Abstract
Background
The extraction of DNA from skeletal remains with good quality and quantity is often challenging for the ability to generate DNA typing. Previous studies demonstrated the DNA extraction with total demineralization from fresh teeth and bones; however, the application in old skeletal remains has been less performed. To obtain good quality and high yield of DNA amount extracted from skeletal remains, the objective of this study was focused on exploring the factors influencing the total demineralization process to obtain developing effective methods.
Results
The concentration of EDTA was found to significantly enhance calcium chelation from the bone while pH of EDTA solution, incubation temperature, incubation time, and volume of EDTA solution were not significant. The optimal condition of total demineralization obtained from Placket-Burmann results represented good-quality DNA and the highest concentration of extracted DNA yield. Subsequently, the STR typing in some bone specimens processed by total demineralization process prior to DNA extraction was improved.
Conclusions
EDTA concentration was a key influencing factor on the total demineralization process to chelate calcium from human skeletal remains. The total demineralization process in old bone specimens probably improved the STR profiles.
Similar content being viewed by others
Background
One of the crucial parts of the personal identification process from skeletal remains is based on the utility of the DNA profiles obtained from the bones or teeth (Latham and Miller 2019). Many techniques such as polymerase chain reaction (PCR), restriction fragment length polymorphism (RFLP), and gel electrophoresis have been extensively used to establish DNA profiling (Renneberg et al. 2017).
A large amount of good-quality extracted DNA from the bones was comparatively complicated and challenging since many factors affected DNA extraction from skeletal remains. First, environmental conditions were the most important factors affecting bone structure and composition. Even though DNA was protected by the hard structure of the bone, it would be degraded by environmental factors such as temperature, UV, humidity, and microorganisms when left in particular conditions for a long time (Samsuwan et al. 2018). Second, parts of the skeletons for DNA extraction were correlated with the success rate of DNA detection from the skeletal remains. Promising parts of the skeletons such as the femur, teeth, tibia, and fibular that would deliver the best success rate for DNA detection were employed (Milos et al. 2007; Mundorff et al. 2009, 2013). Finally, the bones consist of osteocyte and the extracellular matrix surrounding DNA. Seventy-five percent of the extracellular matrix was in the inorganic components consisting of calcium phosphate in the form of hydroxyapatite. This substance supports the hard structure of the bone, resulting in limited DNA extraction (Loreille et al. 2007). Hence, several researchers have discovered the appropriate techniques to eliminate calcium phosphate from the bone structure (Pérez et al. 1989; Serper and Çalt 2002; Sousa and Silva 2005; Sales et al. 2018).
Total demineralization is a process using chelating agents to bind with some types of minerals such as Mg2+ or Ca2+. Likewise, this process gives more advantage to prevent DNA degradation by inhibiting deoxyribonuclease (DNases) (Loreille et al. 2007; Anderung et al. 2008). Ethylene-diaminetetra-acetic acid (EDTA), ethylene-glycol-ether-diaminetetra-acetic acid (EGTA), and 1,2 cyclohexane-diaminetetra-acetic acid (CDTA) have also been recommended for total demineralization (Sousa and Silva 2005). Although many studies have attempted to find appropriate methods for sample preparation prior to DNA extraction, there was still no clear procedure for sample preparation from skeletal remains (Pérez et al. 1989; Serper and Çalt 2002; Sousa and Silva 2005; Sales et al. 2018). A highly efficient DNA extraction method has been developed such as total demineralization plus silica membrane columns method or modified Hi-Flow® silica method, and PrepFiler® BTA kit. Total demineralization of bone powder significantly increased DNA yield and improve STR typing results (Lee et al. 2010; Seo et al. 2010; Hasap et al. 2020). Although the concentration of EDTA in the demineralization process has been reported to affect DNA extraction, other factors including pH of EDTA solution, incubation temperature, incubation time, and volume of EDTA solution have been probably suggested to affect complete total demineralization. Only a few studies have been carried out on all influencing factors on human skeletal remains. Previous studies related to the demineralization process of EDTA have been conducted on human teeth (Pérez et al. 1989; Serper and Çalt 2002; Sousa and Silva 2005) or bovine bones(Sales et al. 2018); however, the application in old human skeletons has been less performed. Thus, this study was conducted to study the factors influencing total demineralization process in DNA extraction from human skeletal remains and developed effective methods with less time-consuming, reasonable cost, and compatible for laboratory to obtain qualified DNA for DNA profiling.
Methods
Skeletal samples
Skeletal samples were collected from unidentified human skeletal remains for personal identification and investigating the cause of death since the 1990s. Four right femurs and one left femur from five male unidentified human skeletal remains were randomly selected. All femurs were cut at the proximal anterior shaft below the lesser trochanter. Then, the skeletal samples were cleaned and ground into bone powder by using the Freezer Mil 6750 with liquid nitrogen. All five powder samples were combined in a pooled sample and distributed in the amount of 100 mg each for further testing.
Total demineralization method based on Plackett-Burman design
Due to the limited experimental samples when considering all factors into account, the program Design Expert version 6.0.8 was applied to find appropriate conditions. The program automatically generated different experimental conditions according to input parameters. Five factors consisting of EDTA concentration, pH of EDTA solution, incubation time, incubation temperature, and volume of EDTA solution were investigated as shown in Table 1. Thirteen experimental runs were generated using the Design-Expert software (version 6.0.8, Stat-Ease Corporation, USA) for five different factors as described above. Each factor was determined at two levels, low (− 1) and high (+ 1) values, based on the literature reported on total demineralization of skeletal remains (Pérez et al. 1989; Hagelberg and Clegg 1991; Żołędziewska et al. 2002; Loreille et al. 2007; Lee et al. 2010; Bilic et al. 2012). Also, the chelated calcium concentration from bone samples acted as a response.
Total demineralization process
One hundred milligrams of bone powder was prepared in a test tube. The demineralization buffer containing varying factors according to Table 1 were added into the test tube based on the Plackett-Burman design matrix. After the demineralization step, the bone solution was centrifuged to separate the supernatants and bone powder residue (Surinyanrattakorn 2011). One milliliter of supernatants was transferred into a new tube to measure the calcium concentrations using atomic absorption spectrophotometry. The remaining bone powder residue was collected for further experiment.
Determination of calcium in demineralization buffer
To determine the calcium level chelated from the bone in demineralization buffer after the completed demineralization process, the calcium concentration was quantified by atomic absorption spectrophotometry (Sales et al. 2018). The amount of calcium in the EDTA solution was calculated comparing the calcium standard curve and expressed as mg/dL.
DNA qualification and quantification
To determine the quality and quantity of the extracted DNA in the bone processed with each demineralization condition, DNA extraction was done by employing a commercial kit (QIAamp®DNA Investigator Kit). After that, two regions of the hemoglobin subunit beta (HBB) gene were amplified by quantitative real-time polymerase chain reaction (qRT-PCR) to assess the quality of the extracted DNA as previously described (Sengüven et al. 2014). The primer sets with different amplicon sizes are shown in Table 2 (Chen et al. 2017). Real-time PCR was performed by Applied Biosystems 7500 Fast Real-Time PCR Systems (Applied Bioscience, USA), with 40 cycles of the following temperature condition: pre-denaturation at 95 °C for 10 min, denaturation at 95 °C for 15 s, annealing at 60 °C for 30 s, and extension at 72 °C for 30 s. The specificity of amplification and quality of amplification was confirmed by analysis of amplifying product on 1.5% agarose gel electrophoresis. In addition, the concentration of the extracted DNA in the bone processed with each demineralization condition was investigated by NanoDrop Spectrophotometer (NanoDrop Technologies, Inc., USA).
STR profiles
To apply the appropriate demineralization method for STR typing, the best demineralization process from the previous experiment was selected. Five individual samples were tested. Each sample was divided into two groups: without and with demineralization process before DNA extraction. The DNA extraction was employed using a commercial kit (QIAamp®DNA Investigator Kit) (QIAGEN, Germany). Sixteen autosomal STR typing were investigated by QIAGEN Investigator® IDplex Plus Kit with Genetic Analyzer 3500 (QIAGEN, Germany). Completed files were analyzed using the Gene Mapper IDX (QIAGEN, Germany) to compare the STR pattern from both two groups.
Statistical analysis
The data were analyzed using descriptive statistics. The comparison of different demineralization methods was also analyzed using the Plackett-Burman design program. The amount of calcium was expressed as mean ± standard deviation (SD) and was compared between the groups using ANOVA and Fisher’s least significant difference (LSD).
Results
The physical characteristics of bone samples were investigated. Bones B and D were relatively yellow-liked color on the external surface while bones A, C, and E were relatively white and yellow (Fig. 1). The age at death of the individuals was considered by the fovea capitis of the femoral head, the pubic symphysis of the hip bone, or the median palatine suture. Bone A represented young to middle adulthood. Bone B was probably associated with a person who was less than 50 years old at the time of death. Bone C was probably 27 to 66 years old. Bone D was probably younger than the individual who belonged to bone C. It could also be estimated that the individual of bone E might be more than 50 years old at the time of death.
The calcium levels in EDTA solution after demineralization was quantitated to represent the highest efficiency of demineralization. The results found that the amount of calcium ranged between 5.90 ± 0.40 and 273.35 ± 39.16 mg/dL. The effect of each factor on the calcium level was represented in the regression analysis for the Plackett-Burman design in Table 3. ANOVA for a model that was not adjusted for curvature was significant (p = 0.0007), allowing for the evaluation of the different factors on calcium level in the extraction buffer. Taken together, the lack of fit error represented a significant effect (p = 0.005) which means the high precision of the design. Also, the model had a coefficient of determination (R2) of 0.93 that could explain 93.0% variability of the data. It can be seen clearly that only one factor (EDTA concentration) was found to significantly enhance calcium chelation from the bone (p value < 0.0001) while the remaining four factors (pH of EDTA solution, incubation temperature, incubation time, and volume of EDTA solution) did not significantly promote calcium chelation from the bone. Moreover, the coefficient estimation of EDTA concentration was 108.85. It could be explained that the calcium level after total demineralization was a direct proportion with the increase of EDTA concentration.
As shown in Fig. 2, the highest and lowest levels of calcium were observed in experimental no. 9 and 7, respectively. The calcium level of the experimental groups 1, 2, 4, 8, 9, and 13 was significantly greater than those of experimental groups 3, 5, 6, 7, 10, 11, and 12. It has corresponded with the result of the previously mentioned experiment that the concentration of the EDTA solution directly influenced the calcium content in the extraction buffer. Although experimental no. 10 used a high concentration of EDTA (0.5 M), the calcium level of this group was lower than the level of the experimental groups using the same concentration of EDTA (0.5 M).
The calcium level in EDTA solution after the demineralization process. Thirteen experimental groups were varied with different factors. The calcium level chelated from the bone in demineralization buffer after the completed demineralization process was quantified by atomic absorption spectrophotometry. *p value less than 0.05 that is statistically significant
From the results, the experimental groups obtaining the significantly highest calcium level was chosen. The quality of extracted DNA was compared between seven conditions for bone demineralization (experimental no. 1, 2, 4, 8, 9, and 13) and without bone demineralization (experimental no. 0). The results showed that the extracted DNA from the bone in experimental no. 4 and 9 were amplified by both primers (113 and 251 bp), whereas no amplification products were observed in the bone without demineralization (Table 4). To confirm the specificity of amplification and quality of amplification, the PCR product was run on 1.5% agarose gel in Figs. 3 and 4. From these data, it was assumed that the amplification of both primer lengths (113 and 251 bp) among the extracted DNA from the demineralization process of experimental no. 4 and 9 provided a good quality of extracted DNA. In addition, the DNA concentration was varied between 7.84 ± 0.30 and 11.05 ± 1.74. The concentration of extracted DNA was not significantly different between each group. However, the extracted DNA from method no. 9 tended to obtain the highest concentration of DNA in the case of the bone processed with the demoralization method (Table 4).
Although the demineralization method no. 4 and 9 represented the appropriate method to chelate calcium with good quality of extracted DNA, experimental no. 9 was selected to perform STR typing since this method required a short processing time for incubation (6 h) which can be easily applied in the laboratory. Therefore, each of the five samples of bones A, B, C, D, and E was tested in two groups; without (I) and with (II) demineralization according to the method used in experimental group 9.
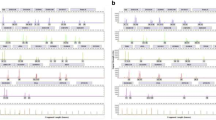
According to the different amplicon sizes, alleles on amelogenin, THO1, TPOX, and D16S539 with amplicon size approximately 70–140 bp can be detected among the extracted DNA of bone sample with/without demineralization process prior STR typing. However, alleles on other loci with amplicon size more than 140 bp could be probably detected among extracted DNA of bone sample with/without demineralization process prior to STR typing. Bones B and D showed a significant increase in percentages of observed alleles of bone with demineralization, from 68.75 to 81.25% and 37.5 to 50%, respectively. Interestingly, some loci can be observed in the bone which was processed with demineralization such as vWA, D7S820, and FGA in bone B as well as vWA, CSF1PO, FGA, and D8S1179 in bone D (Fig. 5). It could be postulated that the demineralization process probably improved STR profiles in some bone specimens (Table 5).
Discussion
DNA extraction from human skeletal remains was a challenging method. Environment, the structure of bone, and process in the laboratory were the influencing factors to the success rate for good quality and quantity DNA. Previous publications have reported the suitable and applicable method for bone preparation before DNA extraction such as demineralization. This process is based on the use of EDTA to chelate some mineral composition in specimens (Balayan et al. 2015). The concentration and pH of EDTA were previously reported to affect the demineralization process of the teeth or on the bovine cortical bone (Sales et al. 2018). Hasap et al. (2020) found that total demineralization combined with DNA purification by commercial kit improved DNA yield and STR profiles (Hasap et al. 2020). However, most tested bone samples would be around the age between less than 1 to 4 years and few samples would be around the ages 5 to 7 years, whereas this research used more than 20 years old preservation of skeletal remains.
Five factors including EDTA concentration, pH of EDTA solution, incubation time, incubation temperature, and volume of EDTA solution were focused on this present study. The result indicated that the concentration of EDTA was the only factor that directly affected the calcium content in the extraction buffer after the DNA extraction process. The amount of calcium removed from bone increased in direct proportion to the concentration of EDTA solution. Our result was concordance with previous reports observing in old sternum human bone (more than 20 years of preservation). DNA profiling was improved when using a high concentration of EDTA (0.5 M) in the demineralization process (Balayan et al. 2015). The comparison of different concentrations of EDTA revealed that the EDTA concentration of 0.5 M was capable to chelate the highest amount of calcium and phosphorus from the bones and teeth (Serper and Çalt 2002; Sales et al. 2018). The rate of calcium chelation by different concentrations of EDTA was rapid within 24 h. However, the increase of EDTA concentration did not result in increased rates of calcium chelation after 48 h of incubation (Kiviranta et al. 1980).
The other factors including the pH of EDTA solution, incubation time, and temperature, as well as the volume of EDTA was found to influence the successful demineralization. Alkaline EDTA can be better captured with calcium ion than acidic EDTA solution (Callis and Sterchi 1998; Sales et al. 2018). Moreover, the over incubation of the bone with EDTA tended to increase DNA degradation resulting in failure STR typing (Jakubowska et al. 2012). So, most studies suggest that the optimal incubation time could be ranged between 24 h and 15 days (Jakubowska et al. 2012; Hasan et al. 2014). However, our study showed that all these factors did not play a significant role in the demineralization process according to the study of Kiviranta et al. (1980) and Seo et al. (2010).
To estimate the quality of the extracted DNA, the investigation of DNA quality especially DNA degradation based on two DNA target sequences of different lengths could be monitored by PCR amplification (Alonso et al. 2004). The highly fragmented DNA samples could be amplified shorter amplicon size of an approximately 62-bp target than a larger amplicon size of about 170–190-bp target (Whitaker et al. 1995; Santos et al. 2009). Moreover, the ratio of PCR product between short and long amplicon size was shown to provide a good estimation of the degree of degradation present in a sample (Swango et al. 2006). We found that extracted DNA with demineralization buffer no. 4 and 9 showed the complete amplification with two amplicon size targets and a high yield of DNA amount.
From this result, we assumed that the completed demineralization process in DNA extraction represented the highest calcium level and quality of extracted DNA might improve STR profiles in skeletal remains as described elsewhere (Bender et al. 2000; Loreille et al. 2007; Bilic et al. 2012; Jakubowska et al. 2012; Balayan et al. 2015; Zupanič Pajnič et al. 2016). Therefore, the demineralization process according to experimental group no. 9 was selected to investigate the STR profiles in five different bone samples. Our result illustrated that the demineralization process probably improved STR profiles in some tested bone specimens (bones B and D).
Besides improving STR typing by total demineralization, it was probable that other factors consisting of bone character and age at death might associate with the quality of DNA. For bones B and D, demineralization improved the quality of some loci showing in STR profiles while it could not increase the quality of STR profiles obtained from bones A, C, and E. To consider the bone character, we observed the color changes on the external surface of the bones, especially changes to the relatively light or white color associated with sun bleaching exposure. As the bones were exposed to sunlight for a long time, the exterior surface appeared more white in color (Pokines and Symes 2013). We assumed that bones A, C, and E were left in the environment for a long time and older in age than bones B and D resulting in DNA degradation by UV and the environment (Latham and Miller 2019). This is consistent with the observation of Żołędziewska et al. (2002). The decalcification process did not require the severely damaged bone due to the smaller loss of the DNA during preparation (Żołędziewska et al. 2002). Regarding age at death, the bone’s age at death was estimated, and our data represented that bones B and D were probably younger than those who belonged to bones A, C, and E. High bone density in younger bones can preserve DNA better than older bones (Latham and Miller 2019).
The limitation of this study is that we cannot retrieve the information of postmortem, which environment of death, postmortem interval, preparing process of bone, and preservation process in the laboratory room. The controlled setting, such as “the body farm” where the environment or postmortem interval of the bone sample can be controlled, was investigated. Additionally, the successful extraction of human skeletal remains depended on many factors such as the environment, the strong bone structure, and the selection of bone fragments used for DNA extraction, some of which cannot be controlled.
Conclusions
Our study demonstrated that EDTA concentration appeared to have an influence on the demineralization process to chelate calcium prior to the extraction of DNA from skeleton remains. The optimal condition for the total demineralization process was 0.5 M EDTA, pH 10, 1 ml at 60 °C in 6 h for obtaining the good quality and high yield of extracted DNA. The total demineralization process in bone preparation before the extraction with a commercial DNA extraction kit probably improves the STR profiles in some bone specimens. However, other factors such as the environment and external structure of the bone affected the quality of STR profiles and should be further studied.
Availability of data and materials
Not applicable
References
Alonso A, Martín P, Albarrán C et al (2004) Real-time PCR designs to estimate nuclear and mitochondrial DNA copy number in forensic and ancient DNA studies. Forensic Sci Int 139:141–149 https://doi.org/10.1016/j.forsciint.2003.10.008
Anderung C, Persson P, Bouwman A et al (2008) Fishing for ancient DNA. Forensic Sci Int Genet 2:104–107 https://doi.org/10.1016/j.fsigen.2007.09.004
Balayan A, Kapoor A, Chaudhary G, Raina A (2015) Evaluation of techniques for human bone decalcification and amplification using sixteen STR markers. Egypt J Forensic Sci 5:30–35 https://doi.org/10.1016/j.ejfs.2014.05.002
Bender K, Schneider PM, Rittner C (2000) Application of mtDNA sequence analysis in forensic casework for the identification of human remains. Forensic Sci Int 113:103–107 https://doi.org/10.1016/S0379-0738(00)00223-1
Bilic A, Loreille O, Parsons TJ, Amory S (2012) Forensic Science International: Genetics automatable full demineralization DNA extraction procedure from degraded skeletal remains. 6:398–406 https://doi.org/10.1016/j.fsigen.2011.08.004
Callis G, Sterchi D (1998) Decalcification of bone: literature review and practical study of various decalcifying agents, methods, and their effects on bone histology. J Histotechnol 21:49–58 https://doi.org/10.1179/his.1998.21.1.49
Chen TB, Xun Z, Lin JP et al (2017) Real-time PCR for quantitative detection of mitochondrial DNA from peripheral blood mononuclear cell in patients with HBV-related hepatocellular carcinoma. Int J Clin Exp Med 10:5629–5636
Hagelberg E, Clegg JB (1991) Isolation and characterization of DNA from archaeological bone. Proc R Soc B Biol Sci 244:45–50 https://doi.org/10.1098/rspb.1991.0049
Hasan M, Hossain T, Majumder AK et al (2014) National Forensic DNA Profiling Laboratory, Dhaka Medical College Campus, Dhaka - 1000, Bangladesh Key words. Dhaka Univ J Biol 23:101–107
Hasap L, Chotigeat W, Pradutkanchana J et al (2020) A novel, 4-h DNA extraction method for STR typing of casework bone samples. Int J Legal Med. https://doi.org/10.1007/s00414-019-02232-9
Jakubowska J, Maciejewska A, Pawłowski R (2012) Comparison of three methods of DNA extraction from human bones with different degrees of degradation. Int J Legal Med 126:173–178 https://doi.org/10.1007/s00414-011-0590-5
Kiviranta I, Tammi M, Lappalainen R et al (1980) The rate of calcium extraction during EDTA decalcification from thin bone slices as assessed with atomic absorption spectrophotometry. Histochemistry 68:119–127 https://doi.org/10.1007/BF00489507
Latham KE, Miller JJ (2019) DNA recovery and analysis from skeletal material in modern forensic contexts. Forensic Sci Res 4:51–59 https://doi.org/10.1080/20961790.2018.1515594
Lee HY, Park MJ, Kim NY et al (2010) Simple and highly effective DNA extraction methods from old skeletal remains using silica columns. Forensic Sci Int Genet 4:275–280 https://doi.org/10.1016/j.fsigen.2009.10.014
Loreille OM, Diegoli TM, Irwin JA et al (2007) High efficiency DNA extraction from bone by total demineralization. Forensic Sci Int Genet 1:191–195 https://doi.org/10.1016/j.fsigen.2007.02.006
Milos A, Selmanović A, Smajlović L et al (2007) Success rates of nuclear short tandem repeat typing from different skeletal elements. Croat Med J 48:486–493
Mundorff AZ, Bartelink EJ, Mar-Cash E (2009) DNA preservation in skeletal elements from the world trade center disaster: recommendations for mass fatality management. J Forensic Sci 54:739–745 https://doi.org/10.1111/j.1556-4029.2009.01045.x
Mundorff AZ, Davoren J, Weitz S (2013) Developing an empirically based ranking order for bone sampling: examining the differential DNA yield rates between human skeletal elements over increasing post mortem intervals. Doc number 241868 1–76
Pérez VC, Cárdenas MEM, Planells US (1989) The possible role of pH changes during EDTA demineralization of teeth. Oral Surgery, Oral Med Oral Pathol 68:220–222 https://doi.org/10.1016/0030-4220(89)90196-5
Pokines J, Symes SA (2013) Manual of forensic taphonomy, 1st edn. CRC Press 496 Pages - 164 B/W Illustrations
Renneberg R, Berkling V, Loroch V (2017) Analytical biotechnology and the human genome. Biotechnol Beginners:343–391 https://doi.org/10.1016/b978-0-12-801224-6.00010-2
Sales E, Silva CERD, Letichevsky S et al (2018) Chemical induced demineralization study in cortical bone. J Instrum:13 https://doi.org/10.1088/1748-0221/13/05/C05010
Samsuwan J, Somboonchokepisal T, Akaraputtiporn T, et al (2018) A method for extracting DNA from hard tissues for use in forensic identification. Biomed Reports 433–438. https://doi.org/10.3892/br.2018.1148
Santos S, Sá D, Bastos E et al (2009) An efficient protocol for genomic DNA extraction from formalin-fixed paraffin-embedded tissues. Res Vet Sci 86:421–426 https://doi.org/10.1016/j.rvsc.2008.08.007
Sengüven B, Baris E, Oygur T, Berktas M (2014) Comparison of methods for the extraction of DNA from formalin-fixed, paraffin-embedded archival tissues. Int J Med Sci 11:494–499 https://doi.org/10.7150/ijms.8842
Seo SB, Zhang A, Kim HY et al (2010) Technical note: efficiency of total demineralization and ion-exchange column for DNA extraction from bone. Am J Phys Anthropol 141:158–162 https://doi.org/10.1002/ajpa.21193
Serper A, Çalt S (2002) The demineralizing effects of EDTA at different concentrations and pH. J Endod 28:501–502 https://doi.org/10.1097/00004770-200207000-00002
Sousa SMG, Silva TL (2005) Demineralization effect of EDTA, EGTA, CDTA and citric acid on root dentin: a comparative study. Braz Oral Res 19:188–192 https://doi.org/10.1590/s1806-83242005000300006
Surinyanrattakorn D (2011) Comparison of DNA extraction methods from unidentified human skeletal remains. Silapakorn University, Thailand
Swango KL, Timken MD, Chong MD, Buoncristiani MR (2006) A quantitative PCR assay for the assessment of DNA degradation in forensic samples. Forensic Sci Int 158:14–26 https://doi.org/10.1016/j.forsciint.2005.04.034
Whitaker JP, Clayton TM, Urquhart AJ, et al (1995) Short tandem repeat typing of bodies from a mass disaster: high success rate and characteristic amplification patterns in highly degraded samples. Biotechniques 18:670-674+676
Żołędziewska M, Gronkiewicz S, Dobosz T (2002) Comparison of various decalcificators in preparation of DNA from human rib bones. Przegląd Antropol Rev 65:75–80
Zupanič Pajnič I, Debska M, Gornjak Pogorelc B et al (2016) Highly efficient automated extraction of DNA from old and contemporary skeletal remains. J Forensic Leg Med 37:78–86 https://doi.org/10.1016/j.jflm.2015.11.001
Acknowledgements
This work was supported by the Faculty of Medicine Research Fund, Chiang Mai University, Chiang Mai, Thailand (Grant number: 023/2563). And we would like to thank the Department of Forensic Medicine, Chiang Mai University, Thailand, for allowing access to their skeletal collection for this research.
Funding
Faculty of Medicine Research Fund, Chiang Mai University, Chiang Mai, Thailand (Grant number: 023/2563)
Author information
Authors and Affiliations
Contributions
Conceptualization: P.B., C.K., and S.K. Methodology: P.B. and S.K. Formal analysis: P.B. and S.K. Data curation: P.B. Writing—original draft preparation: P.B. Writing—review and editing: C.K., S.R., and S.K. Supervision: S.K. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was carried out in accordance with the Code of Ethics of the World Medical Association (Declaration of Helsinki) and approved by the Ethics Committee of the Faculty of Medicine, Chiang Mai University, Thailand (No.EXEMPTION-6521/2019)
Consent for publication
Not applicable
Competing interests
The authors declare that they have no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Booncharoen, P., Khacha-ananda, S., Kanchai, C. et al. Factors influencing DNA extraction from human skeletal remains: bone characteristic and total demineralization process. Egypt J Forensic Sci 11, 2 (2021). https://doi.org/10.1186/s41935-021-00216-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s41935-021-00216-8