Abstract
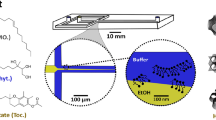
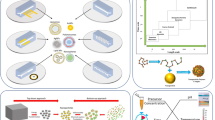
Nanoparticles (NPs) have emerged as a highly useful and clinically translatable drug delivery platform for vast therapeutic payloads. Through the precise tuning of their physicochemical properties, NPs can be engineered to exhibit controlled drug release properties, enhanced circulation times, improved cellular uptake and targeting, and reduced toxicity profiles. Conventional bulk methods for the production of polymeric NPs suffer from the ability to control their size and polydispersity, batch-to-batch variability, significant preparation times, and low recovery. Here, we describe the development and optimization of a high-throughput microfluidic method to produce cargo-less immunomodulatory nanoparticles (iNPs) and their formulation-dependent anti-inflammatory properties for the modulation of lipopolysaccharide (LPS)-induced macrophage responses. Using poly(lactic acid) (PLA) as the core-forming polymer, a rapid and tunable microfluidic hydrodynamic flow-focusing method was developed and optimized to systematically evaluate the role of polymer and surfactant concentration, surfactant chemistry, and flow rate ratio (FRR) on the formation of iNPs. A set of iNPs with 6 different surface chemistries and 2 FRRs was then prepared to evaluate their inherent anti-inflammatory effects using bone marrow-derived macrophages stimulated with the Toll-like receptor 4 agonist, LPS. Finally, a lyophilization study was performed using various cryoprotectants and combinations to identify preferable conditions for iNP storage. Overall, we demonstrate a highly controlled and reproducible method for the formulation of iNPs using microfluidics and their formulation-dependent inherent anti-inflammatory immunomodulatory properties, which represents a potentially promising strategy for the management of inflammation.
Graphical Abstract







Similar content being viewed by others
References
Peer D, Karp JM, Hong S, Farokhzad OC, Margalit R, Langer R. Nanocarriers as an emerging platform for cancer therapy. Nat Nanotechnol. 2007;2:751–60.
Anselmo AC, Mitragotri S. Nanoparticles in the clinic: an update. Bioeng Transl Med. 2019;4:e10143.
Pearson RM, Podojil JR, Shea LD, King NJC, Miller SD, Getts DR. Overcoming challenges in treating autoimmuntity: development of tolerogenic immune-modifying nanoparticles. Nanomed Nanotechnol. 2018;18:282–91.
Pearson RM, Hsu H-J, Bugno J, Hong S. Understanding nano-bio interactions to improve nanocarriers for drug delivery. MRS Bull. 2014;39:227–37.
Rao JP, Geckeler KE. Polymer nanoparticles: preparation techniques and size-control parameters. Prog Polym Sci. 2011;36:887–913.
Sanjay ST, Zhou W, Dou M, Tavakoli H, Ma L, Xu F, Li X. Recent advances of controlled drug delivery using microfluidic platforms. Adv Drug Deliv Rev. 2018;128:3–28.
Shepherd SJ, Issadore D, Mitchell MJ. Microfluidic formulation of nanoparticles for biomedical applications. Biomaterials. 2021;274:120826.
Zhang L, Chen Q, Ma Y, Sun J. Microfluidic methods for fabrication and engineering of nanoparticle drug delivery systems. ACS Appl Bio Mater. 2019;3:107–20.
Shrimal P, Jadeja G, Patel S. A review on novel methodologies for drug nanoparticle preparation: microfluidic approach. Chem Eng Res Des. 2020;153:728–56.
Zhao C-X, He L, Qiao SZ, Middelberg APJ. Nanoparticle synthesis in microreactors. Chem Eng Sci. 2011;66:1463–79.
Hamdallah SI, Zoqlam R, Erfle P, Blyth M, Alkilany AM, Dietzel A, Qi S. Microfluidics for pharmaceutical nanoparticle fabrication: the truth and the myth. Int J Pharm. 2020;584:119408.
Liu D, Cito S, Zhang Y, Wang CF, Sikanen TM, Santos HA. A versatile and robust microfluidic platform toward high throughput synthesis of homogeneous nanoparticles with tunable properties. Adv Mater. 2015;27:2298–304.
Niculescu A-G, Chircov C, Bîrcă AC, Grumezescu AM. Nanomaterials synthesis through microfluidic methods: an updated overview. Nanomaterials. 2021;11:864.
Ma J, Lee SM-Y, Yi C, Li C-W. Controllable synthesis of functional nanoparticles by microfluidic platforms for biomedical applications—a review. Lab Chip. 2017;17:209–26.
Seo M, Matsuura N. Direct incorporation of lipophilic nanoparticles into monodisperse perfluorocarbon nanodroplets via solvent dissolution from microfluidic-generated precursor microdroplets. Langmuir. 2014;30:12465–73.
Casey LM, Kakade S, Decker JT, Rose JA, Deans K, Shea LD, Pearson RM. Cargo-less nanoparticles program innate immune cell responses to toll-like receptor activation. Biomaterials. 2019;218:119333.
Getts DR, Martin AJ, McCarthy DP, Terry RL, Hunter ZN, Yap WT, Getts MT, Pleiss M, Luo X, King NJ, Shea LD, Miller SD. Microparticles bearing encephalitogenic peptides induce T-cell tolerance and ameliorate experimental autoimmune encephalomyelitis. Nat Biotechnol. 2012;30:1217–24.
McCarthy DP, Yap JW, Harp CT, Song WK, Chen J, Pearson RM, Miller SD, Shea LD. An antigen-encapsulating nanoparticle platform for TH1/17 immune tolerance therapy. Nanomed Nanotechnol. 2017;13:191–200.
Bihari P, Vippola M, Schultes S, Praetner M, Khandoga AG, Reichel CA, Coester C, Tuomi T, Rehberg M, Krombach F. Optimized dispersion of nanoparticles for biological in vitro and in vivo studies. Part Fibre Toxicol. 2008;5:14.
Strickson S, Campbell DG, Emmerich CH, Knebel A, Plater L, Ritorto MS, Shpiro N, Cohen P. The anti-inflammatory drug BAY 11-7082 suppresses the MyD88-dependent signalling network by targeting the ubiquitin system. Biochem J. 2013;451:427–37.
Bozza FA, Salluh JI, Japiassu AM, Soares M, Assis EF, Gomes RN, Bozza MT, Castro-Faria-Neto HC, Bozza PT. Cytokine profiles as markers of disease severity in sepsis: a multiplex analysis. Crit Care. 2007;11:R49.
Valencia PM, Pridgen EM, Rhee M, Langer R, Farokhzad OC, Karnik R. Microfluidic platform for combinatorial synthesis and optimization of targeted nanoparticles for cancer therapy. ACS Nano. 2013;7:10671–80.
Chiesa E, Dorati R, Modena T, Conti B, Genta I. Multivariate analysis for the optimization of microfluidics-assisted nanoprecipitation method intended for the loading of small hydrophilic drugs into PLGA nanoparticles. Int J Pharm. 2018;536:165–77.
Valencia PM, Farokhzad OC, Karnik R, Langer R. Microfluidic technologies for accelerating the clinical translation of nanoparticles. Nat Nanotechnol. 2012;7:623–9.
Dobrovolskaia MA, McNeil SE. Immunological properties of engineered nanomaterials. Nat Nanotechnol. 2007;2:469–78.
Dobrovolskaia MA, Aggarwal P, Hall JB, McNeil SE. Preclinical studies to understand nanoparticle interaction with the immune system and its potential effects on nanoparticle biodistribution. Mol Pharm. 2008;5:487–95.
Huang L, Zhao X, Qi Y, Li H, Ye G, Liu Y, Zhang Y, Gou J. Sepsis-associated severe interleukin-6 storm in critical coronavirus disease 2019. Cell Mol Immunol. 2020;17:1092–4.
Schulte W, Bernhagen J, Bucala R. Cytokines in sepsis: potent immunoregulators and potential therapeutic targets—an updated view. Mediators Inflamm. 2013;2013:165974.
Tisoncik JR, Korth MJ, Simmons CP, Farrar J, Martin TR, Katze MG. Into the eye of the cytokine storm. Microbiol Mol Biol Rev. 2012;76:16–32.
Karnik R, Gu F, Basto P, Cannizzaro C, Dean L, Kyei-Manu W, Langer R, Farokhzad OC. Microfluidic platform for controlled synthesis of polymeric nanoparticles. Nano Lett. 2008;8:2906–12.
Rayahin JE, Buhrman JS, Zhang Y, Koh TJ, Gemeinhart RA. High and low molecular weight hyaluronic acid differentially influence macrophage activation. ACS Biomater Sci Eng. 2015;1:481–93.
Dashtimoghadam E, Mirzadeh H, Taromi FA, Nyström B. Microfluidic self-assembly of polymeric nanoparticles with tunable compactness for controlled drug delivery. Polymer. 2013;54:4972–9.
Abdelwahed W, Degobert G, Stainmesse S, Fessi H. Freeze-drying of nanoparticles: formulation, process and storage considerations. Adv Drug Deliv Rev. 2006;58:1688–713.
Errea A, Cayet D, Marchetti P, Tang C, Kluza J, Offermanns S, Sirard J-C, Rumbo M. Lactate inhibits the pro-inflammatory response and metabolic reprogramming in murine macrophages in a GPR81-independent manner. PLoS ONE. 2016;11:e0163694.
Allen RP, Bolandparvaz A, Ma JA, Manickam VA, Lewis JS. Latent, immunosuppressive nature of poly(lactic-co-glycolic acid) microparticles. ACS Biomater Sci Eng. 2018;4:900–18.
Sangsuwan R, Thuamsang B, Pacifici N, Allen R, Han H, Miakicheva S, Lewis JS. Lactate exposure promotes immunosuppressive phenotypes in innate immune cells. Cell Mol Bioeng. 2020;13:541–57.
Behzadi S, Serpooshan V, Tao W, Hamaly MA, Alkawareek MY, Dreaden EC, Brown D, Alkilany AM, Farokhzad OC, Mahmoudi M. Cellular uptake of nanoparticles: journey inside the cell. Chem Soc Rev. 2017;46:4218–44.
Correa S, Boehnke N, Barberio AE, Deiss-Yehiely E, Shi A, Oberlton B, Smith SG, Zervantonakis I, Dreaden EC, Hammond PT. Tuning nanoparticle interactions with ovarian cancer through layer-by-layer modification of surface chemistry. ACS Nano. 2020;14:2224–37.
Yang H, Fung S-Y, Liu M. Programming the cellular uptake of physiologically stable peptide–gold nanoparticle hybrids with single amino acids. Angew Chem Int Ed. 2011;50:9643–6.
Yang H, Fung S-Y, Xu S, Sutherland DP, Kollmann TR, Liu M, Turvey SE. Amino acid-dependent attenuation of Toll-like receptor signaling by peptide-gold nanoparticle hybrids. ACS Nano. 2015;9:6774–84.
Funding
Research reported in this publication was supported by Startup funds provided by the University of Maryland, Baltimore (UMB), and the National Institute of General Medical Sciences of the National Institutes of Health under Award Number R35GM142752. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
NT and RMP contributed to the design of experiments, analysis of data, and compilation of the manuscript. JS and NT contributed to optimization and development of the microfluidic system. SKB and JS executed experiments related to in vitro testing of nanoparticles and cryoprotectant studies. BS operated the SEM and collected images of the nanoparticles. All authors discussed and analyzed the results and contributed to the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Guest Editors: Aliasger Salem, Juliane Nguyen and Kristy Ainslie.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12248_2021_645_MOESM1_ESM.tiff
Figure S1. Six iNP formulations utilizing one of the six various surfactants on the iNP exterior. Chemical structures of (1) poly(ethylene-alt-maleic acid) E60/E400, (2) poly(vinyl alcohol) (PVA), (3) poly(acrylic acid) (PAA), (4) poly(glutamic acid) (PGA), and (5) hyaluronic acid (HA) utilized for iNP synthesis. Supplementary file1 (TIFF 190 kb)
12248_2021_645_MOESM3_ESM.tiff
Figure S3: Comparison of two different nanoparticle synthesis methods: emulsification and microfluidics. Both nanoparticles were synthesized using E400 and PLA and measured for particle size (nm) and PDI. (A) A statistical F test to compare variance was performed, where particle size measurements for emulsification (n=18, SD=144.4) and microfluidics (n=19, SD=51.99) were statistically different (p<0.0001). (B) A statistical F-test to compare variance was performed, where PDI measurements for emulsification (n=18, SD=0.07031) and microfluidics (n=19, SD=0.0414) were statistically different (p<0.0315). Supplementary file3 (TIFF 4880 kb)
12248_2021_645_MOESM4_ESM.tiff
Figure S4. Correlation between particle size and zeta potential of iNPs on pro-inflammatory cytokine secretions. Data was obtained from experiments presented in Figure 5. Supplementary file4 (TIFF 103 kb)
12248_2021_645_MOESM5_ESM.tiff
Figure S5. Correlation between FRR of iNPs on pro-inflammatory cytokine secretions. Data was obtained from experiments presented in Figure 5. Supplementary file5 (TIFF 46 kb)
Rights and permissions
About this article
Cite this article
Truong, N., Black, S.K., Shaw, J. et al. Microfluidic-Generated Immunomodulatory Nanoparticles and Formulation-Dependent Effects on Lipopolysaccharide-Induced Macrophage Inflammation. AAPS J 24, 6 (2022). https://doi.org/10.1208/s12248-021-00645-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-021-00645-2