Abstract
Background
Circadian rhythms are known to influence a variety of biological phenomena such as cell cycle, sleep-wake rhythm, hormone release and other important physiological functions. Given that cell cycle entry of hibernating hematopoietic stem cells (HSCs) plays a critical role in controlling hematopoiesis, we asked functional significance of the clock gene Bmal1, which plays a central role in regulating circadian rhythms as a transcription factor. Here we investigated the necessity of Bmal1 for HSC functions using Bmal1 deficient (Bmal1 −/−) mice.
Findings
Using colony-forming assays in vitro, we found that the frequency of mixed colony formation between Bmal1 +/+ and Bmal1 −/− CD34−KSL cells does not differ significantly. Competitive bone marrow assays also revealed that Bmal1 −/− bone marrow cells competed normally with wild-type cells and displayed long-term multi-hematopoietic lineage reconstitution. In addition, there were no significant differences in the frequencies and hibernation state of bone marrow HSCs between Bmal1 +/+ and Bmal1 −/− mice, suggesting that they are independent of circadian rhythms.
Conclusions
This paper discusses the necessity of circadian rhythms for HSC functions. Our data clearly shows that a key circadian clock gene Bmal1 is dispensable for intrinsic functions of HSCs, such as differentiation, proliferation and repopulating ability.
Similar content being viewed by others
Findings
Background
Hematopoietic stem cells (HSCs) reside in specialized bone marrow (BM) microenvironments, called niches, providing the entire range of blood cells throughout the lifespan [1, 2]. We have recently demonstrated that non-myelinating Schwann cells induce hibernation of HSCs in mouse BM [3, 4]. Occasionally, most HSCs in the BM niche come out of hibernation and undergo cell division on average every one to two months [5, 6]. Although the molecular mechanisms underlying re-entry into the cell cycle remain obscure, recent evidence suggests that the circadian clock regulates HSC trafficking between the BM and peripheral blood (PB) via the sympathetic nervous system [7]. To address the relationship between circadian oscillation and HSC hibernation in BM hematopoiesis, we here considered the possibility that the circadian transcription factor Bmal1 [8] is involved in BM hematopoiesis. Accumulating evidences have suggested that BMAL1 forms heterodimers with CLOCK, binds to E-box sequences in the promoter region and regulates the transcription of a number of clock-controlled genes. We therefore examined differentiation, proliferation and repopulating capacity of HSCs in Bmal1 deficient (Bmal1 −/−) mice which demonstrate complete loss of circadian behavioral rhythms [9] and only half life span of wild-type mice [10]. Our findings led to the conclusion, however, that Bmal1 is dispensable for differentiation, proliferation and repopulating ability of murine HSCs.
Materials and methods
Mice
C57BL/6-Ly5.1 (B6-Ly5.1) and C57BL/6-Ly5.1/5.2-F1 (B6-F1) mice were purchased from Sankyo-Lab Service (Tsukuba, Japan). Bmal1 −/− mice were obtained by mating Bmal1 +/− mice [11] bred and maintained in the Animal Research Facility of the Institute of Medical Science, the University of Tokyo. Animal care in our laboratory was in accord with the guidelines of the University of Tokyo for animal and recombinant DNA experiments.
CFU-Cs assay
PB mononuclear cells were isolated from 400 μl PB on Ficoll-Paque PLUS (GE Healthcare, Buckinghamshire, England) and CFU-Cs assays were performed using MethoCult (STEMCELL Technologies, Vancouver, Canada) according to manufacturer’s protocols. On day 11 of culture, colonies were observed under light microscopy.
Purification of murine HSCs
Mouse CD34−KSL HSCs were purified from BM cells of 8-10-week-old mice. The cells were stained with an antibody cocktail consisting of biotinylated anti-Gr-1, −Mac-1, −CD4, −IL-7R, and -Ter-119 (eBioscience, San Diego, CA), and -B220 and -CD8 monoclonal antibodies (BioLegend, San Diego, CA) (lineage-marker cocktail). Lineage-positive cells were depleted with anti-Biotin MicroBeads (Miltenyi Biotec, Auburn, CA) and LS columns (Miltenyi Biotec). The remaining cells were further stained with fluorescein isothiocyanate (FITC)-conjugated anti-CD34 (BD Bioscience, California, CA), phycoerythrin (PE)-conjugated anti-Sca-1 (eBioscience), and allophycocyanin (APC)-conjugated anti-c-Kit antibodies (BioLegend). Biotinylated antibodies were detected with streptavidin-APC-Cy7 (BioLegend). Analysis and cell sorting were performed on a MoFlo using Summit software (Dako, Glostrup, Denmark) and results were analyzed with FlowJo software (Tree Star, Ashland, OR).
Colony assays and single-cell cultures
CD34−KSL HSCs were clonally deposited into 96-well micro-titer plates containing 200 μl of S-Clone SF-03 (Sanko Junyaku Inc, Tokyo, Japan) supplemented with 10% BSA and cytokines (50 ng/ml mouse SCF, 50 ng/ml human TPO, 20 ng/ml mouse IL-3 and 2 U/ml mouse EPO for colony assays; 50 ng/ml mSCF, 50 ng/ml hTPO for proliferation assays). Colonies were recovered on day 11 of culture, cytospun onto slide glasses and subjected to Hemacolor staining (MERCK, Darmstadt, Germany) for morphological examination. To observe proliferation potential of CD34−KSL cells, cells were counted under light microscopy.
Competitive repopulation assays
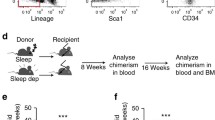
Competitive repopulation assays were performed using the Ly5 congenic mouse system. 1 × 106 BM cells from Bmal1 +/+ or Bmal1 −/− mice (B6-Ly5.2) and the same number of BM competitor cells from B6-F1 mice were transplanted into B6-Ly5.1 mice irradiated at a dose of 9.5 Gy. After transplantation, PB cells of the recipients were stained with PE-conjugated anti-Ly5.1 (BioLegend) and FITC-conjugated anti-Ly5.2 (BD Bioscience). The cells were further stained with PE-Cy7-conjugated anti-Mac-1 and -Gr-1, Pacific Blue (PB)-conjugated anti-B220 and APC-Cy7-conjugated anti-CD3 antibodies (BioLegend) and then analyzed on a FACS Aria (BD Bioscience). The second BMT was performed by transferring 1 × 106 BM cells from femora and tibiae of the primary recipient mice into lethally irradiated Ly5.1 mice. PB cells from the secondary recipient mice were analyzed 4, 8 and 12 weeks after the second BMT.
Cell cycle assays
To analyze the G0 phase, cells were incubated with 1 μg/ml Pyronin Y (Sigma-Aldrich, Saint Louis, Missouri) at 37°C for 30 min and analyzed on a FACS Aria. To investigate the turnover rate of CD34−KSL cells, EdU (invitrogen) was administered continuously to mice in the drinking water (0.5 mg/ml). After 3 weeks, BM cells were assessed with a Click-iT EdU PB Flow Cytometry Assay Kit (invitrogen) according to manufacturer’s protocols and analyzed on a FACS Aria.
White blood cell differentiation
PB cells of 10 or 40-week-old Bmal1 +/+ or Bmal1 −/− mice were stained with PE-conjugated anti-Gr-1, APC-conjugated anti-CD4, FITC-conjugated anti-CD8 (eBioscience), PE-Cy7-conjugated anti-Mac-1, PB-conjugated anti-B220 and APC-Cy7-conjugated anti-CD3 antibodies and then analyzed on a FACS Aria.
Results and discussion
Bmal1 −/− HSCs exhibit comparable differentiation and proliferation potentials in vitro
It has been shown that the mobilization of hematopoietic stem and progenitor cells (HSPCs) from BM is regulated by circadian clock [7]. We therefore considered the possibility that the circadian transcription factor Bmal1 is involved with BM hematopoiesis. Indeed we could detect oscillating CFU-Cs of HSPCs in PB of Bmal1 +/+ mice at Zeitgeber time (ZT) 5 and ZT17, but there were no statistically significant fluctuations in case of Bmal1 −/− mice (Additional file 1: Figure S1A). Thus, oscillating CFU-Cs of HSPCs appear to be regulated by circadian clock, however, it is unclear how Bmal1 affects intrinsic functions of HSCs such as differentiation, proliferation and repopulating capacity. We therefore asked to investigate and clarify these problems.
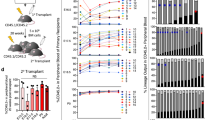
For the present investigation of effects of Bmal1 absence on differentiation of HSCs, we performed colony-forming assays in vitro in which freshly isolated Bmal1 +/+ and Bmal1 −/− CD34−KSL cells were cultured for 11 days in medium supplemented with SCF, TPO, IL-3 and EPO. The resultant frequencies of mixed colonies (nmEM) did not differ significantly between Bmal1 +/+ and Bmal1 −/− CD34−KSL cells (Bmal1 +/+ CD34−KSL cells; 29.17 ± 1.18%, Bmal1 −/− CD34−KSL cells; 34.40 ± 2.22%) (Figure 1A). After close examination, we found that there is no significant morphological difference between the colonies of two groups (Figure 1B). In addition, Bmal1 +/+ and Bmal1 −/− CD34−KSL cells demonstrated comparable proliferation potentials after 7 days culture (Figure 1C).
Normal differentiation in vitro and normal long-term reconstitution ability in vivo of Bmal1 −/− HSCs. A, B) Normal in vitro colony formation capacity of Bmal1 −/− HSCs. Single HSCs from Bmal1 +/+ and Bmal1 −/− mice were cultured with cytokines for 11 days. Data shown are the mean numbers ± SDs of colonies of three independent experiments (n = 3). Colony cells were morphologically identified as neutrophils (n), macrophages (m), erythroblasts (E) and megakaryocytes (M). The scale bar in B is 100 μm. C) Comparable proliferation potentials of Bmal1 −/− HSCs. CD34−KSL HSCs were clonally deposited into 96-well micro-titer plates containing 200 μl of S-Clone SF-03 supplemented with 10% BSA and cultured with the indicated cytokines (50 ng/ml mouse SCF, 50 ng/ml TPO) for 7 days. Cell numbers were counted under a microscope. Data shown are mean numbers ± SEMs of colonies (n = 52). D-F) Comparable long-term reconstitution ability of Bmal1 +/+ and Bmal1 −/− HSCs during serial transplantation. Lethally irradiated recipient B6-Ly5.1 mice were transplanted with 1 × 106 BM cells (harvested at ZT5) from Bmal1 +/+ and Bmal1 −/− mice (Ly5.2) and the same number of BM competitor cells from F1 mice in a competitive repopulation assay. Data shown are the mean ratios ± SDs of donor-derived cells in the PB at 12 weeks after the first BMT (D, n = 7), in the BM at 12 weeks after the first transplantation (E, n = 7), and in the PB at 12 weeks after the second BMT (F, n = 5) of three independent experiments.
Bmal1 is dispensable for Bone marrow reconstitution
To determine the repopulating ability of Bmal1 −/− HSCs in vivo, we designed a competitive repopulation assay. For this purpose, 1 × 106 BM cells from Bmal1 +/+ or Bmal1 −/− mice were transplanted into lethally irradiated recipient mice along with an equal number of BM cells from B6-F1 mice. At 4, 8 and 12 weeks after transplantation, flow cytometric analysis showed a high-level chimerism of B220+ cells in PB of the recipients transplanted with Bmal1 −/− BM cells, but this was not observed in the second Bone Marrow Transplantation (BMT). In addition, there was no statistically significant difference in the chimerism of Gr-1+/Mac-1+ and CD3+ cells. These results suggest that Bmal1 +/+ and Bmal1 −/− BM cells are equally capable of hematopoietic reconstitution (Figure 1D and Additional file 1: Figure S1B). With regard to donor-derived chimerism in the recipient’s BM, there was no significant difference between Bmal1 +/+ and Bmal1 −/−-derived CD34−KSL cells (Figure 1E).
In a second competitive repopulation assay, at 12 weeks after the first BMT, 1 × 106 BM cells from these recipients were transplanted into second recipient mice. At 4, 8 and 12 weeks after the second BMT, no big difference was also seen between the hematopoietic reconstitution ability of both donor-derived cells (Figure 1F and Additional file 1: Figure S1C). Moreover, we performed a third BMT at 12 weeks after the second BMT, but the result was the same as with the second BMT (data not shown).
Normal frequencies and hibernation state of Bmal1 −/− HSCs
Although these results presented here led us to the conclusion that there appears to be no intrinsic circadian rhythm in HSCs, deficiency of Bmal1 might change BM niche and affects the frequencies or cell cycling of HSCs. However, flow cytometry analysis of BM revealed no significant difference in the frequencies of Bmal1 +/+ and Bmal1 −/− CD34−KSL cells at ZT5 and ZT17 (Figure 2A,B). Likewise, the frequencies of KSL cells, Common myeloid progenitor (CMP), Granulocyte-macrophage progenitor (GMP) and Megakaryocyte-erythroid progenitor (MEP) in Bmal1 −/− mice were similar to those in Bmal1 +/+ mice (Additional file 2: Figure S2).
Cell cycling and differentiation of HSCs are normal in arrhythmic Bmal1 deficient mice. A, B) Normal frequency of HSCs in the BM of 8-10-week-old Bmal1 −/− mice. CD34−KSL fractions were assessed by flow cytometry. A) Data shown are representative of CD34−KSL cells at ZT5 and ZT17. B) The mean percentages ± SDs of CD34−KSL cells at ZT5 (n = 4) and ZT17 (n = 3) of two independent experiments. C) Comparable frequency of quiescent cells in HSC populations. HSCs of Bmal1 +/+ and Bmal1 −/− mice were stained with Pyronin Y and analyzed by flow cytometry to give the mean percentages ± SDs of Pyronin Y− cells in the CD34−KSL populations at ZT5 and ZT17 (n = 3) of two independent experiments. D) Normal EdU incorporation in Bmal1 −/− CD34−KSL cells. EdU was administered orally to mice for 3 weeks, and EdU incorporation into HSCs was evaluated using a Click-iT EdU PB Flow Cytometry Assay Kit. Data shown are the mean percentages ± SDs of EdU+ cells in HSC populations (Bmal1 −/− mice; n = 6, Bmal1 −/− mice; n = 3). E) White blood cell differentiation in young (10-week-old) and aged (40-week-old) mice. Each stack in the bar represents a cell type percentage. Gr-1+, granulocytes; Mac-1+, macrophages; B220+, B cells; CD4+, CD4+ T cells; and CD8+, CD8+ T cells (n = 6) of four independent experiments.
To investigate the hibernation status of HSCs in Bmal1 −/− mice, we stained CD34−KSL cells with Pyronin Y [12]. Consistent with our previous work [4], we found that most Bmal1 +/+ and Bmal1 −/− CD34−KSL cells were negative for Pyronin Y staining, indicating normal HSC hibernation state, and that there were no differences depending on circadian rhythm (Figure 2C). In addition, after oral administration of EdU (5-ethynyl-2´-deoxyuridine) to Bmal1 −/− mice for 3 weeks, we could not obtain statistically significant difference in EdU incorporation between Bmal1 +/+ and Bmal1 −/− CD34−KSL, indicating no alteration in cell cycling status (Figure 2D).
Bmal1 deficiency does not affect white blood cell differentiation
It has been reported that life span of Bmal1 −/− mice is only half that of wild-type mice [10], raising the possibility of an altered hematopoietic differentiation program in Bmal1 −/− mice. We therefore examined PB cells of Bmal1 +/+ and Bmal1 −/− mice at 10 and 40 weeks of age. Although most Bmal1 −/− mice died within 40-week-old and the survived 40-week-old Bmal1 −/− mice looked older than their Bmal1 +/+ counterparts, there were no significant changes in the levels of myeloid cells, B cells or T cells (Figure 2E).
Concluding remarks
Recent studies have demonstrated that the central clock in suprachiasmatic nucleus (SCN) regulates the expression of Cxcl12 through sympathetic nervous system [7] and Cxcr4 expression in BM KSL cells or CD150+CD48− cells [13] fluctuates according to circadian rhythms [14]. However, it has been reported that the clock genes are not expressed rhythmically in side population (SP) cells [15], suggesting that Cxcr4 expression may be independent from control of clock genes. Moreover, Yagita et. al. [16] have recently found that circadian clock oscillation is not detected in mouse embryonic stem (ES) cells and induced pluripotent stem (iPS) cells, but is induced during their differentiation. Taken together, these findings appear to support the idea that the absence of circadian rhythm does not affect the function of stem cells in common.
In conclusion, despite the fact that mobilization of HSCs is controlled by circadian rhythm, our results demonstrate that Bmal1 deficiency does not affect differentiation, proliferation and repopulating ability of murine HSCs. Therefore, we propose that circadian gene Bmal1 is dispensable for intrinsic properties of murine HSCs.
Abbreviations
- HSC:
-
Hematopoietic stem cell
- BM:
-
Bone marrow
- PB:
-
Peripheral blood
- HSPCs:
-
Hematopoietic stem and progenitor cells
- BMT:
-
Bone marrow transplantation
- ZT:
-
Zeitgeber time
- CMP:
-
Common myeloid progenitor
- GMP:
-
Granulocyte-macrophage progenitor
- MEP:
-
Megakaryocyte-erythroid progenitor
- SCN:
-
Suprachiasmatic nucleus
- SP:
-
Side population
- ES:
-
Embryonic stem
- iPS:
-
Induced pluripotent stem.
References
Kiel MJ, Morrison SJ: Uncertainty in the niches that maintain haematopoietic stem cells. Nat Rev Immunol. 2008, 8 (4): 290-301. 10.1038/nri2279.
Orkin SH, Zon LI: Hematopoiesis: an evolving paradigm for stem cell biology. Cell. 2008, 132 (4): 631-644. 10.1016/j.cell.2008.01.025.
Yamazaki S, Ema H, Karlsson G, Yamaguchi T, Miyoshi H, Shioda S, Taketo MM, Karlsson S, Iwama A, Nakauchi H: Nonmyelinating Schwann cells maintain hematopoietic stem cell hibernation in the bone marrow niche. Cell. 2011, 147 (5): 1146-1158. 10.1016/j.cell.2011.09.053.
Yamazaki S, Iwama A, Takayanagi S, Morita Y, Eto K, Ema H, Nakauchi H: Cytokine signals modulated via lipid rafts mimic niche signals and induce hibernation in hematopoietic stem cells. EMBO J. 2006, 25 (15): 3515-3523. 10.1038/sj.emboj.7601236.
Bradford GB, Williams B, Rossi R, Bertoncello I: Quiescence, cycling, and turnover in the primitive hematopoietic stem cell compartment. Exp Hematol. 1997, 25 (5): 445-453.
Cheshier SH, Morrison SJ, Liao X, Weissman IL: In vivo proliferation and cell cycle kinetics of long-term self-renewing hematopoietic stem cells. Proc Natl Acad Sci U S A. 1999, 96 (6): 3120-3125. 10.1073/pnas.96.6.3120.
Mendez-Ferrer S, Lucas D, Battista M, Frenette PS: Haematopoietic stem cell release is regulated by circadian oscillations. Nature. 2008, 452 (7186): 442-447. 10.1038/nature06685.
King DP, Takahashi JS: Molecular genetics of circadian rhythms in mammals. Annu Rev Neurosci. 2000, 23: 713-742. 10.1146/annurev.neuro.23.1.713.
Bunger MK, Wilsbacher LD, Moran SM, Clendenin C, Radcliffe LA, Hogenesch JB, Simon MC, Takahashi JS, Bradfield CA: Mop3 is an essential component of the master circadian pacemaker in mammals. Cell. 2000, 103 (7): 1009-1017. 10.1016/S0092-8674(00)00205-1.
Kondratov RV, Kondratova AA, Gorbacheva VY, Vykhovanets OV, Antoch MP: Early aging and age-related pathologies in mice deficient in BMAL1, the core componentof the circadian clock. Genes Dev. 2006, 20 (14): 1868-1873. 10.1101/gad.1432206.
Shimba S, Ogawa T, Hitosugi S, Ichihashi Y, Nakadaira Y, Kobayashi M, Tezuka M, Kosuge Y, Ishige K, Ito Y, Komiyama K, Okamatsu -Ogura Y, Kimura K, Saito M: Deficient of a clock gene, brain and muscle Arnt-like protein-1 (BMAL1), induces dyslipidemia and ectopic fat formation. PLoS One. 2011, 6 (9): e25231-10.1371/journal.pone.0025231.
Huttmann A, Liu SL, Boyd AW, Li CL: Functional heterogeneity within rhodamine123(lo) Hoechst33342(lo/sp) primitive hemopoietic stem cells revealed by pyronin Y. Exp Hematol. 2001, 29 (9): 1109-1116. 10.1016/S0301-472X(01)00684-1.
Kiel MJ, Yilmaz OH, Iwashita T, Terhorst C, Morrison SJ: SLAM family receptors distinguish hematopoietic stem and progenitor cells and reveal endothelial niches for stem cells. Cell. 2005, 121 (7): 1109-1121. 10.1016/j.cell.2005.05.026.
Lucas D, Battista M, Shi PA, Isola L, Frenette PS: Mobilized hematopoietic stem cell yield depends on species-specific circadian timing. Cell Stem Cell. 2008, 3 (4): 364-366. 10.1016/j.stem.2008.09.004.
Tsinkalovsky O, Filipski E, Rosenlund B, Sothern RB, Eiken HG, Wu MW, Claustrat B, Bayer J, Levi F, Laerum OD: Circadian expression of clock genes in purified hematopoietic stem cells is developmentally regulated in mouse bone marrow. Exp Hematol. 2006, 34 (9): 1249-1261.
Yagita K, Horie K, Koinuma S, Nakamura W, Yamanaka I, Urasaki A, Shigeyoshi Y, Kawakami K, Shimada S, Takeda J, Uchiyama Y: Development of the circadian oscillator during differentiation of mouse embryonic stem cells in vitro. Proc Natl Acad Sci U S A. 2010, 107 (8): 3846-3851. 10.1073/pnas.0913256107.
Acknowledgements
We thank Dr. H Yoshitane, Dr. Y Fukada, Y Ishii and Y Yamazaki for technical help and advice, and Dr. M Kasai for critical reading of the manuscript. This work was supported in part by grants from the Ministry of Education, Culture, Sport, Science and Technology, Japan, Japan Science and Technology Corporation (JST).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
AI and SY designed the research and analyzed the data. AI, SY, SS and HN wrote the paper. All authors read and approved the final manuscript.
Electronic supplementary material
12952_2013_156_MOESM1_ESM.pdf
Additional file 1: Figure S1: A) Traffic of HSPCs to bloodstream shows circadian oscillation. Circulating Colony-forming Units in Culture (CFU-Cs) did not oscillate in Bmal1 −/− mice (n = 3) compared with Bmal1 +/+ mice (n = 4). Data shown are the mean percentages ± SDs of two independent experiments. B, C) Comparable long-term reconstitution ability of Bmal1 +/+ and Bmal1 −/− HSCs during serial transplantation. Data shown are the mean ratios ± SDs of donor-derived cells in the PB at 4, 8 weeks after the first (n = 7) and the second BMT (n = 5) of three independent experiments. (PDF 5 MB)
12952_2013_156_MOESM2_ESM.pdf
Additional file 2: Figure S2: Normal frequency of progenitors in the BM of 8-10-week-old Bmal1 −/− mice. KSL, CMP, GMP and MEP fractions were assessed by flow cytometry. The mean percentages ± SDs of KSL cells, CMP, GMP and MEP of Bmal1 +/+ and Bmal1 −/− mice of two independent experiments (n = 3). (PDF 5 MB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Ieyasu, A., Tajima, Y., Shimba, S. et al. Clock gene Bmal1 is dispensable for intrinsic properties of murine hematopoietic stem cells. J Negat Results BioMed 13, 4 (2014). https://doi.org/10.1186/1477-5751-13-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1477-5751-13-4