. J. C. (1977) Mol. Physiol. 34(5), 1311–1327.
Bayer, A. E., et al. (1998) in Carhohydrases from Trichoderma reesei and Other Microorganisms. Claeyssens, M., Nerinckx, W., and Piens, K., eds., The Royal Society of Chemistry, pp. 39–65.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Escovar-Kousen, J.M., Wilson, D. & Irwin, D. Integration of computer modeling and initial studies of site-directed mutagenesis to improve cellulase activity on Cel9A from Thermobifida fusca . Appl Biochem Biotechnol 113, 287–297 (2004). https://doi.org/10.1385/ABAB:113:1-3:287
Issue Date:
DOI: https://doi.org/10.1385/ABAB:113:1-3:287