Abstract
Background
Escherichia coli O157:H7 is one of the major foodborne bacterial pathogens and also a biodefense agent. To ensure food safety and public health, it is very important to develop rapid methods for E. coli O157:H7 detection. In this study, we designed a nanoparticle-labeled biosensor for the rapid detection of E. coli O157:H7 in broth.
Results
Magnetic nanoparticles (MNPs) were conjugated with monoclonal antibodies (Abs) to separate target E. coli O157:H7 cells from broth samples. Gold nanoparticles (AuNPs) were conjugated with polyclonal Abs, and were then introduced to the MNP-target complex to form a sandwich MNP-target-AuNP. By measuring the amount of AuNPs through an electrochemical method, the presence and the amount of the target bacteria were determined. Results showed a sensitivity of 101 colony forming units per milliliter (cfu/ml) with a linear range of 101–106 cfu/ml.
Conclusions
Compared to conventional culture plating methods, the biosensor reduced the detection time from 2 to 4 days to less than 1 hour with a simple target extraction method. The AuNP-labeled biosensor has potential applications in the rapid detection of infectious agents for public health, biodefense, and food/water safety.
Similar content being viewed by others
Background
Rapid detection of pathogenic bacteria is critical to public health, biodefense, and food/water safety. Escherichia coli O157:H7 is one of the major foodborne bacterial pathogens and also a biodefense agent. There were several outbreaks of E. coli O157:H7 in recent years that endangered public health [1–3]. Because conventional culture plating methods for E. coli O157:H7 take two to four days to obtain results, development of rapid detection methods for this organism is important. Biosensors are emerging technologies that have the potential for getting rapid results and that can be employed in the field. There are many biosensor configurations and approaches that are in the research and design stage. These configurations include antibody-based systems [4–8], enzyme-based detection [9, 10] and DNA-based sensors [11, 12]. In addition to speed, biosensors have the potential to generate highly sensitive results. This is especially critical as many bacterial infections could be caused by as low as 10 organisms [13].
The application of nanomaterials in biosensors, such as nanoparticles with optical, electronic and magnetic properties, has drawn interest. Because of their unique characteristics, nanoparticles have been used to enhance sensor sensitivity either by increasing the capture efficiency of the target molecules or by utilizing the optical and electronic properties of the nanostructures to amplify signals. Magnetic nanoparticles were employed for separating targets for bacterial detection [6, 14, 15]. Gold nanoparticles (AuNPs) were used for signal amplification [15, 16]. Polymeric nanoparticles were also introduced for signal amplification [6, 17].
In this paper, we developed an electrochemical biosensor using antibody-modified nanoparticles for the detection of E. coli O157:H7. Two novel nanoparticles were utilized in the biosensor design: 1) polymer-coated magnetic nanoparticles (MNPs) to separate the target bacteria from the sample matrix and carbohydrate-capped AuNPs to label the separated target by forming a sandwich structure and generate the signal. The signal of AuNPs for the corresponding target was measured by differential pulse voltammetry (DPV) on a screen printed carbon electrode (SPCE) chip. The biosensor enabled rapid pathogen detection in 45 min from sample preparation to final readout of results.
Results and discussion
Magnetic separation of target E. coli O157:H7
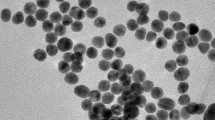
E. coli O157:H7 cells were magnetically captured as shown in Fig. 1. We modified the Fe2O3 nanoparticles with polyaniline (PANI) for direct immobilization of anti-E. coli O157:H7 antibody (Ab). Figure 2 presents the transmission electron microscopy (TEM) images of the Fe2O3 nanoparticle core (Fig. 2a) and the PANI-coated MNPs (Fig. 2b) [18]. Figure 2a reveals that the average diameter of the Fe2O3 nanoparticle core is 20 nm, while Fig. 2b shows that the PANI-coated MNPs have diameters ranging from 50 to 100 nm. The increase of the diameter was due to the formation of PANI around the Fe2O3 core. According to the insets, the electron diffraction pattern in Fig. 2a exhibited a typical maghemite (γ-Fe2O3) nanoparticle structure [19]. In Fig. 2b, the electron diffraction pattern shows a set of rings which are typical for PANI [20], noted that it has less bright spots than in Fig. 2a. This pattern also indicates the coating of Fe2O3 core by PANI.
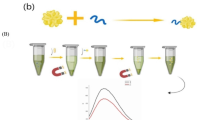
Schematic of the gold nanoparticle (AuNP)-labeled biosensor. Target cells in a sample were captured by magnetic nanoparticle (MNP)-antibody (Ab) conjugates and separated by a magnet. Then the cells were labeled with AuNPs. The MNP-Ab-cell-Ab-AuNP complexes were transferred onto a screen printed carbon electrode that is a chip connected to a potentiostat for electrochemical measurement
Polyaniline (PANI)-coated magnetic nanoparticles (MNPs). Transmission electron microscopy (TEM) images of: (a) Fe2O3 core; (b) PANI-coated MNPs. The insets show the electron diffraction patterns of the nanoparticles [18]. Used with permission from Biosensors & Bioelectronics
Electrostatic interaction has been used to modify the PANI-coated MNPs with antibody. The interaction between the negatively charged Fc fragment of antibody molecules and the positively charged PANI contributes to the conjugation [18]. Figure 3a shows a schematic of the interaction between the PANI-coated MNPs and the antibody.
a Schematic of the conjugation of Polyaniline (PANI)-coated magnetic nanoparticles (MNPs) and antibody; and (b) Scanning electron microscopy (SEM) image of an antibody-conjugated MNP bound to an E.coli O157:H7 cell [6]; used with permission from International Journal of Food Safety, Nutrition and Public Health (Inderscience retains copyright of the original article and figure)
The MNP-Ab was used to magnetically separate the target from the sample through the antibody-antigen interaction. The scanning electron microscopy (SEM, Fig. 3b) confirms the capture of the target [6]. In this image, a rod-shaped E. coli O157:H7 cell is bound to MNP-Ab conjugate.
AuNP-labeled biosensor for detection
Figure 4 shows two typical sensorgrams of native AuNPs and native MNPs. Current peak for AuNPs is at 0.3 V and MNPs is at 0.58 V. Figure 5 shows typical DPV sensorgrams for the detection of E. coli O157:H7 in different cell concentrations (102, 104, and 106 colony forming units per milliliter, cfu/ml). The sensorgrams show a wide curve that seems to include both AuNPs and MNPs. For the analysis, peak current to the left (representing AuNPs) around 0.3 V was chosen for signal reporting. As shown in the graph, peak current for AuNPs increased with increasing cell concentration. Figure 5 confirms the formation of the MNP-cell-AuNP complex. The amount of target cells detected was proportional to the amount of AuNPs.
A linear relationship between signal to noise ratio (SNR) and cell concentration is shown in Fig. 6, with an R2 value of 0.953. The SNR was calculated as the peak current of AuNPs for each concentration divided by the current at 0.3 V of the blank, which was used as the signal of this biosensor. The SNR for 101, 102, 103, 104, 105 and 106 cfu/ml are 1.28, 1.56, 1.69, 1.73, 2.12 and 2.36, respectively. A statistical analysis using t test was conducted and the results are presented in Table 1. As shown, cell concentration at 101 cfu/ml has a P value of 0.0546 (which is close to critical value of P = 0.05). The P value for other concentrations shows that the samples are significantly different from the blank. Therefore, the lowest cell concentration is weakly 101 cfu/ml and strongly at 102 cfu/ml. These results verify that MNP-Ab-cell-Ab-AuNP is an effective approach to highly sensitive detection. The specificity of the biosensor, which largely depends on the monoclonal antibody used in magnetic separation, was evaluated in another study by our lab [21]. An inclusivity of 94 % and an exclusivity of 69 % was obtained [21]. The inclusivity was calculated as the number of positive tests identified by the antibody divided by the actual total positive test number. The exclusivity was calculated as the number of negative tests identified divided by the actual total negative test number. The specificity of the monoclonal and polyclonal Abs-based sandwich configuration was also verified in our previous study [8].
Conclusions
A AuNP-labeled biosensor has been developed for the sensitive and rapid detection of E. coli O157: H7. The lower limit of detection is at 101 cfu/ml with a dynamic range of 101 to 106 cfu/ml. Target extraction and detection are completed within 45 min. Furthermore, sample preparation requires a simple magnet without special equipment. Due to its simplicity and rapid results, this biosensor has the potential for in-field use in quickly screening bacterial contamination in food products (e.g. ground beef, vegetables, and juices) and water systems. The biosensor is also a promising technology for point-of-care screening of diseases and environmental determination of harmful biological agents.
Materials and methods
Reagents and materials
Two kinds of nanoparticles were synthesized: MNPs and AuNPs. Aniline, iron (III) oxide nanopowder, ammonium persulfate, methanol, and diethyl ether (Sigma-Aldrich, St. Louis, MO) were used for the synthesis of the MNPs. Gold (III) chloride trihydrate (Aldrich, MO) and dextrin (Fluka, MO) were used for the synthesis of AuNPs under alkaline conditions [22].
MNPs were functionalized with monoclonal anti-E. coli O157:H7 antibody obtained from Meridian Life Science, Inc (Saco, ME). AuNPs were conjugated with polyclonal anti- E. coli O157:H7 antibody from Meridian Life Science, Inc (Saco, ME). Protein A from Staphylococcus aureus (Sigma-Aldrich, St. Louis, MO) was used as the linkage agent for AuNP and antibody conjugation.
Triton-X100, phosphate buffered saline (PBS), casein, bovine serum albumin (BSA) and sodium phosphate (dibasic and monobasic) were obtained from Sigma-Aldrich (St. Louis, MO). PBS buffer (0.01 M and 0.1 M, pH 7.4), 0.01 M PBS buffer with 0.05 % (w/v) Triton-X100, phosphate buffer (0.1 M sodium phosphate, pH 7.4), 0.01 M PBS buffer with 0.01 % casein, 0.01 M PBS buffer with 0.1 % (w/v) BSA were prepared with deionized water from Millipore Direct-Q system. PBS buffer and phosphate buffer were used in preparing nanoparticle-Ab conjugates and in washing. PBS buffer with casein or BSA was used to block nanoparticle surface against nonspecific binding. PBS buffer with Triton-X100 was used for washing off unbound or nonspecifically bound reactants after capture.
Bacterial culture
E. coli O157:H7 Sakai strain was obtained from the Nano-Biosensors Lab collection at Michigan State University. The colonies from frozen (stored at –70 °C) culture were grown on trypticase soy agar (BD Biosciences, MD) plates. A single colony was isolated and inoculated in tryptic soy broth (BD Biosciences, MD) and grown overnight at 37 °C. One milliliter of the liquid culture was transferred to another tube of tryptic soy broth and incubated overnight at 37 °C. One milliliter of this liquid culture was transferred to a new tube of broth and incubated at 37 °C for 6 h before each experiment. The serial dilutions of bacterial culture were prepared using 0.1 % (w/v) peptone water (Fluka-Biochemika, Switzerland) before each experiment. Viable cells were enumerated by microbial plating on Sorbitol MacConkey agar (SMAC, BD Biosciences, MD).
Apparatus
Electrochemical measurement was performed with a potentiostat/galvanostat (263A, Princeton Applied Research, MA) with a software operating system (PowerSuite, Princeton Applied Research, MA) on a computer connected to the potentiostat. The measurement was performed by introducing each sample onto a SPCE chip (Gwent Inc. England). The SPCE chip consisted of a working carbon electrode and a counter and reference silver/silver chloride electrode. One hundred microliters of each sample were introduced to the electrode area on the SPCE chip.
Synthesis of nanoparticles
PANI coated MNPs were synthesized according to our method [6]. Fifty milliliters of 1 M hydrochloric acid, 10 ml of water and 0.4 ml of aniline monomer were mixed in a flask, and then 0.65 g of iron (III) oxide nanopowder were added to the solution to maintain a final γ-Fe2O3: aniline weight ratio of 1: 0.6. The mixture was put in a beaker filled with ice and sonicated for 1 h. The solution was stirred while it was still on ice. During the stirring, ammonium persulfate (1 g of ammonium persulfate in 20 ml deionized water) was added to the solution slowly for 30 min. The solution was stirred for another 1.5 h. After the reaction, the solution was filtered using 2.5 μm filter paper and washed with 20 % methanol. Hydrochloric acid (1 M) was used to wash until the filtrate became clear, followed by washing with 10 ml of 20 % methanol. The filtrate was filtered again using a 1.2 μm filter paper, and 10 ml of 20 % methanol solution was added to the filter. The hydrochloric acid and methanol wash was repeated. The nanoparticles on the filter paper were left under a fume hood to dry for 24 h at room temperature and stored in a vacuum desiccator after drying.
AuNPs were synthesized under alkaline conditions following the approach published by Anderson et al [22]. Briefly, 20 ml of dextrin stock solution (25 g/l) and 20 ml of sterile water were mixed in a 50 ml sterile orange cap tube (disposable). Five milliliters of HAuCl4 stock solution (8 g/ml) were then added, and the pH of the solution was adjusted to 9 with sterile 10 % (w/v) Na2CO3 solution. The final volume was brought to 50 ml with pH 9 water. The reaction was carried out by incubating the solution in a sterile flask in the dark at 50 °C with continuous shaking (100 rpm) for 6 h. A red solution was obtained at the end of the reaction.
Functionalization of nanoparticles
MNPs were functionalized with a monoclonal anti-E. coli O157:H7 antibody [6]. MNPs (2.5 mg) were suspended in 150 μl of 0.1 M phosphate buffer, and sonicated for 15 min. Monoclonal anti-E. coli O157:H7 antibody (2.5 mg/ml, 100 μl) was added to the suspension, and hybridized on tube rotator for 5 min. Twenty five microliters of PBS buffer (0.1 M) were added. Then the conjugation was carried on for 55 min on the tube rotator. The MNPs were separated from the solution by magnetic separation, and blocked by adding 250 μl of 0.1 M tris buffer with 0.01 % casein and incubated for 5 min. This step was repeated three times, and the suspension was put on tube rotator for 1 h to hybridize. Finally, the MNPs were magnetically separated and resuspended in 2.5 ml of 0.1 M phosphate buffer. The MNP-Ab conjugate was stored at 4 °C before use.
AuNPs were conjugated with a polyclonal anti-E. coli O157:H7 antibody through protein A linkage. Two hundred microliters of 1:2 diluted suspension of AuNPs in water were put into a 2 ml microcentrifuge tube and sonicated for 10 min. Then the suspension was centrifuged for 6 min at 13,000 rpm. The supernatant was removed after centrifugation. To modify the surface of the AuNPs, protein A (0.25 mg/ml) in 200 μl of 0.01 M PBS buffer was used to resuspend the AuNPs. The conjugation was conducted by rotating the mixture for 1 h. The modified AuNPs were separated from the suspension by centrifugation for 6 min at 13,000 rpm. The nanoparticles were washed by adding 200 μl of 0.01 M PBS buffer and centrifuged. After removing the supernatant, 100 μl of 1 mg/ml antibody and 100 μl of 0.01 M PBS buffer were added to the tube and mixed for 60 min by rotating. After separating the AuNP-antibody (AuNP-Ab) conjugates from the supernatant, 200 μl of PBS buffer with 0.1 % (w/v) BSA were added to the tube. The mixture was rotated for 30 min. Finally, the AuNP-Ab conjugates were separated from the suspension by centrifugation, and the final suspension of the conjugates in 200 μl PBS buffer with BSA was stored at 4 °C.
Detection of target pathogenic bacteria
Detection of the target pathogen is presented in Fig. 1. Blank control for the tests was peptone water in the same volume as the sample. Firstly, 400 μl of 0.01 M PBS buffer, 50 μl of cell dilution (or peptone water for the blank) and 50 μl of MNP-Ab conjugates were combined in a 2 ml sterile tube. After 15 min hybridization, PBS buffer (55 μl, 0.01 M) with 0.1 % BSA was added to the mixture as a blocking agent. Then, the MNP-E. coli complexes were magnetically separated from the solution and resuspended in 450 μl of 0.01 M PBS buffer. Secondly, 50 μl of the AuNP-Ab conjugates were introduced to the system, followed by 15 min hybridization. After washing the complexes once with 0.01 M PBS buffer, the complexes were resuspended in 500 μl of PBS buffer with 0.05 % Triton-X100, and let stand for 3 min. Finally, the complexes were magnetically separated from the buffer and resuspended in 500 μl of 0.01 M PBS buffer. One hundred microliters of the suspension were plated on SMAC for cell counting. The rest of the complexes were magnetically separated from the supernatant (400 μl).
Electrochemical measurement
The target bacteria were detected by measuring the electrochemical signal of AuNPs. Each sample from the last section (complexes magnetically separated from supernatant) in 100 μl 1 M hydrochloric acid was introduced to the SPCE chip. An oxidation potential of 1.4 V vs. Ag/AgCl was applied to the working electrode. After oxidation, a differential pulse voltammetric (DPV) measurement was performed. The scan was from 1.5 V to −1.5 V. The potential and currents were recorded. All measurements were performed at room temperature. Each sample was measured three times. At least three samples of each concentration of bacteria were tested.
Abbreviations
- AuNPs:
-
Gold nanoparticles
- MNPs:
-
Magnetic nanoparticles
- DPV:
-
Differential pulse voltammetry
- SPCE:
-
Screen printed carbon electrode
- TEM:
-
Transmission electron microscopy
- PANI:
-
Polyaniline
- Ab:
-
Antibody
- SEM:
-
Scanning electron microscopy
- SNR:
-
Signal to noise ratio
- PBS:
-
Phosphate buffered saline
- BSA:
-
Bovine serum albumin
- SMAC:
-
Sorbitol MacConkey agar
References
United States Centers for Disease Control and Prevention (CDC). Multistate Outbreak of E. coli O157:H7 Infections Linked to Eating Raw Refrigerated, Prepackaged Cookie Dough. 2009. http://www.cdc.gov/ecoli/2009/0630.html. Accessed 9 Jun 2015.
CDC. Investigation Announcement: Multistate Outbreak of E. coli O157:H7 Infections Linked to Romaine Lettuce. 2011. http://www.cdc.gov/ecoli/2011/ecoliO157/romainelettuce/120711/. Accessed 9 Jun 2015.
CDC. Multistate Outbreak of Shiga toxin-producing Escherichia coli O157:H7 Infections Linked to Ground Beef. 2014. http://www.cdc.gov/ecoli/2014/O157H7-05-14/index.html. Accessed 9 Jun 2015.
Luo Y, Nartker S, Miller H, Hochhalter D, Wiederoder M, Wiederoder S, et al. Surface functionalization of electrospun nanofibers for detecting E. coli O157:H7 and BVDV cells in a direct-charge transfer biosensor. Biosens Bioelectron. 2010;26(4):1612–7.
Radke SM, Alocilja EC. A high density microelectrode array biosensor for detection of E. coli O157:H7. Biosens Bioelectron. 2005;20(8):1662–7.
Setterington EB, Cloutier BC, Ochoa JM, Cloutier AK, Jain P, Alocilja EC. Rapid, sensitive, and specific immunomagnetic separation of foodborne pathogens. Int J Food Saf Nutr Public Health. 2011;4(1):83–100.
Wang Y, Wang R, Li Y, Srinivasan B, Tung S, Wang H, et al. Detection of Escherichia coli O157:H7 using interdigitated array microelectrode-based immunosensor. Biol Eng. 2010;2(2):49–62.
Wang Y, Fewins P, Alocilja EC. Electrochemical Immunosensor Using Nanoparticle-based Signal Enhancement for Escherichia coli O157:H7 Detection. Sensors Journal, IEEE. 2015; 15(8):4692-9.
Linman MJ, Sugerman K, Cheng Q. Detection of low levels of Escherichia coli in fresh spinach by surface plasmon resonance spectroscopy with a TMB-based enzymatic signal enhancement method. Sensors Actuators B Chem. 2010;145(2):613–9.
Park S, Kim H, Paek S, Hong JW, Kim Y. Enzyme-linked immuno-strip biosensor to detect Escherichia coli O157:H7. Ultramicroscopy. 2008;108(10):1348–51.
Sun H, Choy TS, Zhu DR, Yam WC, Fung YS. Nano-silver-modified PQC/DNA biosensor for detecting E. coli in environmental water. Biosens Bioelectron. 2009;24(5):1405–10.
Wang L, Liu Q, Hu Z, Zhang Y, Wu C, Yang M, et al. A novel electrochemical biosensor based on dynamic polymerase-extending hybridization for E. coli O157:H7 DNA detection. Talanta. 2009;78(3):647–52.
Council for Agricultural Science and Technology (CAST). Foodborne Pathogens: Risks and Consequences. 1994. Report No. 122.
Li K, Lai Y, Zhang W, Jin L. Fe2O3@Au core/shell nanoparticle-based electrochemical DNA biosensor for Escherichia coli detection. Talanta. 2011;84(3):607–13.
Zhang D, Huarng MC, Alocilja EC. A multiplex nanoparticle-based bio-barcoded DNA sensor for the simultaneous detection of multiple pathogens. Biosens Bioelectron. 2010;26(4):1736–42.
Hao R, Song H, Zuo G, Yang R, Wei H, Wang D, et al. DNA probe functionalized QCM biosensor based on gold nanoparticle amplification for Bacillus anthracis detection. Biosens Bioelectron. 2011;26(8):3398–404.
Jiang X, Wang R, Wang Y, Su X, Ying Y, Wang J, et al. Evaluation of different micro/nanobeads used as amplifiers in QCM immunosensor for more sensitive detection of E. coli O157:H7. Biosens Bioelectron. 2011;29(1):23–8.
Pal S, Alocilja EC. Electrically active polyaniline coated magnetic (EAPM) nanoparticle as novel transducer in biosensor for detection of Bacillus anthracis spores in food samples. Biosens Bioelectron. 2009;24(5):1437–44.
Hyeon T, Lee S, Park J, Chung Y, Bin NH. Synthesis of highly crystalline and monodisperse maghemite nanocrystallites without a size-selection process. J Am Chem Soc. 2001;123(51):12798–801.
Shaikh SF, Lim JY, Mane RS, Han S, Ambade SB, Joo O. Wet-chemical polyaniline nanorice mass-production for electrochemical supercapacitors. Synth Met. 2012;162(13–14):1303–7.
Cloutier BC. Development of mitigation strategies toward preventative postures in food defense. Ph.D. United States, Michigan: Michigan State University; 2012.
Anderson MJ, Torres-Chavolla E, Castro BA, Alocilja EC. One step alkaline synthesis of biocompatible gold nanoparticles using dextrin as capping agent. J Nanopart Res. 2011;13(7):2843–51.
Acknowledgements
This project was funded by the Michigan State University Strategic Partnership Grant through A-CAPPP (Anti-Counterfeiting and Product Protection Program) and by the MSU Targeted Support Grants for Technology Development Project. The authors would like to thank Patrick Fewins for his assistance in the experiments.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
YW contributed to experimental design, carried out the experiments and drafted the manuscript. ECA conceived and designed the experiments, participated in discussions, and drafted the manuscript. Both authors read and approved the final manuscript.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Wang, Y., Alocilja, E.C. Gold nanoparticle-labeled biosensor for rapid and sensitive detection of bacterial pathogens. J Biol Eng 9, 16 (2015). https://doi.org/10.1186/s13036-015-0014-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13036-015-0014-z