Abstract
Background
Violacein is a deep violet compound that is produced by a number of bacterial species. It is synthesized from tryptophan by a pathway that involves the sequential action of 5 different enzymes (encoded by genes vioA to vioE). Violacein has antibacterial, antiparasitic, and antiviral activities, and also has the potential of inducing apoptosis in certain cancer cells.
Results
Here, we describe the construction of a series of plasmids harboring the complete or partial violacein biosynthesis operon and their use to enable production of violacein and deoxyviolacein in E.coli. We performed in vitro assays to determine the biological activity of these compounds against Plasmodium, Trypanosoma, and mammalian cells. We found that, while deoxyviolacein has a lower activity against parasites than violacein, its toxicity to mammalian cells is insignificant compared to that of violacein.
Conclusions
We constructed E. coli strains capable of producing biologically active violacein and related compounds, and propose that deoxyviolacein might be a useful starting compound for the development of antiparasite drugs.
Similar content being viewed by others
Background
Violacein is a violet indolocarbazole pigment that is produced by bacteria such as Chromobacterium violaceum, which are commonly found in water and soil throughout the world [1,2,3,4,5,6]. Violacein has antipyretic [7, 8], ulcer-protective [8], antibacterial [9,10,11], antifungal [3, 12], trypanocidal [13, 14], antileishmanial [15], antinematode [16], and antiviral [17] activities. It also has the potential of inducing apoptosis in certain cancer cells [2, 18]. Violacein kills wild-type and drug-resistant strains of the malaria parasite, Plasmodium falciparum and is therapeutically against malaria in mice [19]. These characteristics suggest that violacein has considerable research potential and may represent a chemical scaffold for the developemt of clinically useful drugs.
Commercially, violacein is usually isolated from Chromobacterium [20,21,22] or Janthinobacterium [23, 24]; however, this process is costly and there are reports of rare but deadly infections caused by these bacteria [25,26,27,28]. Hence, there has been considerable interest in the development of safe, and efficient routes to the biosynthesis of this compound [1, 29,30,31,32,33,34].
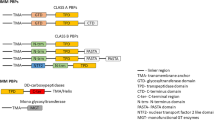
The violacein biosynthetic pathway from L-tryptophan (Fig. 1) requires the expression of five genes: vioA, vioB, vioC, vioD, and vioE [35,36,37,38,39]. It should be noted that VioC enzyme is involved in both the production of deoxyviolacien from protodeoxyviolaceinic acid and in the generation of violacein from protoviolaceinic acid. Several studies have shown that transforming and expressing a complete metabolic pathway into a different bacterial host may lead to improved production of violacein [40,41,42,43]. For example, Rodrigues and co-workers [32,33,34] have successfully engineered Escherichia coli to produce high yields of violacein and the side-product deoxyviolacein. This was accomplished by cloning the complete vioABCDE and the partial vioABCE operons (respectively) from C. violaceum into pBADMycHisB, which allows the induction of the operon by L-arabinose [32, 34]. These authors also metabolically engineered the host’s tryptophan production to maximize the yield of violacein and deoxyviolacein [32, 34, 42]. More recently Jones and co-workers [44] and Xu and co-workers [45] utilized violacein biosynthesis as a model for metabolic pathway balancing and optimization. Employing different approaches, they fine-tuned the expression of violacein-producing enzymes, leading to an improvement in the production of the compound by up to 30-fold when compared to previously reported work.
We have generated a synthetic operon containing the coding sequences of each of the five genes required for violacein biosynthesis, with a codon-usage optimized for E. coli (http://parts.igem.org/Part:BBa_K274002). We also constructed strains lacking vioD, to promote the accumulation of deoxyviolacein. A similar approach was employed by Rodrigues and co-workers for the production of high yields of violacein and deoxyviolacein [32,33,34]. We produced and purified violacein and deoxyviolacein, and characterized their toxicity and antiplasmodial activity in wild-type and drug-resistant Plasmodium falciparum strains; we also determined their activity against Trypanosoma cruzi.
Methods
Bacterial strains and plasmids
We constructed plasmids expressing violacein and deoxyviolacein by sub-cloning the synthetic violacein operon (Part: BBa_K274002) designed by Shuna Gould for the iGEM09_Cambridge project (http://parts.igem.org/Part:BBa_K274002). The synthetic violacein operon is comprised of the 5 coding sequences specifying violacein pathway enzymes (vioA, vioB, vioC, vioD and vioE), each preceded by a ribosome-binding site. The operon was designed with a BamHI site in the space between vioB and vioC open reading-frames (ORFs), a BglII site between the vioC and vioD ORFs, and a BclI site between the vioD and vioE reading-frames. Since cleavage of the BamHI, BglII, and BclI sites generate compatible cohesive ends, this facilitated the construction of three different operons: vioABCE; vioABDE; vioABE. The synthetic operons are flanked by EcoRI and PstI restriction endonuclease sites, enabling the use of these two enzymes to readily subclone the entire (vioABCDE) and partial operons (vioABCE, vioABDE, vioABE) into the EcoRI and NsiI sites of pBAT4 (Fig. 1).
Production and purification of violacein and deoxyviolacein
We transformed E. coli BL21(DE3) (New England Biolabs) cells with plasmids expressing the synthetic vioABCDE (for production of both violacein and deoxyviolaein) or vioABCE (for production of deoxyviolacein alone) operons. The leaky expression from the T7 promoter in these plasmids was enough to allow sufficient synthesis of the enzymes in the violacein biosynthetic pathway.
We picked individual colonies, inoculated 50 mL cultures in 2× YT (16 g/L Tryptone, 10 g/L yeast extract, 5 g/L sodium chloride) with 100 mg/L ampicillin, and incubated for 16 h at 37 °C. These pre-cultures were inoculated into 20 L of 2xYT supplemented with 100 mg/L of ampicillin and 100 mg/L of L-tryptophan in a Sartorius Biostat Cplus fermenter. Cultures were grown for 5 h at 37 °C, with agitation (400 rpm) and air influx of 4 L/min. The temperature was then reduced to 20 °C (to avoid excessive growth and foaming overnight) and the cultures were incubated for a further 16 h. Cells were harvested by centrifugation. Violacein and deoxyviolacein were extracted by resuspending the bacterial pellets in 500 mL of 90% v/v acetone. Cell suspensions in acetone were filtered to produce crude violacein and deoxyviolacein extracts.
Violacein and deoxyviolacein purification
Deoxyviolacein
The acetone cell extract was evaporated to dryness. The crude residue was suspended in acetone and dry-loaded onto silica gel (SiO2). Deoxyviolacein was purified by column chromatography on silica gel (SiO2), first washing with petroleum ether (boiling point = 40–60 °C) and eluting with a 1:1 solution of ethyl acetate and petroleum ether. Deoxyviolacein was obtained as a purple solid and was analytically pure (100%). The identity and purity of deoxyviolacein were confirmed by 1H NMR (Additional file 1: Figure S1). We calculated the purity by integrating related peaks and comparing the areas. No other analysis was run because the data is consistent with that from the literature [46]. The apparent purity of deoxyviolacein allowed its quantitation and that of violacein (see below). Minor contamination by inorganic compounds cannot be excluded but these would need to be soluble in acetone.
Violacein
The acetone cell extract was evaporated to dryness. The crude residue was suspended in acetone with sonication and dry-loaded onto silica gel (SiO2). Violacein was purified by column chromatography on silica gel (SiO2), first washing with petroleum ether (boiling point = 40–60 °C) and eluting with 4:6, 1:1, and 6:4 solutions of ethyl acetate and petroleum ether (boiling point = 40–60 °C). Violacein was obtained as a crude mixture with approximately 12% deoxyviolacein (estimated from 1H NMR). The identity of violacein was confirmed by 1H NMR (Additional file 1: Figure S1) and was consistent with the literature [46].
Plasmodium falciparum drug sensitivity assays
We cultivated Plasmodium falciparum 3D7 and W2 strains in complete RPMI (RPMI 1640, Sigma, USA), supplemented with 10% plasma (AB−) and 2% Haematocrit (O+). Plasmodium cultures were synchronized twice with sorbitol, and drug sensitivity tests were performed on cultures enriched for ring-stage parasites. The parasitemia of the cultures was adjusted to 1% and drug sensitivity screens were performed in 96-well plates, with the following drug concentrations: for violacein and chloroquine [47] 8 concentrations of a 2× serial dilution, starting with 5 μM were employed; for deoxyviolacein, the same number of serial dilutions were tested with a starting concentration of 50 μM. All experiments were performed in triplicate and included solvent controls as well as untreated erythrocytes. After 48 h of incubation at 37 °C, cultures were labeled with SYBR® Green and analyzed by flow cytometry. IC50s were calculated using GraphPad Prism version 5.01.
Trypanosoma cruzi drug sensitivity assays
In vitro drug sensitivity assays on Trypanosoma cruzi were performed as described by Ferreira [48]. Briefly, we performed the assays using T. cruzi strain Tulahuen (parasites engineered to express E. coli β-galactosidase, lacZ [49], that catalyzes a colorimetric reaction when biologically active). Trypomastigotes were grown on monolayers of human fibroblasts, and epimastigotes were grown in liver infusion tryptone with 10% fetal calf serum, penicillin and streptomycin (to prevent contamination). Cultures assayed for β-galactosidase activity were grown in RPMI 1640 medium without phenol red plus 10% fetal calf serum, penicillin, and streptomycin.
Drug-sensitivity assays were performed in 96-well tissue culture plates (Becton Dickinson). Human fibroblasts were seeded at 2 × 103 per well in 80 μL volumes (RPMI 1640 without phenol red) and incubated overnight. The next day, β–galactosidase-expressing trypomastigotes were added at 1 × 104 per well in 20 μL of RPMI 1640 without phenol red. After 24 h, violacein or deoxyviolacein (10 mM stocks in DMSO) were added to the cultures in serial dilutions in 50 μL volumes (RPMI 1640 without phenol red). Each dilution was tested in triplicate. After 72 h of incubation, the plates were inspected under an inverted microscope to check the growth of the controls and sterility. Then, 50 μL of the substrate containing chlorophenol red-β-D-galactopyranoside (CPRG) and Nonidet P-40 (0.1% final concentration) was added to all wells. β–galactosidase activity led to a change in the color of the medium from yellow to red, a change that was quantified through measuring the absorbance at 570 nm in an automated plate reader [48]. Wells containing violacein and deoxyviolacein (without phenol red) were used to normalize for the purple color of the compounds. Data were transferred into Sigma Plot to determine IC50 values. The drug benzimidazole was used as a positive control and untreated parasite cultures were used as negative control for these assays.
Cytotoxicity of violacein and deoxyviolacein
The cytotoxicity of the compounds used in this work was evaluated using an MTT [3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide assay. MTT is a yellow tetrazole compound that is reduced to a purple formazan in the mitochondria of living cells. Hence, the proportion of living eukaryotic cells in a given culture can be quantified by monitoring its absorbance at 590 nm.
We cultured HepG2 (human hepatoma) and COS-7 (kidney from African Green Monkey) cell lines in the presence of different concentrations of the test compounds, to evaluate their cytotoxicity. We incubated 104 cells in 200 μL cultures (96-well plates) in 5% v/v CO2 at 37 °C, in RPMI with Gentamicin (40 mg/L), and 10% of heat-inactivated fetal calf serum (FCS). The final concentrations of test compounds were: violacein (10 μM, 5 μM, 2.5 μM, 1.25 μM and 0.62 μM) and deoxyviolacein (200 μM, 100 μM, 50 μM, 25 μM and 12.5 μM).
After 48 of incubation, we added 15 μL of 5 mg/L MTT and incubated for 4 h in 5% v/v CO2 at 37 °C. The plate was then centrifuged at 1500 rpm for 5 min, the supernatant discarded, the cells washed with PBS, and resuspended in 50 μL of isopropanol. Wells containing violacein and deoxyviolacein (without MTT) were used to normalize for the purple color and precipitation of the compounds.
We cultured HepG2 (human hepatoma) and CHO-745 (Chinese hamster ovary) cell lines in the presence of different concentrations of the test compounds, to evaluate their morphology. We incubated 104 cells in 200 μL cultures (96-well plates) in 5% v/v CO2 at 37 °C, in RPMI with Gentamicin (40 mg/L), and 10% of heat-inactivated fetal calf serum (FCS). The final concentrations of test compounds were: violacein (0 μM, 0.8 μM, 4 μM, 20 μM and 100 μM) and deoxyviolacein (0 μM, 0.8 μM, 4 μM, 20 μM and 100 μM).
Results
Activity of purified violacein and deoxyviolacein against T. cruzi
We evaluated the biological activity of violacein and deoxyviolacein produced in our recombinant E. coli strains expressing the complete or partial synthetic violacein operon, and found that violacein could efficiently inhibit the trypanosomatids, with an IC50 of 1.51 μM ± 0.4, whereas the IC50 for deoxyviolacein was above 50 uM. As a comparison, the IC50 determined in the same experiment for the anti T. cruzi drug benzimidazole (N-benzyl-2-(2-nitro-1H- imidazol-1-yl)acetamide) was 3.07 μM ±0.6. The IC50 values of the compounds against the T. cruzi Tulahuen strain in the in vitro assay represent the means of at least three individual experiments.
Antiplasmodial activity of purified violacein and deoxyviolacein
Initially, we tested higher concentrations (> 5 μM) of violacein in a Plasmodium sensitivity test, and noted that samples treated with 10 or 50 μM of violacein could not be analyzed through flow cytometry as all erythrocytes ruptured in the presence of those doses of the compound. Violacein was active against both chloroquine-sensitive and chloroquine-resistant Plasmodium strains (IC50 ~ 0.4 μM against 3D7 and ~ 0.5 μM against W2 parasites). Deoxyviolacein, on the other hand, showed a mild activity against Plasmodium strains when compared with violacein (IC50 ~ 11 μM against 3D7 and ~ 14 μM against W2 parasites). The chloroquine control curve performed in both strains showed IC50 values comparable with those described in the literature (Fig. 2).
Cytotoxicity of violacein and deoxyviolacein
When performing drug sensitivity assays on erythrocytes infected with P.falciparum, we noticed that healthy and infected erythrocytes ruptured when treated with 10 μM or more of violacein. In contrast, erythrocytes treated with 50 μM deoxyviolacein did not show any obvious morphological changes compared to the untreated cells. We evaluated the morphological changes in HepG2 and CHO-745 cells upon exposure to violacein and deoxyviolacein, but once more detected no morphological changes to cells treated with 20 μM deoxyviolacein and minor changes to cells treated with 100 μM deoxyviolacein (Additional file 2: Figures. S2 and S3).
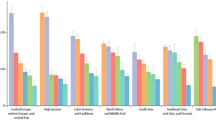
We performed viability assays to investigate the toxicity of violacein and deoxyviolacein to COS-7 and HepG2 cell lines. Our experiments confirmed the cytoxicity of violacein against both cell lines (IC50 of ~ 2.5 μM against COS-7 and ~ 1.4 μM against HepG2), with a stronger effect on the tumor cell line. In contrast, deoxyviolacein showed low toxicity against mammalian cell lines, as cells were able to grow well even in the presence of concentrations of deoxyviolacein 20 times higher then its IC50 in Plasmodium strains (Fig. 3). We were unable to test higher deoxyviolacein concentrations since it precipitated under our experimental conditions.
Cytotoxicity of violacein and deoxyviolacein. MTT assay used to investigate the cytotoxicity of violacein and deoxyviolacein to (a) COS-7 and (b) HepG2 cells, indicated that deoxyviolacein shows no measurable toxicity to mammalian cells, whereas the IC50 for violacein is approximately 2.5 μM and 1.4 μM for COS-7 and HepG2 cells, respectively
Discussion
We have constructed E. coli strains producing either a mixture of violacein or dexyviolacein alone, using enzymes encoded by synthetic operons. The biological activity of these compounds against Trypanosoma, Plasmodium and mammalian cells was assessed. The synthesis of these pigment compounds was easily monitored by their color and it was possible to observe, for example, that pigment formation was greatly enhanced by intense aeration.
We purified deoxyviolacein using the conditions described in the Methods section, achieving close to 100% purity in just a few purification steps. Violacein, however, was contaminated with approximately 12% deoxyviolacein. Hence, in all experiments where we describe the biological effects of violacein, we had some deoxyviolacein as a contaminant. As the biological activity of deoxyviolacein was consistently lower than that of violacein, we inferred that this contamination would not interfere with the interpretation of the results.
We performed in vitro assays to investigate the antiplasmodial activity of violacein and deoxyviolacein, using both 3D7 (wild-type) and W2 (chloroquine-resistant) Plasmodium falciparum strains. We found that the IC50 of violacein was ca. 0.5 μM, whereas that of deoxyviolacein was ca. 10 μM. The 3D7 and W2 strains were equally sensitive to violacein and deoxyviolacein, whilst 3D7 was around 30 times more sensitive to chloroquine (IC50 ~ 20 nM) than W2 (IC50 ~ 0.5uM). Hence, the mechanism conferring resistance to chloroquine in W2 lines did not affect their sensitivity to violacein or deoxyviolacein.
When treating Plasmodium-infected erythrocytes with violacein, we found that higher concentrations of the pigment caused the complete rupture of the red blood cells. Hence, we decided to investigate the cytotoxicity of violacein and deoxyviolacein. In spite of their very similar structure, violacein was very toxic to mammalian cells, whereas deoxyviolacein showed selective toxicity against Plasmodium (the parasite was at least 20× more sensitive to this compound than were the mammalian cells) than violacein (the parasite showing only ca. 5× greater sensitivity to this compound than did mammalian cells). The IC50 of violacein produced from our synthetic operon is about ~ 1.4 μM for HepG2 and ~ 2.5 μM for COS-7 cell lines, in agreement with published results, indicating a degree of specificity of the compound against cancer cells. On the other hand, the HepG2 cell line showed no significant viability loss when treated with deoxyviolacein concentrations close to 200 μM. When treating the Trypanosoma cruzi Tulahuen strain with violacein and deoxyviolacein, we also observed much higher biological activity with violacein (IC50 of 1.51 μM ± 0.4) than with deoxyviolacein (IC50 > 50 μM).
Conclusions
We have utilized synthetic operons encoding enzymes for complete or partial pathways for the biosynthesis of violacein or deoxyviolacein in E. coli strains, and investigated the biological activity of the produsts. Deoxyviolacein, in spite of its lower antiparasitic activity, might be a better starting point than violacein for the development of a novel antiparasitic drug due to its low toxicity to human cells. However, it is important to note that we did not investigate the import of either compound by the target cells, and so cannot exclude the possibility that deoxyviolacein’s lower toxicity was due to an inefficient import into mammalian or parasite cells.
References
Choi SY, Yoon KH, Lee JI, Mitchell RJ. Violacein: properties and production of a versatile bacterial pigment. Biomed Res Int. 2015;2015:465056.
Masuelli L, Pantanella F, La Regina G, Benvenuto M, Fantini M, Mattera R, Di Stefano E, Mattei M, Silvestri R, Schippa S, et al. Violacein, an indole-derived purple-colored natural pigment produced by Janthinobacterium lividum, inhibits the growth of head and neck carcinoma cell lines both in vitro and in vivo. Tumour Biol. 2016;37(3):3705–17.
Sasidharan A, Sasidharan NK, Amma DB, Vasu RK, Nataraja AV, Bhaskaran K. Antifungal activity of violacein purified from a novel strain of Chromobacterium sp. NIIST (MTCC 5522). J Microbiol. 2015;53(10):694–701.
Smith HJ, Foreman CM, Akiyama T, Franklin MJ, Devitt NP, Ramaraj T. Genome sequence of janthinobacterium sp. CG23_2, a violacein-producing isolate from an Antarctic supraglacial stream. Genome Announc. 2016;4(1).
Myeong NR, Seong HJ, Kim HJ, Sul WJ. Complete genome sequence of antibiotic and anticancer agent violacein producing Massilia sp. strain NR 4-1. J Biotechnol. 2016;223:36–7.
Lu Y, Wang L, Xue Y, Zhang C, Xing XH, Lou K, Zhang Z, Li YD, Zhang G, Bi J, et al. Production of violet pigment by a newly isolated psychrotrophic bacterium from a glacier in Xinjiang China. Biochem Eng J. 2009;43(2):131–41.
Duran N, Justo GZ, Ferreira CV, Melo PS, Cordi L, Martins D. Violacein: properties and biological activities. Biotechnol Appl Biochem. 2007;48(Pt 3):127–33.
Antonisamy P, Kannan P, Ignacimuthu S. Anti-diarrhoeal and ulcer-protective effects of violacein isolated from Chromobacterium violaceum in Wistar rats. Fundam Clin Pharmacol. 2009;23(4):483–90.
Lichstein HC, VANdS VF. The antibiotic activity of violacein, prodigiosin, and phth iocol. J Bacteriol. 1946;52:145.
Aruldass CA, Masalamany SR, Venil CK, Ahmad WA: Antibacterial mode of action of violacein from Chromobacterium violaceum UTM5 against Staphylococcus aureus and methicillin-resistant Staphylococcus aureus (MRSA). Environ Sci Pollut Res Int. 2018;25(6):5164–5180. https://doi.org/10.1007/s11356-017-8855-2. Epub 2017 Mar 31.
Dodou HV, de Morais Batista AH, Sales GWP, de Medeiros SC, Rodrigues ML, Nogueira PCN, Silveira ER, Nogueira NAP. Violacein antimicrobial activity on Staphylococcus epidermidis and synergistic effect on commercially available antibiotics. J Appl Microbiol. 2017;123(4):853–60.
Becker MH, Brucker RM, Schwantes CR, Harris RN, Minbiole KP. The bacterially produced metabolite violacein is associated with survival of amphibians infected with a lethal fungus. Appl Environ Microbiol. 2009;75(21):6635–8.
Duran N, Campos V, Riveros R, Joyas A, Pereira MF, Haun M. Bacterial chemistry-III: preliminary studies on trypanosomal activities of Chromobacterium violaceum products. Anais da Academia Brasileira de Ciencias. 1989;61(1):31–6.
Duran N, Antonio RV, Haun M, Pilli RA. Biosynthesis of a trypanocide by Chromobacterium violaceum. World J Microbiol Biotechnol. 1994;10(6):686–90.
Leon LL, Miranda CC, De Souza AO, Duran N. Antileishmanial activity of the violacein extracted from Chromobacterium violaceum. J Antimicrob Chemother. 2001;48(3):449–50.
Ballestriero F, Daim M, Penesyan A, Nappi J, Schleheck D, Bazzicalupo P, Di Schiavi E, Egan S. Antinematode activity of Violacein and the role of the insulin/IGF-1 pathway in controlling violacein sensitivity in Caenorhabditis elegans. PLoS One. 2014;9(10):e109201.
Andrighetti-Frohner CR, Antonio RV, Creczynski-Pasa TB, Barardi CR, Simoes CM. Cytotoxicity and potential antiviral evaluation of violacein produced by Chromobacterium violaceum. Memorias do Instituto Oswaldo Cruz. 2003;98(6):843–8.
de Carvalho DD, Costa FT, Duran N, Haun M. Cytotoxic activity of violacein in human colon cancer cells. Toxicol in Vitro : an international journal published in association with BIBRA. 2006;20(8):1514–21.
Lopes SC, Blanco YC, Justo GZ, Nogueira PA, Rodrigues FL, Goelnitz U, Wunderlich G, Facchini G, Brocchi M, Duran N, et al. Violacein extracted from Chromobacterium violaceum inhibits Plasmodium growth in vitro and in vivo. Antimicrob Agents Chemother. 2009;53(5):2149–52.
Riveros R, Haun M, Duran N. Effect of growth conditions on production of violacein by Chromobacterium violaceum (BB-78 strain). Braz. J. Med. Biol. Res. = Rev. Bras. Pesqui. Med. Biol. / Sociedade Bras. Biol. [et al]. 1989;22(5):569–77.
Blosser RS, Gray KM. Extraction of violacein from Chromobacterium violaceum provides a new quantitative bioassay for N-acyl homoserine lactone autoinducers. J Microbiol Methods. 2000;40(1):47–55.
Antonio RV, Creczynski-Pasa TB. Genetic analysis of violacein biosynthesis by Chromobacterium violaceum. Genet. Mol. Res. : GMR. 2004;3(1):85–91.
Pantanella F, Berlutti F, Passariello C, Sarli S, Morea C, Schippa S. Violacein and biofilm production in Janthinobacterium lividum. J Appl Microbiol. 2007;102(4):992–9.
Ambrozic Avgustin J, Zgur Bertok D, Kostanjsek R, Avgustin G. Isolation and characterization of a novel violacein-like pigment producing psychrotrophic bacterial species Janthinobacterium svalbardensis sp. nov. Antonie Van Leeuwenhoek. 2013;103(4):763–9.
Ti TY, Tan WC, Chong AP, Lee EH. Nonfatal and fatal infections caused by Chromobacterium violaceum. Clin. Infect. Dis. : an official publication of the Infectious Diseases Society of America. 1993;17(3):505–7.
Ponte R, Jenkins SG. Fatal Chromobacterium violaceum infections associated with exposure to stagnant waters. Pediatr Infect Dis J. 1992;11(7):583–6.
Patijanasoontorn B, Boonma P, Wilailackana C, Sitthikesorn J, Lumbiganon P, Chetchotisakd P, Noppawinyoowong C, Simajareuk K. Hospital acquired Janthinobacterium lividum septicemia in Srinagarind Hospital. J. Med. Assoc. Thai. = Chotmaihet thangphaet. 1992;75(Suppl 2):6–10.
Farrell K, Van Werkhooven M, Gratten M, Aiken GH. Chromobacterium violaceum infections. Report of a fatal case. P. N. G. Med. J. 1979;22(4):94–5.
Jiang PX, Wang HS, Zhang C, Lou K, Xing XH. Reconstruction of the violacein biosynthetic pathway from Duganella sp. B2 in different heterologous hosts. Appl Microbiol Biotechnol. 2010;86(4):1077–88.
Fang MY, Zhang C, Yang S, Cui JY, Jiang PX, Lou K, Wachi M, Xing XH. High crude violacein production from glucose by Escherichia coli engineered with interactive control of tryptophan pathway and violacein biosynthetic pathway. Microb Cell Factories. 2015;14:8.
Chuang J, Boeke JD, Mitchell LA. Coupling yeast golden gate and VEGAS for efficient assembly of the violacein Pathway in saccharomyces cerevisiae. Methods Mol Biol. 2018;1671:211–25.
Rodrigues AL, Trachtmann N, Becker J, Lohanatha AF, Blotenberg J, Bolten CJ, Korneli C, de Souza Lima AO, Porto LM, Sprenger GA, et al. Systems metabolic engineering of Escherichia coli for production of the antitumor drugs violacein and deoxyviolacein. Metab Eng. 2013;20:29–41.
Rodrigues AL, Gocke Y, Bolten C, Brock NL, Dickschat JS, Wittmann C. Microbial production of the drugs violacein and deoxyviolacein: analytical development and strain comparison. Biotechnol Lett. 2012;34(4):717–20.
Rodrigues AL, Becker J, de Souza Lima AO, Porto LM, Wittmann C. Systems metabolic engineering of Escherichia coli for gram scale production of the antitumor drug deoxyviolacein from glycerol. Biotechnol Bioeng. 2014;111(11):2280–9.
Sanchez C, Brana AF, Mendez C, Salas JA. Reevaluation of the violacein biosynthetic pathway and its relationship to indolocarbazole biosynthesis. Chembiochem : a European journal of chemical biology. 2006;7(8):1231–40.
Brady SF, Chao CJ, Handelsman J, Clardy J. Cloning and heterologous expression of a natural product biosynthetic gene cluster from eDNA. Org Lett. 2001;3(13):1981–4.
August PR, Grossman TH, Minor C, Draper MP, MacNeil IA, Pemberton JM, Call KM, Holt D, Osburne MS. Sequence analysis and functional characterization of the violacein biosynthetic pathway from Chromobacterium violaceum. J Mol Microbiol Biotechnol. 2000;2(4):513–9.
Fuller JJ, Ropke R, Krausze J, Rennhack KE, Daniel NP, Blankenfeldt W, Schulz S, Jahn D, Moser J. Biosynthesis of violacein, structure and function of l-Tryptophan Oxidase VioA from chromobacterium violaceum. J Biol Chem. 2016;291(38):20068–84.
Shinoda K, Hasegawa T, Sato H, Shinozaki M, Kuramoto H, Takamiya Y, Sato T, Nikaidou N, Watanabe T, Hoshino T. Biosynthesis of violacein: a genuine intermediate, protoviolaceinic acid, produced by VioABDE, and insight into VioC function. Chem Commun (Camb). 2007;40:4140–2.
Wilkinson B, Micklefield J. Mining and engineering natural-product biosynthetic pathways. Nat Chem Biol. 2007;3(7):379–86.
Wang HS, Wang FZ, Zhu XF, Yan YC, Yu XH, Jiang PX, Xing XH. Biosynthesis and characterization of violacein, deoxyviolacein and oxyviolacein in heterologous host, and their antimicrobial activities. Biochem Eng J. 2012;67:148–55.
He L, Xiu Y, Jones JA, Baidoo EE, Keasling JD, Tang YJ, Koffas MA. Deciphering flux adjustments of engineered E. coli cells during fermentation with changing growth conditions. Metab Eng. 2017;39:247–56.
Sun H, Zhao D, Xiong B, Zhang C, Bi C. Engineering Corynebacterium glutamicum for violacein hyper production. Microb Cell Factories. 2016;15(1):148.
Jones JA, Vernacchio VR, Lachance DM, Lebovich M, Fu L, Shirke AN, Schultz VL, Cress B, Linhardt RJ, Koffas MA. ePathOptimize: a combinatorial approach for transcriptional balancing of metabolic pathways. Sci Rep. 2015;5:11301.
Xu P, Rizzoni EA, Sul SY, Stephanopoulos G. Improving metabolic pathway efficiency by statistical model-based multivariate regulatory metabolic engineering. ACS Synth Biol. 2017;6(1):148–58.
Wille G, Steglich W. A short synthesis of the bacterial pigments violacein and deoxyviolacein. Synthesis. 2001;5:759–62.
Millet J, Alibert S, Torrentino-Madamet M, Rogier C, Santelli-Rouvier C, Bigot P, Mosnier J, Baret E, Barbe J, Parzy D, et al. Polymorphism in Plasmodium falciparum drug transporter proteins and reversal of in vitro chloroquine resistance by a 9,10-dihydroethanoanthracene derivative. Antimicrob Agents Chemother. 2004;48(12):4869–72.
Ferreira RS, Dessoy MA, Pauli I, Souza ML, Krogh R, Sales AI, Oliva G, Dias LC, Andricopulo AD. Synthesis, biological evaluation, and structure-activity relationships of potent noncovalent and nonpeptidic cruzain inhibitors as anti-Trypanosoma cruzi agents. J Med Chem. 2014;57(6):2380–92.
Buckner FS, Verlinde CL, La Flamme AC, Van Voorhis WC. Efficient technique for screening drugs for activity against Trypanosoma cruzi using parasites expressing beta-galactosidase. Antimicrob Agents Chemother. 1996;40(11):2592–7.
Acknowledgements
Not applicable.
Funding
This study has supported by Sao Paulo Research Foundation (FAPESP) Grant 2012/16525-2 to FTMC and Grant 2015/03553-6 to EB, contract from the European Commission under the FP7 Collaborative Program, UNICELLSYS to SGO, UK Biotechnology and Biological Sciences Research Council (BB/F008228/1) to SGO and The Bill and Melinda Gates foundation (OPP1087646 to EB and SGO). TAT was supported by a CNPq fellowship and FTMC is a CNPq research fellow level 1C.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Author information
Authors and Affiliations
Contributions
JA provided the original synthetic violacein operon and advised on violacein expression. EB subcloned the complete and partial violacein operons and performed large-scale crude violacein and deoxyviolacein preparations. AR optimized violacein and deoxyviolacein extractions and JES and DRS planned and performed compound purification. EB, FTMC and TAT planned and performed the Plasmodium falciparum and mammalian cytotoxicity in vitro assays. RK and AA designed and performed the Trypanosoma cruzi in vitro assays. EB and SGO conceived the project and wrote the manuscript. All authors read, edited and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional files
Additional file 1:
Figure S1. Chemical structure (www.chemspider.com) and 1H spectra of Violacein and Deoxyviolacein. Chemical structure (www.chemspider.com) and 1H spectra of Violacein and Deoxyviolacein. (PDF 216 kb)
Additional file 2:
Figure S2. Cytotoxicity of deoxyviolacein and violacein to HepG2 cells. Morphological changes of HepG2 cells treated with 0.8 to 100 uM of deoxyviolacein and violacein. Figure S3. Cytotoxicity of deoxyviolacein and violacein to CHO-745 cells. Morphological changes of CHO-745 cells treated with 0.8 to 100 uM of deoxyviolacein and violacein. (PDF 4205 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Bilsland, E., Tavella, T.A., Krogh, R. et al. Antiplasmodial and trypanocidal activity of violacein and deoxyviolacein produced from synthetic operons. BMC Biotechnol 18, 22 (2018). https://doi.org/10.1186/s12896-018-0428-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12896-018-0428-z