Abstract
Background
White mold disease, caused by Sclerotinia sclerotiorum the devastating pathogen, attacks green beans (Phaseolus vulgaris L.) and several crops worldwide. The present investigation was carried out to introduce some antagonistic microorganisms as novel antifungal substances to be an alternative and secure method to effectively control the disease.
Results
Three Streptomyces species, i.e., S. griseus (MT210913 “DG5”), S. rochei (MN700192 “DG4”), and S. sampsonii (MN700191 “DG1”) were isolated, biologically, molecularly characterized, and evaluated in vitro and in vivo. Molecularly, polymerase chain reaction (PCR) amplification and nucleotide sequencing were used to characterize the pathogen and bio-agents. PCR amplification of the pathogen and Streptomyces species (bioagents) exhibited amplicons of around 535 bp and 1300 bp, respectively. The nucleotide sequence analysis of the three Streptomyces spp. indicated that S. rochei was closely related to S. griseus, and both had a distance relationship with S. sampsonii. The evaluation of bioagents was carried out against S. sclerotiorum. Reduction percentages in the mycelial growth of the pathogen ranged between 60.17 and 52.30%, indicating that S. rochie gave the highest inhibition percent. Incorporations of Streptomyces spp. culture filtrate components into culture media proved that S. sampsonii was more efficient as a bioagent in reducing mycelial growth pathogen by 84.50%. The effectiveness of the bioagent volatile compounds inhibited the pathogen growth at a rate of 54.50-72.54%, respectively, revealing that S. rochei was the highest inhibitor followed by S. griseus. The parasitic activity of Streptomyces spp. upon S. sclerotiorum showed deformation, contraction, and collapse when observed by light and scanning electron microscopy. Molecular characterization of the 3 Streptomyces spp. revealed that S. griseus was closely related to S. sampsonii (96%), secondly ranked by S. rochei (93.1%). Viability and germination of pathogen sclerotia were reduced when they dipped into the Streptomyces spore suspensions for 10, 20, and 30 days. Application of the 3 Streptomyces spp. in the field proved a great potential to control the disease.
Conclusions
The results suggested that the 3 Streptomyces strains and their secondary metabolites can be potential biocontrol agents and biofertilizers for controlling S. sclerotiorum, the causative agent of bean white mold disease.
Similar content being viewed by others
Background
Beans are a staple food source of protein for billions of people worldwide and are suspected of infection by various diseases. The white rot (mold) caused by the destructive soil-borne pathogen Sclerotinia sclerotiorum (Lib.) de Bary, affects over 600 plant species including almost all dicotyledonous and some monocotyledonous plants (Shitou et al. 2020). It is considered one of the most important limiting factors in producing green beans worldwide (Zheng et al. 2019). In Egypt, bean (Phaseolus vulgaris L.) plants are grown usually in high moist and cool conditions. In both dry and green beans, the pathogen causes white mold disease leading to losses ranged from 30 to 100% depending on the favorable conditions (Mohamed and Atallah 2020). Such conditions (cold weather) seemed to be subsidizing factors for white rot infection, thus the growers have been battling large yield losses due to a disease caused by the fungus (Alsum et al. 2017). Globally, many challenges confront the management strategies of S. sclerotiorum due to the long-term persistence and tolerance of reproductive structures (sclerotia), a wide host range, and unpredictability of infection (Arfaoui et al. 2018). Due to its unique life cycle, the pathogen infects host plants either through aeciospores that can be discharged reluctantly upwards from apothecia into the air or by mycelium emerging from infected tissue and germinated sclerotia (Zheng et al. 2019). Synthetic pesticides are capable to effectively control plant diseases, but some fungicides result in severe environmental and health problems.
Out of all rhizosphere microbes, actinomycetes are considered as potential biocontrol agents because they exhibit many useful features. In this context, actinomycetes have been utilized to defend various plants against a wide range of phytopathogenic fungi and bacteria as well as human and insects (Zhao et al. 2019). The genus Streptomyces, a unique subgroup of actinomycetes bacteria, is best-known for their ability to produce a variety of weapons as bioactive secondary metabolites, i.e., antibiotics, antifungal, antibacterial, antioxidants, antiparasitic, anticancer, plant growth-promoting substances, insecticidal (Dongli et al. 2019; Zhao et al. 2019).
Recently, microbial antagonists such as Streptomyces spp. have been widely applied for the biocontrol of plant diseases (Katarzyna et al. 2018). The interaction of Streptomyces with the pathogenic fungi is usually due to the production of cell wall-degrading enzymes such as cellulases, chitinases, amylases, and glucanases. Also, the parasitic activity of Streptomyces species upon phytopathogenic fungi was demonstrated by light and scanning electron micrographs (Zamoum Miyada et al. 2017 and Kong et al. 2019). Furthermore, Streptomyces spp. are recognized for the production of an exceptionally large number of the natural bioactive secondary metabolites (70-80%) and volatile organic compounds “VOCs” well-known for their pharmaceutical or agrochemical applications (Salwan and Sharma 2020; Sharma et al. 2020). Nowadays, the bioformulation of VOCs derived from Streptomyces spp. available are used as bio-fungicide and inhibited hyphal growth against different soilborne pathogenic fungi (Sharma et al. 2020).
The highlights of the present investigation were conducted to determine the capabilities of parasitic activities, the potency of secondary metabolites, and volatile compounds of different Streptomyces isolates in vitro and in vivo to introduce alternative fungicides such as biological agents to manage S. sclerotiorum. For this reason, the utilization of antagonistic microorganisms and/or their novel antibacterial, antifungal, and antiviral substances becomes necessary as an alternative and secure method to effectively control the fungal and bacterial diseases of crops.
Methods
Collection, isolation, and purification of the pathogen (S. sclerotiorum)
Pure culture of S. sclerotiorum (green bean isolate) was obtained from infected green bean plants grown in the fields of Fayoum governorate, Egypt. The fungus was isolated from the sclerotia, formed on the pods of the diseased plants. The sclerotia were surface sterilized by sodium hypochlorite (0.5%) for 2-3 min, washed several times with sterilized distilled water, then dried between sterilized filter papers, and transferred directly into plate 9 cm contained Cook’s medium. The purified S. sclerotiorum isolate was identified according to Kora et al. (2003) and grown on Cook (1970), which contained 15 g glucose; 1 g K2HPO4; 0.5 g K2Cl; 0.5 g MgSO4.7H2O; 0.01 g FeSO4.7H2O; 2 g asparagine; 20 g agar/l liter distilled water). The fungus was kept onto the slant tubes medium at 5 °C for further experiments.
Molecular identification of the pathogen
Total genomic DNA extraction was carried out as described by Toda et al. (1999). Polymerase chain reaction (PCR) was conducted by the universal primers, i.e., forward primer ITS-1(5′-TCC GTA GGT GAACCT GCG G-3′) and reverse primer ITS-4 (5′-TCC TCC GCT TAT TGATAT GC-3′) as described by White et al. (1990). Amplification targeted is the sequence divergence within the ribosomal DNA internal transcribed spacer (rDNA-ITS) regions of the fungal. PCR was conducted according to the method described by Hayakawa et al. (2006) in GATC Biotech German Company (Under License of Sigma). PCR products were separated on 2 %agarose gel electrophoresis. The DNA ladder (100 bp) was also loaded in the gels to estimate the proper band size of amplified products and photographed. PCR cleanup was carried out to the PCR product, using GeneJET™ PCR Purification Kit (Thermo K0701). Finally, the sequencing of the PCR product was made, using ABI 3730xl DNA sequencer plus forward and reverse primers. The sequenced PCR product was confirmed, using NCBI (The National Center for Biotechnology Information) mega blast for the species identity. According to the percentage of homogencity between our isolate and identified isolates, the name of species was detected. After identification, the isolate has been deposited at the National Center for Biotechnology Information (NCBI) under the GenBank accession number (MT645495). The nucleotide sequences of the corresponding 16S rRNA gene were compared with 15 sequences of other fungus isolates from different geographical regions that were kept in the GenBank. Alignment program and phylogenetic tree were conducted to compute evolutionary distances for both sequences of nucleotides of the isolated fungus using DNAMan Ver.7.
Isolation and purification of Actinobacteria isolates
Twenty-one random rhizosphere soil samples were collected from different plant sources (Table 1). Also, samples were collected from phyllosphere plants and compost as well as saltwater from different Egyptian governorates. Isolation of Actinobacteria from soil samples was accomplished according to Lawrence (1956) and Hamada Eman (2006). One gram or milliliter of each collected sample was suspended in a flask containing 19-ml sterilized water to be a dilution of 1/20 and shaken in a rotary shaker (150 rpm). Afterward, two drops of each previously mentioned suspension were transferred separately from each flask to cylinder 25 ml in capacity containing 10 ml of phenol diluted by water (1:140) just to reduce the bacterial and fungal contamination. Subsequently, 0.1 ml of diluted soil suspension was streaked separately onto Petri dishes each containing 15 ml of solidified starch nitrate agar (StNA) medium contained (g/l); 20 g soluble starch; 2 g KNO3; 0.5 g NaCl; 1 g K2HPO4; 0.5 g MgSO4.7H2O; 3 g CaCO3; 0.001 g FeSO4.7H2O; 20 g agar; pH 7.2. Five plates were used for each sample, incubated at 28 °C, and examined periodically. After 3-5 days, colonies of actinomycetes on the plates were picked based on their morphological characteristics and purified on the same medium, then incubated for 7 days. Actinobacteria colonies were transferred to Petri dishes containing starch nitrate agar (ST.N.A.) medium supplemented with sodium propionate to suppress any bacterial or fungal contamination, then incubated at 25 °C (Crook et al. 1950; Waksman 1959).
Molecular characterization of Streptomyces isolates
Three Streptomyces species were characterized according to a molecular biological protocol by DNA extraction, amplification (PCR), and sequencing of amplification of 16S rDNA gene. DNA extraction of antagonist bacteria was implemented using Maxima Hot Start PCR Master Mix (Thermo K1051) according to the manufacturing instructions of GATC Biotech German Company. The target base pair DNA fragment generated from genomic DNA was amplified using forward primer “63f” (5′-CAGGCCTAACACATGCAAGTC-3′) and reverse primer “1387r” (5′-GGGCGGWGTGTACAAGGC-3′) as designated by Marchesi et al. (1998). PCR reaction was performed by a first initial denaturation at 95 °C for 10 min/one cycle, followed by denaturation 95 °C/30 s/35-cycles, annealing 57 °C/1 min/35-cycles, and extension 72 °C/1.5 min/35-cycles and final extension step at 72 °C/10 min/1-cycle. PCR amplicons were analyzed using 2.0% agarose gel electrophoresis in 1x TAE buffer staining with ethidium bromide. The amplified bands were visualized by UV illumination using a gel documentation system. The DNA Marker (100 bp) was also loaded in the gels to estimate the proper band size of amplified products and photographed. Then, PCR cleanup was carried out to the PCR product, using GeneJET™ PCR Purification Kit (Thermo K0701). Finally, the sequencing of the PCR products was achieved utilizing the ABI 3730xl DNA sequencer. The sequenced PCR product was confirmed, using NCBI (The National Center for Biotechnology Information) mega blast for the species identity. According to the percentage of homogencity between our Streptomyces isolates and identified isolates, names of the genus and species were determined as Streptomyces griseus, S. rochei, and S. sampsonii. After identification, species were deposited at the National Center for Biotechnology Information (NCBI) under the GenBank accession numbers, i.e., S. griseus (MT210913 “DG5”), S. rochei (MN700192 “DG4”), and S. sampsonii (MN700191 “DG1”). The nucleotide sequences of the corresponding 16S rRNA gene were compared with various sequences of other Streptomyces genera from different geographical regions that were kept in the GenBank. The nucleotide sequence of Bacillus subtilis (AJ276351.1) was used as out-group. For the alignment program, sequences retrieved were aligned by the most similar type strains obtained from the GenBank, using CLUSTAL W method by DNAMan Ver.7. While the phylogenetic tree was statistically tested, using the Maximum Like Hood method (Bootstrap phylogeny analysis with 1000 replications).
Determine the efficacy of Streptomyces spp. in vivo
This experiment was carried out at the greenhouse. Five seeds (Green bean “Phaseolus vulgaris L.” cv. Polista) were sown in pots (25 cm in diameter) containing 2.5 kg Peatmoss/sand. Inocula of S. sclerotiorum were prepared as the following: S. sclerotiorum was subcultured on Cook’s medium and kept in an incubator at 22 °C/15 days for the development of sclerotia. Five sclerotia were placed to infest each pot and 3 replicates (each replicate contained 4 pots)/treatment. Each treatment was applied to 60 plants. Concerning the application of Streptomyces species as bio-agents, the six species, i.e., Streptomyces griseus strain DG5 (MT210913), S. rochei strain DG4 (MN700192), S. sampsonii strain DG1 (MN700191), and Streptomyces spp. (no. 2, 27, and 30) were grown on a liquid starch casein medium for 7 days. All Streptomyces spp. were applied at the rate of 1/100-liter water (1 ml contain 15 × 106 cfu). Applications of Streptomyces were sprayed on 15-day-old plants 2 times (15 days interval). Data were recorded after 45 days of planting. All pots were irrigated periodically using the same amount of irrigation and fertilization.
Antifungal activity assays of Streptomyces spp.
Various techniques were carried out to determine the bioactivities of the 3 isolated Streptomyces spp. against S. sclerotiorum utilized dual culture techniques, mechanism of parasitism, culture filtrate, and volatile organic compounds (VOCs).
Detection of antagonistic activity by dual culture techniques
A dual culture technique was carried out to determine the antagonistic activity of Streptomyces spp. against the pathogen. Streptomyces spp. (S. griseus strain DG5 (MT210913), S. rochei strain DG4 (MN700192), and S. sampsonii strain DG1 (MN700191) were tested separately as antagonistic against S. sclerotiorum in vitro. Petri dishes (9 cm in diameter) each contained 15 ml of starch nitrate agar medium were inoculated on one side using a loopful of Streptomyces isolate and incubated at 28 °C for 4 days. The Petri dishes on opposite sides were inoculated by 5 mm mycelial disk from a 7-day-old culture of S. sclerotiorum. Plates inoculated only with S. sclerotiorum served as controls. Three replicate plates were used for each species. All inoculated plates were incubated at 22 °C for 7 days then examined. The percentages of reduction in mycelium growth of the pathogen were calculated and compared with control treatment as elaborated by Abd El-Moity (1985) as follows:
where X is the % of reduction in the growth of pathogenic fungus
G1 is the growth of the pathogenic fungus in the control plate
G2 is the growth of the pathogenic fungus in treated plates with antagonist
Evaluation of the parasitic mechanism of Streptomyces rochei
The parasitic activity of Streptomyces rochei upon S. sclerotiorum was confirmed by light and scanning electron micrographs (SEM). The slide technique provides a clear view of the interaction between Streptomyces spp. and S. sclerotiorum malformation, lyses, or parasitism. In this technique, a sterilized microscope glass slide was covered by a thin film sterilized diluted starch nitrate (0.1%). A loopful of S. rochei was stretched at one side, whereas the mycelial disk of S. sclerotiorum was inoculated at the other side of the microscope glass slide. Inoculated slides were placed in sterilized Petri dishes contained filter paper saturated with 10 ml of sterilized water just to maintain humidity, then incubated at (22 ± 2 °C). Slides were observed periodically using a light microscope with a fixed camera to examine the interaction between the pathogen and antagonist. The mycoparasitism of S. rochei on the mycelium of S. sclerotiorum was studied using SEM. The preparation of the samples was undertaken according to Loliam et al. (2013). Examinations were carried out, using the light microscopy (CHS, Olympus optical Co. Ltd.) and SEM (QUANTA FEG 250, Japan) in the National Research Center.
Impact of Streptomyces culture filtrates on mycelial growth suppression of S. sclerotiorum
Streptomyces species (S. griseus, S. rochei, and S. sampsonii) were grown separately in flasks containing 100 cm starch casein liquid (StC) medium consisted of 10 g soluble starch; 0.3 g casein; 2 g KNO3; 2 g NaCl; 2 g K2HPO4; 0.05 g MgSO4.7H2O; 0.02 g CaCO3; 0.01 g FeSO4 7H2O; distilled water 1000 ml, then adjusted to pH 7.2 and incubated at 28 °C for 7 days. Flasks were shaken on a rotary shaker (at 130 rpm). The culture broth was centrifuged (at 6000 rpm for 10 min), then filtered through a 0.2 μm. Each of the culture filtrates was mixed separately at a rate of 30% to sterilized Cook’s medium when reached 45 °C. Then the disks of S. sclerotiorum mycelium were placed onto the center of the plates containing Streptomyces culture filtrate. The diameter of mycelial growth was measured after 7 days of incubation at 22 °C. Three replicates were used for each genus of Streptomyces. The inhibition percentages of mycelial growth were determined as described by the equation of Gamliel et al. (1989) as the following:
where r is the colony radius of the pathogen on Cook’s medium incorporated with the culture filtrate of the Streptomyces
R is the colony radius of the fungal pathogen on Cook’s medium without Streptomyces culture filtrate.
Antifungal activity of volatile compounds produced by Streptomyces
The volatile organic compounds (VOCs) produced by Streptomyces spp. (S. griseus, S. rochei, S. sampsonii) was tested by the double-sealed plate method according to Fiddaman et al. (1993). Petri dishes (9 cm in diameter), each containing 15 ml of soluble starch casein agar medium was inoculated by Streptomyces spp., separately using loopful. After incubation at 28 °C for 3 days, a second Petri plate containing Cook’s medium was inoculated by a 5-mm desk of S. sclerotiorum and placed over the plate inoculated with the Streptomyces. The two dishes were sealed together by a parafilm and incubated at 22 °C. A Petri dish containing agar medium without bioagent was placed over the plate containing Cook’s medium, inoculated with the fungal pathogen served as control. % inhibition in radial growth of the tested fungus was measured after 7 days as previously mentioned (Gamliel et al. 1989).
Impact of Streptomyces spp. broth on the fungus sclerotia germination
Inhibition of sclerotia germination was conducted using Streptomyces spp. broth as the following: Mycelium of S. sclerotiorum was inoculated on Cook’s medium plates and then incubated at 22 °C. After 2 weeks, the developed sclerotia of S. sclerotiorum were harvested in the same sizes. Streptomyces spp. (S. griseus, S. rochei, and S. sampsonii) were grown in flasks containing 100 cm starch casein liquid media at 28 °C for 7 days. The sclerotia were soaked into individual streptomyces suspension (107 cfu/ml) for 10, 20, and 30 days. The soaked sclerotia were collected and cultured onto the center of the plates contained Cook’s medium. The plates were incubated at 22 °C for 7 days to test the germination of sclerotia and percentage of reduction in sclerotinia mycelial growth as described by Gamliel et al. (1989) as previously mentioned.
Impact of single or combined treatments of Streptomyces spp. under field (in vivo) conditions
A field experiment was carried out during the 2 successive seasons 2017/18 and 2018/19 at the farming plots located at Dahshur District, Giza Governorate, Egypt, where the soil was light sandy soil naturally infested with the pathogen. S. griseus strain DG5, S. rochei strain DG4, and S. sampsonii strain DG1 were applied individually or in combinations as a broth (containing cell-free extract, mycelia + spores). Three replicates were used for each treatment containing 100 bean seeds cv. Paulista. The plants were irrigated and fertilized. Corporal Max was used as a positive control (commercial fungicide produced by the Central Laboratory for Organic Agriculture). All Streptomyces spp. were applied at the rate of 1/100-liter water (1 ml contain 15 × 106 cfu). The application was carried out 4 times (15 days interval) on the plants after 30 days post sowing. Control treatment plants were sprayed by water similarly. The percentages of disease incidence were calculated and the percentages of reduction in white rot disease incidence were calculated after the development of natural disease symptoms on the control plants (60 days post sowing). Also, the total chlorophyll was measured according to Askar and Treptow (1993). The total yield was recorded for all pickings during the harvesting period.
Statistical analysis
Statistical analyses were performed by SPSS (2008), version 17.0, using the General Linear Model’s procedure. Significant differences among treatments were evaluated using Duncan’s multiple range test (P ≤ 0.05).
Results
Identification, purification, and molecular characterization of the pathogen
A pure culture of S. sclerotiorum was started from one sclerotia obtained from infected pods of green bean plants grown in the fields of Fayoum governorate, Egypt, and kept for further experiments. PCR amplification with the primers (ITS-1/ITS-4) resulted in an approximately 553-bp product within the internal transcribed spacer (rDNA-ITS) region of the fungal isolate tested (Fig. 1a). The sequence of the isolate was submitted to NCBI GenBank and gave an accession number MT645495. Phylogenetic tree of the Egyptian isolate S. sclerotiorum and other 15 isolates spread worldwide has been constructed based on nuclear ribosomal internal transcribed spacer the ITS rDNA (amplicons from primers ITS-1 and ITS-4) from a subset of isolates (Egyptian isolate and other 15 worldwide isolates) using DNAMan Ver.7. The result revealed that the nucleotide sequence identity of the ITS rDNA gene between the Egyptian isolate of S. sclerotiorum is closely related to some worldwide isolates and ranged from 98 to 100%. Meanwhile, the lowest homology percentage with other corresponding nucleotide sequences from different geographic origins was recorded at 93% as shown in Fig. 1b. The result indicated that PCR amplification of the ITS gene, using a specifically designed primer pair (ITS-1/ITS-4), followed by sequencing and phylogenetic analysis identified the isolate as S. sclerotiorum.
a Products of polymerase chain reaction amplification isolate using the universal primers ITS-1 and ITS-1. A product (amplicon) of 553 bp length was amplified in Egyptian isolate (Is). M = Leader and Is = isolated fungus. b Phylogenetic tree obtained from the alignment of nucleotide sequences of the Egyptian isolate and fifteen sequences from GenBank. All 15 sequences show their GenBank codes and countries of origin
Molecular characterization of the Streptomyces spp.
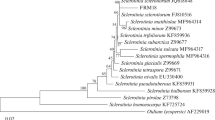
The 16S rRNA gene was successfully amplified by both primer pairs (63f and 1387r) from the 3 tested Streptomycetes, i.e., S. griseus strain (MT210913 “DG1”), S. rochei strain (MN700192 “DG1”) and S. sampsonii strain (MN700191 “DG1”). PCR amplification of its 16S rDNA gene sequence exhibited amplicons of around 1300 bp (Fig. 2a). The result indicated that the PCR technique, as a molecular method, has been extensively used for the identification of Streptomycetes species. It is favored over the traditional methods due to its high efficiency, accuracy rapidity specificity in amplification of the target gene. The alignment program and phylogenetic tree using the Maximum Like Hood method based on 16S rRNA gene sequences showed the close and distance relationships between the nucleotide sequences of Streptomycetes strains in the present study with the nucleotide sequences of available strains in the NCBI GenBank database as shown in (Table 2 and Fig. 2b). Concerning the 3 Egyptian Streptomycetes species, S. griseus strain had the highest sequence similarity (96%) with S. sampsonii strain, while the lowest similarity (93.1%) with S. rochei strain. However, the result indicated the divergence in nucleotide sequence between Streptomyces isolates with Bacillus subtilis out-group exceeded 82.9 %. Further, S. griseus MT210913 shared 100% sequence identity with S .griseus MT043745 in the GenBank. While S. griseus homology was 97% with S. luteus, S. zinciresistens (96.5%), and S. rochei (96.8%), which present in the NCBI GenBank. Homology of S. rochei MN700192 was (95.5%) with S. luteus, S. zinciresistens (95.4%), and S. rochei (95.7%), which present in the NCBI GenBank. Whereas S. sampsonii MN700191 had 99% homology with each of S. sampsonii MK564728 and S. sampsonii MH727710 in the GenBank. The aforementioned result indicated that S. griseus MT210913 was closely related to S. sampsonii MN700191 (96%), secondly ranked by S. rochei MN700192 (93.1%).
a Products of polymerase chain reaction amplification isolates using the universal primers 63-forward and 1387-reverse. Products (amplicons) of 1300 bp length were amplified from the three isolates of Streptomyces (Is). M = Leader, 1,2 and 3 = isolated S. sampsonii, S. rochei, and S. griseus. b Phylogenetic tree based on 16S rRNA gene sequence of S. rochei, S. griseus, and S. sampsonii with the reference Streptomycetes 16S rRNAgene available in the GenBank
Screening of Streptomyces spp. antifungal activity against S. sclerotiorum under greenhouse conditions
Out of 24 Streptomyces spp. isolated from the plant rhizosphere, only 6 isolates proved to be effective as antagonistic agents against the devastating pathogen (Sclerotinia sclerotiorum “Lib.” de Bary) infecting green bean. Among the 6 tested isolates, 3 turned to be more efficient. Data presented in Table 3 revealed that the percentages of disease incidence were reduced to 16.67, 10, and 0.0%, when green bean plants were treated by S. rochei, S. griseus, and S. sampsonii, respectively. Among the 3 Streptomyces spp., S. sampsonii was the most effective bio-agent, followed by S. griseus and S. rochei reached 100, 69.6, and 49.9%, respectively, in the reduction of the disease. The results indicated that the 3 Streptomyces spp. might have many useful and vital bioactive compounds, which act as a wide variety of modes of antifungal against S. sclerotiorum the causal agent of white disease of bean.
In vitro evaluation of the antifungal activity of Streptomyces spp.
In this context, 4 techniques, i.e., dual culture techniques, culture filtrate, mechanism of parasitism, and volatile organic compounds (VOCs) were applied as the following:
In vitro inhibitory potential of Streptomyces spp.
Results of dual culture technique in Table 4 and Figs. 3, 4, and 5 indicated that all Streptomyces spp. significantly inhibited linear growth of pathogenic fungus, S. sclerotiorum compared with control treatment. Percentages of growth reduction of S. sclerotiorum ranged between 60.17 and 52.30% (Fig. 3). S. rochie showed the highest percentage of reduction in mycelial growth. Obtained results displayed that the plate assays are an indication of whether strains can act as biocontrol agents.
Scanning electron micrographs of the deformation of the fungal mycelium. (a) Untreated mycelium of S. sclerotiorum, (b) Bioagent S. rochei induced pores in the mycelium of S. sclerotiorum, and (c) S. rochei induced swollen, deformation, distortion, and lysis of the pathogen mycelium as well as it is grown inside the hyphae
On the other hand, the antagonistic activity of the Streptomyces spp. upon S. sclerotiorum was demonstrated by light and scanning electron micrographs (SEM). Light and scanning electron micrographs showed that S. rochei was colonized on S. sclerotiorum mycelium as well as penetrated and grown inside the hyphae. The growth of S. sclerotiorum could be suppressed by S. rochei appeared to alter the structure of S. sclerotiorum mycelia by lysing hyphal tips that caused swelling, pores, irregular membrane border and shrinkage of the hyphal tip as well as grown inside hyphae of the pathogen (Figs. 4 and 5). This result supports the potency and efficacy of S. rochei as a bioagent, able to produce secondary metabolites, and penetrated as well as destroy and lysis the hyphae of S. sclerotiorum, the devastating soil-borne fungal pathogen.
Antifungal activity of the culture filtrate
The secondary metabolites produced by these 3 Streptomyces spp. were extracted and partially purified. Suppression of mycelial growth of S. sclerotiorum by culture filtrate of Streptomyces spp. proved that they secreted high antifungal activity bioactive secondary metabolites expressed in vitro significant reduction in the growth of pathogenic fungus than in the control treatment. Data in Table 4 showed that the highest percentage of reduction in mycelial growth (84.50%) was obtained when the culture filtrate of S. sampsonii was used (Fig. 6).
Inhibitory effects of VOCs produced by Streptomyces spp. on mycelium growth of S. sclerotiorum
Growth inhibition of S. sclerotiorum was tested by the 3 Streptomyces spp. The growth inhibition of the pathogen was ranged between 54.50 and 72.54%. S. rochei showed the highest inhibition percentage in mycelial growth, followed by S. griseus (64.70%) as shown in Table 4. This result indicated that S. sclerotiorum was suppressed but not completely killed after exposure to volatile organic compounds (VOCs) of Streptomyces spp. Test of growth inhibition in the sealed plates method exhibited that all the Streptomyces spp. were able to produce secondary metabolites such as antifungal or VOCs inhibited and deformed the growth of S. sclerotiorum mycelium (Figs. 7 and 8).
Impact of Streptomyces spp. broth on the fungus sclerotia germination
The 3 antagonistic Streptomyces spp. inhibited significantly the viability and sclerotia germination of S. sclerotiorum compared with the control. Also, the radial mycelial growth of the germinated sclerotia was reduced after soaking into the spore suspensions of Streptomyces spp. separately for 10, 20, and 30 days (Fig. 9). The highest inhibition in mycelial growth of the germinated sclerotia was obtained with S. sampsonii by 90.32, 100, and 100%, when soaked for 10, 20, and 30 days, respectively. In this trial, S. sampsonii, followed by S. griseus proved to be a more inhibitory effect than S. rochei, which reduced mycelial growth to 35.32, 42.38, and 90.32%, respectively after same exposure periods (Table 5).
Application of the three Streptomyces spp. under the field conditions (in vivo)
To check the ability of selected 3 Streptomyces spp. to control the white rot disease caused by S. sclerotiorum, bioagents were applied on bean plants grown in the field as inocula in the form of suspension (containing cell free extract, mycelia+ spores). The tested Streptomyces spp. exhibited a significant reduction of S. sclerotiorum in the field. These results indicated a good correlation between in vitro inhibitory action and in vivo activity. Data in Table 4 showed that all Streptomyces spp. either in a single form or mixture resulted in a significant reduction in disease incidence compared with check treatment. The lowest intensity of disease incidence was achieved by S. rochei DG4 (12%), followed by S. griseus DG5 (13%) and (25%) in check treatment (Table 6). Non-significant differences in percentages of disease incidence and percentages of disease reduction were noticed when S. griseus DG5 and S. rochei DG4 were used separately. A clear significant effect was recorded when the mixtures of Streptomyces spp. were used in comparison with using any single isolate. The highest percentages of disease reduction were noticed when the mixtures of DG4 + DG5 or DG1 + DG5 were used, recording 76.00 and 77.33%, respectively. Clear significant deference was noticed when the mixture of the three Streptomyces spp. (DG1, DG4, and DG5) compared with any other treatment. Using S. rochei (DG4) alone, it proved to be a good protective bioagent and increased the chlorophyll content and pods yield. Also, both of S. rochei and S. griseus played an important role in controlling S. sclerotiorum due to their various modes of action, i.e., hyperparasitism, bioactive metabolites, and volatiles.
Total chlorophyll and yield in the treated plants were measured and compared with the control treatment. Obtained data in Table 5 indicated that all treatments resulted in a reduction of disease incidence and increase in chlorophyll content as well as pods yield than the control. The increase of chlorophyll means increasing in photosynthesis which is reflected as a rise in enzyme level and sugars consequently as well as flourishing in the plant growth and total yield. The relation between reduction in disease incidence and increase of total chlorophyll and yield was very clear when using the most effective treatment (DG1 + DG4 + DG5) in contrast with the control. Further, mixing different isolates increases the scope of the mode of action consequently increases the efficacy of the treatment. Thus, the use of biological control agents (Streptomyces spp.) is considered as an alternative and sustainable strategy to control S. sclerotiorum.
Discussion
White mold disease incited by S. sclerotiorum is notably challenging. Thus, the present research aimed to isolate, characterize, and introduce new useful biological agents to manage the disease. Based on the present investigation findings, Streptomyces spp. revealed antifungal activity against S. sclerotiorum under in vitro conditions as well as successful biocontrol in the greenhouse experiment.
Regarding characterization of the pathogen S. sclerotiorum, the results of PCR amplification (553-bp product), sequence, and phylogenetic tree of the Egyptian isolate S. sclerotiorum shared with other 15 isolates are in agreement with Prova et al. (2018) as they demonstrated that the PCR amplified 564 bp product in the rDNA-ITS region of the fungal S. sclerotiorum isolated from Hyacinth Bean (Lablab purpureus). Also, a consistent trend indicated by several investigators highlights the effectiveness of the nuclear ribosomal internal transcribed spacer (ITS) region sequences located between the nuclear small- and large-subunit rRNA genes in species identification (Prova et al. 2018).
Concerning molecular characterization of the Streptomyces spp., the 16S rRNA gene sequences (1300 bp) were checked and aligned, then statistically tested using the Maximum Like Hood method. The aforementioned results indicated that the three isolated Streptomyces spp. were closely related together; they had close and/or distant relationships with the nucleotide sequences of available strains in the NCBI GenBank database. These results are in harmony with those observed by Marchesi et al. (1998); Anderson and Wellington (2001); and Rintala et al. (2001), who demonstrated that PCR detection had many features in Streptomycetes species identification compared with the morphological, biochemicals, and physiological methods. Further, in consensus with the obtained results, various workers elaborated that the phylogeny tree in the Streptomycetes based on 16S rRNA gene sequences proved to be a useful characterization method for determining the relationships among Streptomycetes species (Sharma et al. 2020; Yu et al. 2020).
In a preliminary experiment to evaluate antifungal activity against S. sclerotiorum under greenhouse conditions, the results proved that the 3 Streptomyces spp. might have antifungal activity against S. sclerotiorum the causal agent of white disease of the bean. A similar trend was observed by those obtained by Zamoum Miyada et al. (2017) who stated that Streptomyces spp. are regarded as significant biological resources, due to their secondary metabolites; these antimicrobial compounds play roles in protecting plants against different pathogens. Furthermore, Srivastava et al. (2015) revealed that S. rochei strain SM3 suppressed mycelial growth and disease development of S. sclerotiorum at an approximate rate of 74%. In vitro antagonistic activity trials of Streptomyces spp. were implemented to ensure the greenhouse results and prove antifungal bioactive compounds to be applied in the field as follows:
In vitro inhibitory potential of Streptomyces spp.
Obtained results, when using the dual culture technique, showed that Streptomyces spp. could act as biocontrol agents. Such results are consistent with those by Dongli et al. (2019) who declared that Streptomyces spp. NEAU-S7GS2 induced a significant inhibitory activity against the mycelial growth of the phytopathogenic fungi S. sclerotiorum and Rhizoctonia solani affecting soybean (Glycine max) plants at a rate of 99.1 and 65.3%, respectively, compared with the control. Moreover, Evangelista-Martínez et al. (2020) showed that the antagonistic activity of the Streptomyces isolate CACIS-1.5CA was similar to the commercial strain Streptomyces lydicus WYEC 108 against the pathogenic fungi; Colletotrichum sp., Alternaria sp., A. sp., Botrytis sp., Rhizoctonia sp., and Rhizopus sp. with percentages ranging from 30 to 63%. Furthermore, Yu et al. (2020) showed that S. triticiradicis caused significant inhibitory effects against 4 phytopathogenic fungi, i.e., Colletotrichum orbiculare, Corynespora cassiicola, S. sclerotiorum, and Exserohilum turcicum with an inhibition rate ranging from 48.4 to 69.0%. Further, many investigators stated that Streptomyces spp. caused a reduction in mycelial growth of S. sclerotiorum due to antifungal substances produced by Streptomyces isolates as bioactive compounds and secondary metabolites such as enzymes and antibiotics (Baharlouei et al. 2013).
Examination of the antagonistic activity of the Streptomyces spp. upon S. sclerotiorum using light and scanning electron micrographs (SEM) supported the potency and efficacy of S. rochei as bioagent, which was able to produce secondary metabolites, and penetrated as well as destroyed and lysis the hyphae of the fungal pathogen. Similary, other investigators (Yang et al. 2019) working with Aspergillus flavus examined severe destruction of cell membrane integrity, irregular morphological shapes after exposure to VOCs produced by Streptomyces philanthi RM-1-138 and S. alboflavus TD-1. Moreover, Dongli et al. (2019) mentioned that the SEM and light microscopy examination revealed that Streptomyces NEAU-S7GS2 led to cell wall degradation in S. sclerotiorum, indicating its production of extracellular lysis enzymes. Similarly, Evangelista-Martínez et al. (2020) examined the morphological changes as irregular membrane border, deformation, shrinkage, and collapsed conidia of postharvest fungal pathogens such as Alternaria sp., Botrytis sp., Rhizoctonia sp., and Colletotrichum musae after exposure to bioactive extract of Streptomyces sp. CACIS-1.5CA.
Antifungal activity of the culture filtrate
The applications of Streptomyces spp. culture filtrate in the present experiments demonstrated their ability to reduce mycelial growth of S. sclerotiorum due to their secretion of secondary metabolites. It could be attributed to the fact that Streptomyces spp. produce a variety of extracellular hydrolytic enzymes (degrading cell wall enzymes such as extracellular hydrolytic enzymes) (degrading cell wall enzymes such as cellulases, hemicellulases, chitinases, amylases, and glucanases) and antibiotics as secondary metabolites (Elleuch et al. 2010). Further, Kanini et al. (2013) tested various Streptomyces isolates from diverse Greek habitats for their antifungal activity against the common phytopathogenic fungus Fusarium oxysporum f. sp. lycopersici affecting tomato. They mentioned that the isolate was characterized molecularly as S. rochei ACTA1551 strongly suppressed the fungal growth when examined in antagonistic bioassays in vitro. Also, S. rochei secreted antimicrobial compounds able to inhibit the growth of all common microbial indicators (Escherichia coli, Bacillus subtilis, Pseudomonas fluorescens, and Saccharomyces cerevisiae). Additionally, Li et al. (2018) stated that the antifungal activity of S. sampsonii crude extract inhibited the hyphal growth and destroyed fungal cell walls of 4 different pathogenic fungi, i.e., Cylindrocladium scoparium, Cryphonectria parasitica, Neofusicoccum parvum, and Fusarium oxysporum). Nevertheless, many researchers recorded that the Actinobacteria proved to be great importance in the field of biotechnology, as producers of benefits and bioactive secondary metabolites, i.e., antifungal, insecticidal, antibacterial, enzymes such as chitinase, antibiotics, and antihelminthic activities with extensive industrial, medical use, and as a tool for eco-friendly management in agricultural applications of plant pathogens (Katarzyna et al. 2018; Salwan and Sharma 2020; and Sharma et al. 2020).
Inhibitory effects of VOCs produced by Streptomyces spp. on mycelium growth of S. sclerotiorum
All of the tested Streptomyces spp. showed more or less antifungal activity against the pathogen both on Petri plate and pot assays. In the same respect, Wu et al. (2015) revealed that volatile organic compounds produced by Streptomyces spp. were reported to induce growth abnormalities in S. sclerotiorum. The produced VOCs by S. albulus NJZJSA2 not only inhibited S. sclerotiorum mycelial growth by 100% but also significantly inhibited the sclerotia germination. Similarly, Cordovez et al. (2015), in their studies on Streptomyces spp. over a disease-suppressive soil, identified Actinobacteria as the most dynamic phylum in a soil suppressive to the fungal root pathogen Rhizoctonia solani. All Streptomyces strains were able to significantly retard the growth of R. solani, an economically important soil-borne fungal pathogen of many crops. VOCs produced by S. canus strains (W47 and W214) were the most inhibitory of radial hyphal growth at a rate of 57 and 41%, respectively, and significantly enhanced plant shoot and root biomass. On the other hand, some investigators discussed that this phenomenon depends on the diversity of the pathogens that have different levels of resistance or susceptibility to VOCs (Gotorvila et al. 2017).
Impact of Streptomyces spp. broth on the fungus sclerotia germination
Application of the 3 antagonistic Streptomyces spp. inhibited significantly the viability and sclerotia germination, which reflected on the growth of the mycelium. The result in consensus with Errakhi et al. (2007), who stated that the highest inhibition in sclerotial germination of Sclerotium rolfsii was obtained by streptomyces isolate J-2 by 93 and 88%, when applied as biomass and filtrate culture, respectively. While the lowest inhibition in sclerotial germination was obtained with the isolate B-5 which was 47, 44, and 17% with biomass inocula, culture filtrate, and spore suspension, respectively. These results are consistent with those recorded by Dongli et al. (2019) who revealed that Streptomyces sp. NEAU-S7GS2 completely inhibited sclerotia germination S. sclerotiorum. Furthermore, Smoliñska and Kowalska (2018) indicated that the antibiosis is likely to be a major mechanism employed by Streptomyces spp. on control S. sclerotiorum sclerotia. Certain bacterial genera can be used as a biological control against S. sclerotiorum by inhibiting the germination of ascospores by the production of antimicrobial substances or direct growth on the ascospores (Huang and Erickson 2008).
Application of the three Streptomyces spp. under the field conditions (in vivo)
Field experiments clearly showed that Streptomyces spp. had the potential to be used as highly effective biocontrol agents due to their various beneficial properties against White mold disease and contribution in enhancement of plant growth as well as they are inexpensive, long lasting, and safe toward the environment and living organisms. Relevant to this work, the results obtained by several investigators, highlighted the contribution of Streptomyces genera to plant growth and protection. Another role of Streptomyces has been revealed by Jogaiah et al. (2016) who coated the seeds of the pearl millet with S. griseus SJ_UOM-07-09 and Streptosporangium roseum SJ_UOM-18-09 to protect the plants against downy mildew disease caused by Sclerospora graminicola. The two strains possessed a direct anti-mildew activity in inhibiting the sporangial formation, zoospore release from sporangia, and enhanced the induced resistance to protect plants from disease. Application of (DG1 + DG4 + DG5) treatment led to a reduction in disease incidence and noticeable total chlorophyll and yield increase compared to control. Furthermore, mixing different isolates improved treatment efficacy. Thus, Streptomyces spp. proved to be as an alternative and sustainable strategy to control S. sclerotiorum. This result supports the earlier explanations that the mixtures of bio-agents with different mechanisms in disease control will have an additive efficient impact in improving disease control compared to their separate application (Huang and Erickson 2008).
Conclusions
Obtained findings demonstrated that Streptomyces spp. (S. griseus, S. rochei, and S. sampsonii) had a great potential to be used as biocontrol agents against S. sclerotiorum in both in vitro and in vivo. The 3 isolates seemed to be a promising contribution to the development of new methods for the management of plant diseases and reducing the deleterious impacts of using chemical pesticides in the future.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- PCR:
-
Polymerase chain reaction
- rDNA-ITS:
-
Ribosomal DNA internal transcribed spacer
- SEM:
-
Scanning electron microscope
- S. :
-
Sclerotinia
- Spp.:
-
Species
- StNA:
-
Starch nitrate agar
- S:
-
Streptomyces
- VOCs:
-
Volatile organic compounds
References
Abd El-Moity TH (1985) Effect of single and mixture of Trichoderma harzianum isolates on controlling three different soil borne pathogens. Egypt J Micrbiol Special Issue pp 111–120
Alsum BA, Elsheshtawi M, Elkahky MT, Elgorban AM, Bakri M, Alkhulafi M (2017) Management of Sclerotinia white rot of beans with antagonistic microorganisms. J Anim Plant Sci 27(2):542–549
Anderson AS, Wellington EMH (2001) The taxonomy of Streptomyces and related genera. Int J Syst Evol Microbiol 51:797–814
Arfaoui A, Hadrami AE, Daayf F (2018) Pre-treatment of soybean plants with calcium stimulates ROS responses and mitigates infection by Sclerotinia sclerotiorum. Plant Physiol Biochem 122:121–128. https://doi.org/10.1016/j.plaphy.2017.11.014
Askar A, Treptow H (1993) Analytical methods. In: Askar A, Treptow H (eds) Quality assurance in tropical fruit processing. Springer-Verlag, New York, pp 27–28
Baharlouei A, Sharifi-Sirchi GR, Shahidi Bonjar GH (2013) Biological control of Sclerotinia sclerotiorum (oilseed rape isolate) by an effective antagonist Streptomyces. Afr J Biotechnol 10(30):5785–5794, http://www.academicjournals.org/AJB. https://doi.org/10.5897/AJB10.2017
Cook RC (1970) Physiological aspects of sclerotium growth in Sclerotinia sclerotiorum. Trans Brit Mycol Soc 54:361–365
Cordovez V, Carrion VJ, Etalo DW, Mumm R, Zhu H, van Wezel GP Raaijmakers JM (2015) Diversity and functions of volatile organic compounds produced by Streptomyces from a disease-suppressive soil. Front Microbiol. 6:1081.doi: https://doi.org/10.3389/fmicb.2015.01081
Crook P, Carpenter CC, Klens PF (1950) The use of sodium propionate in isolating actinomycetes from soil. Science 112:656
Dongli L, Rui Y, Yansong F, Xiangjing W, Zhang J, Wensheng X (2019) Antifungal, plant growth-promoting, and genomic properties of an endophytic actinobacterium Streptomyces sp. NEAU-S7GS2. Frontiers in. Microbiology 10:2077
Elleuch L, Shaaban M, Smaoui S, Mellouli L, Karray-Rebai I, Fguira LF, Shaaban KA, Laatsch H (2010) Bioactive secondary metabolites from a new terrestrial Streptomyces sp.TN262. Appl Biochem Biotechnol 162:579–593
Errakhi R, Bouteau F, Lebrihi A, Barakate M (2007) Evidences of biological control capacities of Streptomyces spp. against Sclerotium rolfsii responsible for damping-off disease in sugar beet (Beta vulgaris L.). World J Microbiol Biotechnol 23:1503–1509. https://doi.org/10.1007/s11274-007-9394-7
Evangelista-Martínez Z, Contreras-Leal EA, Corona-Pedraza LF, Gastélum-Martínez E (2020) Biocontrol potential of Streptomyces sp.CACIS-1.5CA against phytopathogenic fungi causing postharvest fruit diseases. Egyptian J Biol Pest Control 30:117
Fiddaman PJ, O’Neill TM, Rossall S (1993) Screening of bacteria for the suppression of Botrytis cinerea and Rhizoctonia solani on lettuce (Lactuca sativa) using disk bioassays. Annals Appl Biol 137(3):223–235
Gamliel A, Katan J, Cohen F (1989) Toxicity of chloronitrobenzenes to Fusarium oxysporum and Rhizoctonia solani as related to their structures. Phytopathology 17:101–105
Gotorvila A, Teixidó N, Di FA, Usall J, Ugolini L, Torres R, Mari M (2017) Antifungal effect of volatile organic compounds produced by Bacillus amyloliquefaciens CPA-8 against fruit pathogen decays of cherry. Food Microbiol. 64:219–225
Hamada Eman SM (2006) Utilization actinomycees as effective biocontrol agent in controlling Rhizoctonia pathogen attack potatoes. M.Sc. Thesis, Fac. Science., Benha Univ., 116 p.
Hayakawa T, Toda T, Ping Q, Mghalu JM, Yaguchi S, Hyakumachi M (2006) A new subgroup of Rhizoctonia AGD, AG-D III, obtained from Japanese zoysia grass exhibiting symptoms of a new disease. Plant Dis 90:1389–1394
Huang HC, Erickson RS (2008) Factors affecting biological control of Sclerotinia sclerotiorum by fungal antagonists. J. Phytopathology 156:628–634
Jogaiah S, Kurjogi M, Govind SR, Huntrike SS, Basappa VA, Phan T (2016) Isolation and evaluation of proteolytic actinomycete isolates as novel inducers of pearl millet downy mildew disease protection. Sci Rep 6:30789. https://doi.org/10.1038/srep30789
Kanini GS, Efstathios AK, Alexandros LS, Amalia DK (2013) Streptomyces rochei ACTA1551, an indigenous Greek isolate studied as a potential biocontrol agent against Fusarium oxysporum f. sp. lycopersici. BioMed Res Int. https://doi.org/10.1155/2013/387230
Katarzyna JK, Aleksandra RM, Adam G, Joannam Z, Jolanta S (2018) Secondary metabolites of actinomycetes and their antibacterial, antifungal and antiviral properties. Polish J Microbiol 67(3):259–272
Kong D, Xia W, Ju N, Guoqing N (2019) Regulation of antibiotic production by signaling molecules in streptomyces. Front Microbiol 10. https://doi.org/10.3389/fmicb.2019.02927
Kora C, McDonald MR, Boland JG (2003) Sclerotinia rot of carrot: an example of phenological adaptation and bicyclic development by Sclerotinia sclerotiorum. Plant Dis. 87(5):456–470
Lawrence CH (1956) A method of isolating Actinomycetes from scabby potato tissue and soil with minimal contamination. Can. J. Bot. 34:44–47
Li S, Zhang B, Zhu H, Zhu T (2018) Cloning and expression of the chitinase encoded by ChiKJ406136 from Streptomyces sampsonii (Millard&Burr) Waksman KJ40 and its antifungal Effect. Forests. 9:699. https://doi.org/10.3390/f9110699
Loliam B, Morinaga T, Chaiyanan S (2013) Biocontrol of Pythium aphanidermatum by the cellulolytic actinomycetes Streptomyces rubrolavendulae S4. Sci Asia 39:584–590
Marchesi JR, Sato T, Weightman AJ, Martin TA, Fry JC, Hiom SJ, Wade WG (1998) Design and evaluation of useful bacterium-specific PCR primers that amplify genes coding for bacterial 16S rRNA. Appl Environ Microbiol 64(2):795–799
Miyada Z, Yacine G, Nasserdine S, Mathieu F, Zitouni A (2017) Development of formulations based on Streptomyces rochei strain PTL2 spores for biocontrol of Rhizoctonia solani damping-off of tomato seedlings. Biocontrol Sci Technol 27(6):723–738
Mohamed ASA, Atallah O (2020) Controlling common bean white mould caused by Sclerotinia sclerotiorum (Lib.) de Bary. Zagazig J Agric Res 47(1):101–117
Prova A, Akanda A, Islam S, Hossain M (2018) Characterization of Sclerotinia sclerotiorum, an emerging fungal pathogen causing blight in hyacinth bean (Lablab purpureus). Plant Pathol J 34(5):367–380. https://doi.org/10.5423/PPJ.OA.02.2018.0028
Rintala H, Nevalainen A, Rönkä E, Suutari M (2001) PCR primers targeting the 16S rRNA gene for the specific detection of Streptomycetes. Mol Cell Probes 15(6):337–347
Salwan R, Sharma V (2020) Bioactive compounds of Streptomyces: biosynthesis to applications In: Studies in Natural Products Chemistry, Rahman A (ed.) Vol. (67), 1st Ed. Elsevier 467–491, Chennai, India
Sharma V, Sharma A, Malannavar A, Salwan R (2020) Molecular aspects of biocontrol species of streptomyces in agricultural in: molecular aspects of plant beneficial microbes in agriculture. Sharma V, Richa Salwan R, Tawfeeq Al-Ani LK (eds.), Academic Press pp 89–109, Chennai, India
Shitou X, Yan X, Ryan H, Julia Z, Lei Q, Xin L (2020) The notorious soilborne pathogenic fungus Sclerotinia sclerotiorum: an update on genes studied with mutant analysis. Pathogens 9:27. https://doi.org/10.3390/pathogens9010027
Smoliñska U, Kowalska B (2018) Biological control of the soil-borne fungal pathogen Sclerotinia sclerotiorum a review. J Plant Pathol 100:1–12. https://doi.org/10.1007/s42161-018-0023-0
Spss (2008). Statistical Package For Social Science. Version 17.0.0 Spss Corporation
Srivastava S, Patel S, Singh HB, Sinha A, Sarma K (2015) Streptomyces rochei SM3 induces stress tolerance in chickpea against Sclerotinia sclerotiorum and NaCl. J Phytopathol. 163:583–592. https://doi.org/10.1111/jph.12358
Toda T, Hyakumachi M, Suga H, Kageyama K, Tanaka A, Tani T (1999) Differentiation of Rhizoctonia AG-D isolates from turfgrass into subgroups I and II based on rDNA and RAPD analysis. Eur J Plant Pathol 105:835–846
Waksman SA (1959) The actinomycetes Vol. 1 Nature, occurrence and activities. The Williams & Wilkins Co, Baltimore, p 327
White TJ, Bruns T D, Lee SB, Taylor JW (1990). Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: PCR Protocols: A Guide to Methods and Applications, by M. A. Innis, D. H. Gelfand, J. J. Sninsky and T. J. White,(eds.) pp. 315-322. Academic Press, New York
Wu Y, Yuan J, Yaoyao E, Raza W, Shenc Q, Huang Q (2015) Effects of volatile organic compounds from Streptomyces albulus NJZJSA2 on growth of two fungal pathogens. J. Basic Microbiol 55:1104–1117
Yang M, Lu L, Pang J, Hu Y, Guo Q, Li Z, Wu S, Liu H, Wang C (2019) Biocontrol activity of volatile organic compounds from Streptomyces alboflavus TD-1 against Aspergillus flavus growth and aflatoxin production. J Microbiol 57(5):396–404. https://doi.org/10.1007/s12275-019-8517-9
Yu Z, Han C, Yu B, Zhao J, Yan Y, Huang S, Liu C, Xiang W (2020) Taxonomic characterization, and secondary metabolite analysis of Streptomyces triticiradicis sp. nov.: A novel actinomycete with antifungal activity. Microorganisms 8:77. https://doi.org/10.3390/microorganisms8010077
Zhao J, Han L, Yu M, Cao P, Li D, Guo X, Liu Y, Wang X, Xiang W (2019) Characterization of Streptomyces sporangiiformans sp. nov., a Novel Soil actinomycete with antibacterial activity against Ralstonia solanacearum. Microorganisms 7:360. https://doi.org/10.3390/microorganisms7090360
Zheng W, Lu-Yue M, Jun C, Yu-Long L, Li-Na D, Ke-Ming Z, Yan-Hua Y, Xiao-Li T (2019) Recent advances in mechanisms of plant defense to Sclerotinia sclerotiorum. Front Plant Sci 10. https://doi.org/10.3389/fpls.2019.01314
Acknowledgements
We would like to acknowledge Dr. Ahmed Abd-El Aziz Kheder, who provided a profound contribution in the part of molecular biology.
Funding
The authors declare that the current study is self-funded.
Author information
Authors and Affiliations
Contributions
GG suggested the plan of study and followed up on the carried-out work in addition to scripting the manuscript. DG executed the plan of work, analyzed the output data, and contributed in writing the manuscript. AA assisted in writing the manuscript. MR reviewed the manuscript. NS and TA helped in the design of the work plan until they both, sadly, passed away. All authors except NS and TA (passed away) read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gebily, D.A.S., Ghanem, G.A.M., Ragab, M.M. et al. Characterization and potential antifungal activities of three Streptomyces spp. as biocontrol agents against Sclerotinia sclerotiorum (Lib.) de Bary infecting green bean. Egypt J Biol Pest Control 31, 33 (2021). https://doi.org/10.1186/s41938-021-00373-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s41938-021-00373-x