Abstract
Background
Abundant pseudogenes are a feature of mammalian genomes. Processed pseudogenes (PPs) are reverse transcribed from mRNAs. Recent molecular biological studies show that mammalian long interspersed element 1 (L1)-encoded proteins may have been involved in PP reverse transcription. Here, we present the first comprehensive analysis of human PPs using all known human genes as queries.
Results
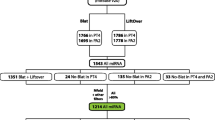
The human genome was queried and 3,664 candidate PPs were identified. The most abundant were copies of genes encoding keratin 18, glyceraldehyde-3-phosphate dehydrogenase and ribosomal protein L21. A simple method was developed to estimate the level of nucleotide substitutions (and therefore the age) of PPs. A Poisson-like age distribution was obtained with a mean age close to that of the Alu repeats, the predominant human short interspersed elements. These data suggest a nearly simultaneous burst of PP and Alu formation in the genomes of ancestral primates. The peak period of amplification of these two distinct retrotransposons was estimated to be 40-50 million years ago. Concordant amplification of certain L1 subfamilies with PPs and Alus was observed.
Conclusions
We suggest that a burst of formation of PPs and Alus occurred in the genome of ancestral primates. One possible mechanism is that proteins encoded by members of particular L1 subfamilies acquired an enhanced ability to recognize cytosolic RNAs in trans.







Similar content being viewed by others
References
Vanin EF: Processed pseudogenes: characteristics and evolution. Annu Rev Genet. 1985, 19: 253-272. 10.1146/annurev.ge.19.120185.001345.
Mighell AJ, Smith NR, Robinson PA, Markham AF: Vertebrate pseudogenes. FEBS Lett. 2000, 468: 109-114. 10.1016/S0014-5793(00)01199-6.
Gonçalves I, Duret L, Mouchiroud D: Nature and structure of human genes that generate retropseudogenes. Genome Res. 2000, 10: 672-678. 10.1101/gr.10.5.672.
Harrison PM, Hegyi H, Balasubramanian S, Luscombe NM, Bertone P, Echols N, Johnson T, Gerstein M: Molecular fossils in the human genome: identification and analysis of the pseudogenes in chromosomes 21 and 22. Genome Res. 2002, 12: 272-280. 10.1101/gr.207102.
Chen C, Gentles AJ, Jurka J, Karlin S: Genes, pseudogenes, and Alu sequence organization across human chromosomes 21 and 22. Proc Natl Acad Sci USA. 2002, 99: 2930-2935. 10.1073/pnas.052692099.
Brosius J: RNAs from all categories generate retrosequences that may be exapted as novel genes or regulatory elements. Gene. 1999, 238: 115-134. 10.1016/S0378-1119(99)00227-9.
Lahn BT, Page DC: Retroposition of autosomal mRNA yielded testis-specific gene family on human Y chromosome. Nat Genet. 1999, 21: 429-433. 10.1038/7771.
Betrán E, Wang W, Jin L, Long M: Evolution of the phosphoglycerate mutase processed gene in human and chimpanzee revealing the origin of a new primate gene. Mol Biol Evol. 2002, 19: 654-663.
Weiner AM, Deininger PL, Efstratiadis A: Nonviral retroposons: genes, pseudogenes, and transposable elements generated by the reverse flow of genetic information. Annu Rev Biochem. 1986, 55: 631-661. 10.1146/annurev.bi.55.070186.003215.
Okada N: SINEs: Short interspersed repeated elements of the eukaryotic genome. Trends Ecol Evol. 1991, 6: 358-361. 10.1016/0169-5347(91)90226-N.
Smit AF: The origin of interspersed repeats in the human genome. Curr Opin Genet Dev. 1996, 6: 743-748. 10.1016/S0959-437X(96)80030-X.
Okada N, Hamada M, Ogiwara I, Ohshima K: SINEs and LINEs share common 3' sequences: a review. Gene. 1997, 205: 229-243. 10.1016/S0378-1119(97)00409-5.
Weiner AM: SINEs and LINEs: the art of biting the hand that feeds you. Curr Opin Cell Biol. 2002, 14: 343-350. 10.1016/S0955-0674(02)00338-1.
International Human Genome Sequencing Consortium: Initial sequencing and analysis of the human genome. Nature. 2001, 409: 860-921. 10.1038/35057062.
Moran JV, Holmes SE, Naas TP, DeBerardinis RJ, Boeke JD, Kazazian HH: High frequency retrotransposition in cultured mammalian cells. Cell. 1996, 87: 917-927.
Kazazian HH, Moran JV: The impact of L1 retrotransposons on the human genome. Nat Genet. 1998, 19: 19-24.
Jurka J: Sequence patterns indicate an enzymatic involvement in integration of mammalian retroposons. Proc Natl Acad Sci USA. 1997, 94: 1872-1877. 10.1073/pnas.94.5.1872.
Esnault C, Maestre J, Heidmann T: Human LINE retrotransposons generate processed pseudogenes. Nat Genet. 2000, 24: 363-367. 10.1038/74184.
Wei W, Gilbert N, Ooi SL, Lawler JF, Ostertag EM, Kazazian HH, Boeke JD, Moran JV: Human L1 retrotransposition: cis preference versus trans complementation. Mol Cell Biol. 2001, 21: 1429-1439. 10.1128/MCB.21.4.1429-1439.2001.
Pavlícek A, Paces J, Elleder D, Hejnar J: Processed pseudogenes of human endogenous retroviruses generated by LINEs: their integration, stability, and distribution. Genome Res. 2002, 12: 391-399. 10.1101/gr.216902. Article published online before print in February 2002.
Dewannieux M, Esnault C, Heidmann T: LINE-mediated retrotransposition of marked Alu sequences. Nat Genet. 2003, 35: 41-48. 10.1038/ng1223.
Britten RJ: Evidence that most human Alu sequences were inserted in a process that ceased about 30 million years ago. Proc Natl Acad Sci USA. 1994, 91: 6148-6150.
Kapitonov V, Jurka J: The age of Alu subfamilies. J Mol Evol. 1996, 42: 59-65.
Sarrowa J, Chang DY, Maraia RJ: The decline in human Alu retroposition was accompanied by an asymmetric decrease in SRP9/14 binding to dimeric Alu RNA and increased expression of small cytoplasmic Alu RNA. Mol Cell Biol. 1997, 17: 1144-1151.
Schmid CW: Does SINE evolution preclude Alu function?. Nucleic Acids Res. 1998, 26: 4541-4550. 10.1093/nar/26.20.4541.
Batzer MA, Deininger PL: Alu repeats and human genomic diversity. Nat Rev Genet. 2002, 3: 370-379. 10.1038/nrg798.
Boeke JD: LINEs and Alus - the polyA connection. Nat Genet. 1997, 16: 6-7.
Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, Sutton GG, Smith HO, Yandell M, Evans CA, Holt RA, et al: The sequence of the human genome. Science. 2001, 291: 1304-1351. 10.1126/science.1058040.
Hubbard T, Barker D, Birney E, Cameron G, Chen Y, Clark L, Cox T, Cuff J, Curwen V, Down T, et al: The Ensembl genome database project. Nucleic Acids Res. 2002, 30: 38-41. 10.1093/nar/30.1.38.
Hogenesch JB, Ching KA, Batalov S, Su AI, Walker JR, Zhou Y, Kay SA, Schultz PG, Cooke MP: A comparison of the Celera and Ensembl predicted gene sets reveals little overlap in novel genes. Cell. 2001, 106: 413-415.
Hesse M, Magin TM, Weber K: Genes for intermediate filament proteins and the draft sequence of the human genome: novel keratin genes and a surprisingly high number of pseudogenes related to keratin genes 8 and 18. J Cell Sci. 2001, 114: 2569-2575.
Zhang Z, Harrison P, Gerstein M: Identification and analysis of over 2000 ribosomal protein pseudogenes in the human genome. Genome Res. 2002, 12: 1466-1482. 10.1101/gr.331902.
Sved J, Bird A: The expected equilibrium of the CpG dinucleotide in vertebrate genomes under a mutation model. Proc Natl Acad Sci USA. 1990, 87: 4692-4696.
Graur D, Li W-H: Fundamentals of Molecular Evolution. 2000, Sunderland, MA: Sinauer Associates, 2
Nachman MW, Crowell SL: Estimate of the mutation rate per nucleotide in humans. Genetics. 2000, 156: 297-304.
Shoshani J, Groves CP, Simons EL, Gunnell GF: Primate phylogeny: morphological vs. molecular results. Mol Phylogenet Evol. 1996, 5: 102-154. 10.1006/mpev.1996.0009.
Kay RF, Ross C, Williams BA: Anthropoid origins. Science. 1997, 275: 797-804. 10.1126/science.275.5301.797.
Smit AF, Tóth G, Riggs AD, Jurka J: Ancestral, mammalian-wide subfamilies of LINE-1 repetitive sequences. J Mol Biol. 1995, 246: 401-417. 10.1006/jmbi.1994.0095.
Boissinot S, Furano AV: Adaptive evolution in LINE-1 retrotransposons. Mol Biol Evol. 2001, 18: 2186-2194.
Yoshihama M, Uechi T, Asakawa S, Kawasaki K, Kato S, Higa S, Maeda N, Minoshima S, Tanaka T, Shimizu N, Kenmochi N: The human ribosomal protein genes: sequencing and comparative analysis of 73 genes. Genome Res. 2002, 12: 379-390. 10.1101/gr.214202.
Kimura M: The Neutral Theory of Molecular Evolution. 1983, Cambridge: Cambridge University Press
Takahata N: Molecular phylogeny and demographic history of humans. In Humanity from African Naissance to Coming Millennia. Edited by: Tobias PV, Raath MA, Moggi-Cecchi J, Doyle GA. 2001, Firenze: Firenze University Press, 299-305.
Kajikawa M, Okada N: LINEs mobilize SINEs in the eel through a shared 3' sequence. Cell. 2002, 111: 433-444.
Ivics Z, Hackett PB, Plasterk RH, Izsvák Z: Molecular reconstruction of Sleeping Beauty, a Tc1-like transposon from fish, and its transposition in human cells. Cell. 1997, 91: 501-510.
Chou H-H, Hayakawa T, Diaz S, Krings M, Indriati E, Leakey M, Paabo S, Satta Y, Takahata N, Varki A: Inactivation of CMP-N-acetylneuraminic acid hydroxylase occurred prior to brain expansion during human evolution. Proc Natl Acad Sci USA. 2002, 99: 11736-11741. 10.1073/pnas.182257399.
Kazazian HH: An estimated frequency of endogenous insertional mutations in humans. Nat Genet. 1999, 22: 130-10.1038/9638.
Zhang J, Rosenberg HF, Nei M: Positive Darwinian selection after gene duplication in primate ribonuclease genes. Proc Natl Acad Sci USA. 1998, 95: 3708-3713. 10.1073/pnas.95.7.3708.
Suga H, Koyanagi M, Hoshiyama D, Ono K, Iwabe N, Kuma K, Miyata T: Extensive gene duplication in the early evolution of animals before the parazoan-eumetazoan split demonstrated by G proteins and protein tyrosine kinases from sponge and hydra. J Mol Evol. 1999, 48: 646-653.
Gu X, Wang Y, Gu J: Age distribution of human gene families shows significant roles of both large- and small-scale duplications in vertebrate evolution. Nat Genet. 2002, 31: 205-209. 10.1038/ng902.
Kent WJ, Haussler D: Assembly of the working draft of the human genome with GigAssembler. Genome Res. 2001, 11: 1541-1548. 10.1101/gr.183201.
Kent WJ: BLAT - the BLAST-like alignment tool. Genome Res. 2002, 12: 656-664. 10.1101/gr.229202. Article published online before March 2002.
Hattori M, Fujiyama A, Taylor TD, Watanabe H, Yada T, Park HS, Toyoda A, Ishii K, Totoki Y, Choi DK, et al: The DNA sequence of human chromosome 21. Nature. 2000, 405: 311-319. 10.1038/35012518.
Human Genome Research Group: Chromosome 21. [http://hgp.gsc.riken.go.jp/data_tools/chr21.html]
Dunham I, Shimizu N, Roe BA, Chissoe S, Hunt AR, Collins JE, Bruskiewich R, Beare DM, Clamp M, Smink LJ, et al: The DNA sequence of human chromosome 22. Nature. 1999, 402: 489-495. 10.1038/990031.
Human chromosome 22 project overview. [http://www.sanger.ac.uk/HGP/Chr22]
UCSC genome bioinformatics. [http://www.genome.ucsc.edu]
Repbase update. [http://www.girinst.org/Repbase_Update.html]
Nei M, Gojobori T: Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol. 1986, 3: 418-426.
Ophir R, Itoh T, Graur D, Gojobori T: A simple method for estimating the intensity of purifying selection in protein-coding genes. Mol Biol Evol. 1999, 16: 49-53.
Bustamante CD, Nielsen R, Hartl DL: A maximum likelihood method for analyzing pseudogene evolution: implications for silent site evolution in humans and rodents. Mol Biol Evol. 2002, 19: 110-117.
NCBI Reference sequences. [http://www.ncbi.nlm.nih.gov/RefSeq/]
Saitou N, Nei M: The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987, 4: 406-425.
Acknowledgements
We thank Katsuhiko Murakami (RIKEN-GSC) for helpful discussions and Kei-ichi Kuma and Takashi Miyata (Kyoto University) for providing the data on the average nucleotide substitution rates of 31 pairs of human PPs. This work was partially supported by the Ministry of Education, Culture, Sports, Science and Technology of Japan, Grant-in-Aid for Scientific Research. This work was also supported by a grant from BIRD of Japan Science and Technology Corporation (JST) for K.O.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ohshima, K., Hattori, M., Yada, T. et al. Whole-genome screening indicates a possible burst of formation of processed pseudogenes and Alu repeats by particular L1 subfamilies in ancestral primates. Genome Biol 4, R74 (2003). https://doi.org/10.1186/gb-2003-4-11-r74
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1186/gb-2003-4-11-r74