Abstract
Background
The bacterium Bacillus subtilis, which is not a natural riboflavin overproducer, has been converted into an excellent production strain by classical mutagenesis and metabolic engineering. To our knowledge, the enhancement of riboflavin excretion from the cytoplasm of overproducing cells has not yet been considered as a target for (further) strain improvement. Here we evaluate the flavin transporter RibM from Streptomyces davawensis with respect to improvement of a riboflavin production strain.
Results
The gene ribM from S. davawensis, coding for a putative facilitator of riboflavin uptake, was codon optimized (ribM opt ) for expression in B. subtilis. The gene ribM opt was functionally introduced into B. subtilis using the isopropyl-β-thiogalactopyranoside (IPTG)-inducible expression plasmid pHT01: Northern-blot analysis of total RNA from IPTG treated recombinant B. subtilis cells revealed a ribM opt specific transcript. Western blot analysis showed that the his6-tagged heterologous gene product RibM was present in the cytoplasmic membrane. Expression of ribM in Escherichia coli increased [14C]riboflavin uptake, which was not affected by the protonophore carbonyl cyanide m-chlorophenylhydrazone (CCCP). Expression of ribM opt supported growth of a B. subtilis ΔribB::Ermr ΔribU::Kanr double mutant deficient in riboflavin synthesis (ΔribB) and also deficient with respect to riboflavin uptake (ΔribU). Expression of ribM opt increased roseoflavin (a toxic riboflavin analog produced by S. davawensis) sensitivity of a B. subtilis ΔribU::Kanr strain. Riboflavin synthesis by a model riboflavin B. subtilis production strain overproducing RibM was increased significantly depending on the amount of the inducer IPTG.
Conclusions
The energy independent flavin facilitator RibM could in principle catalyze riboflavin export and thus may be useful to increase the riboflavin yield in a riboflavin production process using a recombinant RibM overproducing B. subtilis strain (or any other microorganism).
Similar content being viewed by others
Background
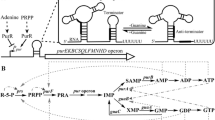
Riboflavin (vitamin B2) is a direct precursor to the cofactors flavin mononucleotide (FMN) and flavin adenine dinucleotide (FAD). Riboflavin is synthesized by plants and many microorganisms; it is not synthesized by animals [1]. Many Gram-positive bacteria seem to be capable of acquiring riboflavin from the environment, whereas most Gram-negative bacteria depend on the endogenous synthesis of this vitamin [2]. Riboflavin transporters (uptake systems) have been identified and characterized in B. subtilis [3], in Lactococcus lactis [4, 5] and in a few other bacteria. Three classes of riboflavin transporters seem to exist: (1) homologs of ribU of B. subtilis, (2) homologs of ribM of Corynebacterium glutamicum and (3) homologs of impX of Fusobacterium nucleatum [6]. The latter class has not been functionally characterized [2]. B. subtilis RibU is part of a modular multi-subunit riboflavin transporter and belongs to the recently identified family of energy-coupling factor (ECF) transporters [7–11]. L. lactis RibU [4] has also been included in the latter classification, the driving force behind transport activity was shown to be ATP hydrolysis [9]. Notably, RibU of Staphylococcus aureus has been crystallized and its three-dimensional structure has been determined [8]. B. subtilis RibU is a proton-riboflavin symporter with high affinity for its substrate (K m = 5-20 nM) [3]. RibU is strikingly different from the Corynebacterium glutamicum riboflavin transporter RibM, which was characterized as an energy-independent facilitator for riboflavin with much lower affinity (K m = 11 μM) [3]. RibM from C. glutamicum is similar (40% at the amino acid level) to RibM (23.7 kDa) from S. davawensis. The gene for the latter protein is present in the S. davawensis riboflavin biosynthetic gene cluster ribBMAH which is controlled by an FMN riboswitch [12] directly upstream of ribB [13]. S. davawensis is the only known producer of the riboflavin analog roseoflavin, which has antibiotic activity [14]. Highly similar (> 65% similarity) RibM proteins (all containing five putative trans membrane domains) are present in other species of the genus Streptomyces. The gene for the flavin facilitator ribM from S. davawensis was codon optimized for expression in B. subtilis. The gene ribM was functionally characterized and was found to encode a transporter for riboflavin and roseoflavin. Finally, ribM was evaluated as a possible tool to enhance the productivity of a B. subtilis riboflavin production strain.
Results
Expression of ribM from S. davawensis in E. coli increased riboflavin uptake
The wild-type gene ribM from S. davawensis was expressed in E. coli using the plasmid pNCO113ribM, employing an IPTG controlled T5 promotor [15]. E. coli does not contain an endogenous uptake system for flavins [16]. Riboflavin import was monitored using [14C]riboflavin. It was found that expression of ribM resulted in a significant increase in the cellular radioactivity shortly after adding the labelled substrate (Figure 1A) strongly suggesting that ribM codes for a functional flavin transporter. The uptake activity was 1.5 pmol riboflavin × OD cells-1 min-1 in the ribM expressing strain. The control strain (not containing ribM) showed an uptake activity of 0.04 pmol riboflavin × OD cells-1 min-1. Riboflavin uptake mediated by S. davawensis RibM was only slightly affected by FAD and the protonophore CCCP (Figure 1B). Roseoflavin, FMN and a 10-fold excess of unlabelled riboflavin apparently reduced [14C]riboflavin uptake suggesting that roseoflavin and FMN are substrates for RibM. Similar results were generated in our previous study analysing RibM from C. glutamicum [3], where it was additionally shown that riboflavin uptake was not affected by the absence of glucose or the addition of sodium azide. In summary, we conclude that RibM from S. davawensis is a transporter for riboflavin and roseoflavin, and that transport appears to occur independently of metabolic energy.
Characterization of RibM from Streptomyces davawensis as a riboflavin transporter. (A) The gene ribM from S. davawensis was overexpressed in E. coli BL21 using the plasmid pNCO113ribM (grey triangles). An Escherichia coli strain containing the empty vector pNCO113 (black circles) served as a control. E. coli strains were grown in riboflavin-free M9 medium. The cells were collected by centrifugation, suspended in transport buffer (50 mM K2HPO4/KH2PO4, 50 mM MgCl2, pH 7.0) and the experiment was started by adding [14C]riboflavin to a final concentration of 1.6 μM. Samples were filtered, washed with water and the radioactivity was determined by liquid scintillation counting. "1 OD cells" means "the amount of cells that when present in 1 ml will lead to an OD600 reading of 1.0". (B) Uptake experiments were performed as described for (A) with 2.2 μM [14C]riboflavin in the presence of a 10-fold excess of riboflavin (ribM+RF), flavin mononucleotide (ribM+FMN), flavin adenine dinucleotide (ribM+FAD), roseoflavin (ribM+RoF) or carbonyl cyanide m-chlorophenylhydrazone (ribM+CCCP; 130 μM). The uptake activity in the absence of competitors was 4.8 pmol pmol riboflavin × OD cells-1 min-1 (ribM). In the presence of CCCP uptake activity was 4.3 pmol riboflavin × OD cells-1 min-1. "1 OD cells" means "the amount of cells that when present in 1 ml will lead to an OD600 reading of 1.0".
Introduction of ribM from S. davawensis in B. subtilis
The ribM gene from S. davawensis has a relatively high G+C-content (72%) and was optimized with respect to the codon usage of B. subtilis in order to allow efficient heterologous expression. The artificial gene was named ribM opt and was inserted into the expression vector pHT01 to give pHT01ribMopt. Expression using pHT01 is based on the strong σA-dependent promoter (P grac ) preceding the groE operon of B. subtilis which has been converted into an efficiently controllable (IPTG-inducible) promoter by addition of the lac operator from E. coli. A B. subtilis wild-type strain (Marburg 168) and a riboflavin overproducing B. subtilis strain (BSHP) were transformed with pHT01ribMopt. From the resulting strains, B. subtilis 168 < pHT01ribMopt > and BSHP < pHT01ribMopt > plasmids were isolated. DNA sequencing revealed the presence of the ribM opt gene under control of P grac in all strains.
The gene ribM opt from S. davawensis was functionally expressed in B. subtilis
B. subtilis 168 < pHT01ribMopt > and B. subtilis 168 < pHT01 > (control) were grown to the exponential growth phase in LB and treated with IPTG (100 μM). Cells were harvested at different time points after induction (3 h, 6 h and 9 h) and total RNA was prepared. Northern blot analysis using a ribM opt specific oligonucleotide probe revealed, that the heterologous gene ribM opt was expressed in B. subtilis 168 < pHT01ribMopt > (Figure 2A). The 0.8 kb transcript was found in IPTG treated cells only. The strongest signal was observed 3 h after induction (Figure 2A, lane 4). In this lane an additional band at 1.8 kb appeared, possibly formed by aggregates of not fully denatured ribM opt transcripts. The data showed that the ribM opt transcript was present even 9 h after induction (in the stationary growth phase). Subsequently, cytoplasmic fractions and membrane fractions of cell-free extracts of B. subtilis 168 < pHT01ribMopt > and B. subtilis 168 < pHT01 > (control) were subjected to Western blot analysis. The synthesis of his6-tagged RibM was monitored using anti-penta-his antibodies (Figure 2B). His6-tagged RibM was found in the membrane fraction of IPTG (100 μM) treated B. subtilis 168 < pHT01ribMopt > only, indicating that RibM was directed to the membranes (Figure 2B, lane 4). The addition of higher amounts of IPTG (1 mM) did not enhance the amount of heterologous RibM in recombinant B. subtilis strains (data not shown). Thus (if not otherwise indicated), in the following experiments, stimulation of RibM synthesis in B. subtilis routinely was carried out using 100 μM IPTG.
Monitoring expression of the gene ribM from Streptomyces davawensis in recombinant Bacillus subtilis strains containing the plasmid pHT01ribM opt . (A) Cell samples (normalized to an OD600 of 1) were taken 3, 6 and 9 h after treatment with IPTG and total RNA was prepared. Control cells were not treated with the inducer. The level of ribM opt -transcript was assessed by Northern blot analysis using a ribM opt -specific digoxigenin labelled DNA probe. The predicted size of the ribM opt transcript is 0.8 kb (see arrow). Lanes 1, 6 and 10: B. subtilis 168 < pHT01 >; Lanes 2, 7 and 11: B. subtilis 168 < pHT01 > induced with 1 mM IPTG; Lanes 3, 8, and 12: B. subtilis 168 < pHT01ribMopt >; Lanes 4, 9 and 13: B. subtilis 168 < pHT01ribMopt > induced with 1 mM IPTG; Lane 5, no sample. (B) After growth in LB and treatment with IPTG B. subtilis < pHT01ribMopt > cells were collected and cell free extracts were prepared. Cytoplasmic fractions and membrane fractions were subjected to SDS-PAGE, followed by transfer to a nitrocellulose membrane and analysis with anti-penta-his mouse monoclonal antibodies/goat anti-mouse IgG alkaline phosphatase (AP)-coupled secondary antibodies. Lane M, protein marker (in kDa); lane 1, B. subtilis < pHT01 > cytoplasmic fraction; lane 2, B. subtilis 168 < pHT01 > membrane fraction; lane 3, B. subtilis 168 < pHT01ribMopt > cytoplasmic fraction; lane 4, B. subtilis 168 < pHT01ribMopt > membrane fraction; (C-terminally his6-tagged RibM from Streptomyces davawensis, calculated molecular mass: 24.7 kDa).
Synthesis of RibM in B. subtilis mutants supported growth and enhanced roseoflavin sensitivity
In the following experiment the B. subtilis double mutant ΔribB::Ermr (ribB was deleted and instead an erythromycin resistance gene was introduced) ΔribU::Kanr (ribU was deleted and replaced by a kanamycin resistance gene) was used. This strain carries deletions in ribB, encoding riboflavin synthase (EC 2.5.1.9) catalyzing the last step of riboflavin biosynthesis [17], and in ribU, the energy-dependent riboflavin transporter [3]. The double mutant was transformed with pHT01ribMopt or the empty plasmid and both transformants were compared in growth assays on LB plates. Only the strains transformed with pHT01ribMopt could grow on LB, which contains about 0.5 μM riboflavin, or on LB supplemented with (additional) 0.1 μM riboflavin or 1 μM riboflavin (Figure 3). Upon addition of 10 μM riboflavin both transformants could grow. Growth of the control strain, however, was reduced. At 100 μM riboflavin no difference in growth was observed indicating that at this concentration the vitamin crosses the cytoplasmic membrane independently of a dedicated transport system. Similar experiments were performed in liquid Spizizen minimal media containing glucose or sucrose (both at 10 g/L) and 1 μM riboflavin in the presence or absence of the inducer IPTG. In all experiments, only the pHT01ribMopt containing strains were able to grow (data not shown). We conclude from these experiments that RibM was responsible for flavin transport, giving further evidence that RibM from S. davawensis is a functional transporter in B. subtilis.
The overproduction of RibM allowed growth of a Δ ribU ::Kanr Δ ribB ::Ermr Bacillus subtilis strain. Streaks (top) and drops (bottom, about 50,000 cells) of B. subtilis ΔribU::Kanr ΔribB::Ermr cells expressing ribM from plasmid pHT01ribMopt were applied to LB plates (about 0.5 μM riboflavin) containing the indicated (additional) amount of riboflavin and 100 μM IPTG. Growth was recorded after incubation for 36 h at 37°C. As controls, strains were transformed with the empty vector pHT01. Apparently, only the strains transformed with pHT01ribMopt could grow on culture media (LB) with low amounts of riboflavin.
The mutant strain ΔribU::Kanr (deficient in flavin uptake) was transformed with pHT01ribMopt and tested with respect to roseoflavin sensitivity on LB plates containing 100 μM IPTG (Figure 4A). Roseoflavin is a toxic riboflavin analog. Clearly, the presence of ribM enhanced roseoflavin transport and thus roseoflavin sensitivity of ΔribU::Kanr < pHT01ribMopt > at 10 μM roseoflavin. The addition of > 50 μM roseoflavin reduced growth of the ΔribU::Kanr control strain containing empty pHT01, indicating that the flavin (like riboflavin) is able to permeate over the cytoplasmic membrane without catalysis mediated by a facilitator. Similar experiments were carried out using liquid LB cultures. B. subtilis ΔribU::Kanr containing pHT01ribMopt was grown in the presence of the inducer IPTG and different concentrations of roseoflavin (Figure 4B). In LB broth, the effect of ribM opt was most obvious at 50 μM roseoflavin where the ribM opt containing strain grew to OD600 of about 2.5 and the strain containing the empty vector grew to OD600 of 4.2. A higher concentration of roseoflavin (100 μM) inhibited growth of both strains, pointing towards a transporter independent diffusion of roseoflavin (as observed for riboflavin).
Overproduction of RibM enhanced roseoflavin sensitivity of a Δ ribU ::Kanr B. subtilis strain. (A) Streaks (top) and drops (bottom, about 50,000 cells) of B. subtilis ΔribU::KanR cells expressing ribM from plasmid pHT01ribMopt were applied to LB plates containing the indicated amounts of the toxic riboflavin analog roseoflavin and 100 μM IPTG. Growth was recorded after incubation for 36 h at 37°C. As controls, strains were transformed with the empty vector pHT01. At 10 μM roseoflavin it was most obvious that the presence of RibM increased roseoflavin sensitivity due to import of the toxic compound. (B) B. subtilis ΔribU::KanR expressing ribM from plasmid pHT01ribMopt was grown in LB broth in the presence of the indicated amounts of roseoflavin and 100 μM IPTG. Growth was recorded at μ = 600 nm. As controls, ΔribU::KanR strains containing empty pHT01 were analyzed. At 50 μM roseoflavin, the effect of RibM was most obvious (see bracket). Cells (black triangles) producing RibM imported toxic roseoflavin and consequently grew to an OD600 of about 2.5 only. The control strain (open triangles) was less affected by roseoflavin and grew to an OD600 of 4.2.
RibM enhanced riboflavin production in a high performance B. subtilis production strain
The high-performance riboflavin production strain BSHP, which produces about 350 mg/L riboflavin, was transformed with pHT01ribMopt and pHT01. Two transformant BSHP strains containing pHT01ribMopt ("ribM1" and "ribM2") and four transformant strains containing pHT01 (controls) were isolated and grown in shake flasks in a medium routinely used for the analysis of the performance of industrial riboflavin production strains. For each strain three different flasks were inoculated and analyzed (n = 3). The cells were treated with IPTG in order to control synthesis of RibM. The final cell densities of all cultures were very similar (about OD600 of 4). At the end of fermentation residual sugar was determined. In all cultures the carbon source was completely metabolized. The total amount of riboflavin (mg/L) (intracellular riboflavin and riboflavin present in the fermentation broth) in the cultures was measured (Figure 5). At 0 μM IPTG the riboflavin concentration was similar in cultures of the pHT01ribMopt containing strains ribM1 and ribM2 when compared to the control pHT01 (the results of the three other control strains were very similar and thus the data were not included in the figure). Similar results were obtained at 10 μM IPTG. At 100 μM IPTG and at 1 mM IPTG, however, a significant increase at the 5% level of significance was found. The data revealed that the differences in productivity were 11% (100 μM IPTG) and 18% (1 mM IPTG). The latter values were determined by comparing the average amount of riboflavin synthesized by ribM1 and ribM2 to the amount produced by the control. Surprisingly, at 1 mM IPTG the amount of riboflavin in cultures of the control strain containing the empty plasmid pHT01 was significantly lower as compared to the values obtained at 0 μM IPTG, 10 μM IPTG and 100 μM IPTG. We do not have an explanation for this. As mentioned above, the control experiment was done with four different independent transformant strains with similar results. It is known that reduced growth positively affects the accumulation of metabolites. Growth, however, was very similar in all cultures (the carbon source was completely metabolized) and therefore a growth limitation cannot explain the apparent increase of the riboflavin synthesis. All in all our data suggest that the presence of RibM enhanced riboflavin production under these experimental conditions.
Introduction of the heterologous flavin facilitator RibM from Streptomyces davawensis enhanced riboflavin synthesis in the high-performance riboflavin production strain BSHP. Two BSHP strains containing pHT01ribMopt (designated as ribM1 and ribM2) and one strain containing pHT01 (control) were grown in shake flasks in the medium used for industrial riboflavin production. For each strain three shake flask experiments were carried out (n = 3). Different amounts of IPTG were used to induce synthesis of RibM. At the end of fermentation (30 h) residual sugar and the optical density at μ = 600 nm were measured. In all shake flasks the carbon source was completely metabolized and the strains had grown to a very similar cell density. For the cultures induced with 100 μM and 1 mM IPTG the t test gave the result that the RibM containing strains produced more riboflavin (mg/L), which was significant at the 5% level.
Discussion
Our results suggest that RibM from S. davawensis mediates flavin (riboflavin/roseoflavin) translocation via an energy independent facilitated diffusion mechanism. The probable physiological role of RibM is the acquirement of riboflavin from the environment. The genes ribBMAH from S. davawensis form a transcription unit and are expressed only when riboflavin is limiting in the growth medium [13]. The genes ribB (riboflavin synthase, alpha-chain; EC 2.5.1.9), ribA (bifunctional GTP cyclohydrolase II/3,4-dihydroxy-2-butanone-4-phosphate synthase; EC 3.5.4.25) and ribH (lumazine synthase; EC 2.5.1.9) are responsible for riboflavin biosynthesis. The fact that ribM is cotranscribed with the genes ribB, ribA and ribH suggests that a transporter is produced in parallel to the riboflavin biosynthetic enzymes. This appears not to be economical. If no riboflavin is present in the cytoplasm, the corresponding biosynthetic enzymes must be synthesized. The coproduction of a riboflavin transporter, however, is reasonable only when external riboflavin indeed is present. In turn, if riboflavin is present in the growth medium, the production of biosynthetic enzymes apparently constitutes a waste of metabolic energy. Notably, in the Streptomycetes S. avermitilis, S. coelicolor, S. scabiei and S. griseus ribBMAH clusters (containing ribM transporter genes) are present as well. It seems that Streptomycetes play it safe with respect to riboflavin supply. If riboflavin is growth limiting, proteins for transport and biosynthesis are synthesized in parallel to ensure availability of the vitamin.
The transport activity and direction of an energy independent facilitator depends on the concentration of the metabolite in the cytoplasm and the surrounding medium, respectively. Thus, RibM in principle may also confer roseoflavin resistance to S. davawensis (by exporting roseoflavin from the cytoplasm), which naturally produces this antibiotic in the stationary growth phase. The export activity of RibM (or other riboflavin transporters) in principle may also be of importance with respect to metabolic engineering of riboflavin production strains. In order to test this idea, ribM was functionally expressed in a production strain of B. subtilis, an organism, which is used for the commercial production of riboflavin [18]. The introduction of ribM into a high performance B. subtilis riboflavin production strain apparently improved riboflavin production. Although the increase was relatively small, for a large scale production process the improvement may be relevant. We suggest that the export activity of RibM leads to reduced levels of riboflavin in the cytoplasm possibly enhancing the carbon flux through the pathway. This may enhance productivity during the exponential growth phase (when riboflavin concentration outside is low) or during stationary growth (when riboflavin levels inside the cells are extremely high). Wild-type B. subtilis cells rapidly and almost quantitatively convert intracellular riboflavin to FMN and FAD [19], a reaction, which is catalyzed by the bifunctional flavokinase/FAD synthetase RibC. All production strains have a strongly reduced RibC activity (1%) [20, 21]. Thus, production strains contain unusually high amounts of free riboflavin, which, in principle, might reduce the activity of the riboflavin biosynthetic enzymes.
The introduction of a protein catalyzing riboflavin export may also improve current production processes employing other microorganisms such as Ashbya gossypii or Candida famata [22]. For A. gossypii interesting physiological studies suggest riboflavin export across the plasma membrane [23]. To our knowledge the corresponding exporter has not yet been identified.
Successful examples for strain optimization on the basis of metabolite export (L-threonine and L-lysine) have been reported for Corynebacterium glutamicum [24–26]. Our work is the first to use a similar approach for optimizing one of the most successful biotechnological processes, the commercial synthesis of riboflavin [2].
Conclusions
The gene ribM from S. davawensis encodes a membrane protein which is able to catalyze the uptake of riboflavin but also of the antibiotic roseoflavin, a structural riboflavin analog. The functional expression of ribM in E. coli and B. subtilis apparently was possible without the cosynthesis of other protein components. Furthermore, our previous [3] and our present data data suggest that RibM proteins are energy independent flavin facilitators. Consequently, S. davawensis RibM could in principle also catalyze riboflavin export and be useful to increase the riboflavin yield in a riboflavin production process. Most riboflavin currently is produced using genetically engineered microorganisms, whereby B. subtilis is an important host. Classic random mutagenesis and methods of metabolic engineering have been used in order to optimize B. subtilis for the production of riboflavin. Most effort was directed at impairment of regulation of the riboflavin biosynthetic operon and amplification of the copy number of the structural genes ribGBAH(T) [2]. However, import of substrates and export of the product have not yet been considered as strategies for further improvement. In order to partially fill this gap, we introduced ribM into a high-performance B. subtilis riboflavin production strain. We could show that the gene was actively transcribed and that the gene product RibM was directed to the cytoplasmic membrane. Shake flask experiments routinely used to evaluate the performance of riboflavin overproducing strains suggest that upon induction with 100 μM IPTG, the amount of riboflavin at the end of growth indeed is higher as compared to the controls. Possibly, riboflavin transporters from other microorganisms may even be more useful and may show even better results.
Methods
Bacterial strains, plasmids and growth conditions
E. coli DH5α was used as a host for gene cloning experiments and was aerobically grown at 37°C on lysogeny broth (LB) [27, 28]. E. coli BL21 [29] was used as a host for the uptake experiments with [14C]riboflavin. A recombinant E. coli BL21 strain overexpressing ribM from S. davawensis was generated by transformation using the plasmid pNCO113ribM which was described earlier [13]. The plasmid pET21 was obtained from Stratagene (Waldbronn, Germany). If not otherwise indicated B. subtilis was aerobically grown at 37°C in 2 × Spizizen's minimal medium [30] supplemented with 0.02% casamino acids, 2% yeast extract and 10% glucose or in LB. B. subtilis 168 (trpC2) [31] is a wild-type strain with respect to riboflavin biosynthesis and uptake and was used as a control. The B. subtilis double mutant ΔribB::Ermr ΔribU::Kanr is auxotrophic for riboflavin and does not contain a functional riboflavin uptake system (ribU) [3]. It was grown in media supplemented with 20 mg/L riboflavin, 1 μg/ml erythromycin and 5 μg/ml kanamycin. B. subtilis 168 ΔribU::Kanr [3] was grown in media supplemented with 5 μg/ml kanamycin. The B. subtilis high-performance riboflavin production strain BSHP was constructed by introducing additional copies of the B. subtilis ribGBAHT genes controlled by strong constitutive phage promoters (P spo ) into the genome of B. subtilis strain 3979 [18, 32].
Expression of S. davawensis ribM in B. subtilis
B. subtilis strains overproducing RibM (B. subtilis < pHT01ribMopt >) were generated using the expression vector pHT01 (Mobitech, Göttingen, Germany) replicating in Bacillus species from the pUB110 origin [33]. The gene ribM from S. davawensis was codon-optimized by GENEART (Regensburg, Germany) using overlapping oligonucleotides and PCR. In addition, codons specifying a his6-tag were introduced at the 3'-end. The gene (ribM opt ) (# FR719838, European Nucleotide Archive) was delivered in pGA4ribMopt. The latter plasmid was used as a template for PCR amplification of ribM opt using the modifying oligonucleotides RibMopt fw BamHI (5'-ACA GGA TCC ATG AAT TGG CTG AAT AGC-3') and RibMopt rv AatII (5'-ATT GAC GTC CTA TTA GTG GTG GTG ATG GTG-3'). The PCR product was purified and digested with BamHI and AatII to allow cloning in pHT01. The resulting plasmid pHT01ribMopt was used to transform B. subtilis using a standard protocol [34]. Expression of ribM opt in B. subtilis was stimulated by adding IPTG in the early exponential growth phase. For selection of pHT01ribMopt 30 μg ml-1 chloramphenicol was added to the growth media. Growth was monitored using a photometer at μ = 600 nm.
Uptake experiments in E. coli
E. coli BL21 cells were grown in M9 minimal medium [27] supplemented with antibiotics as required. The growth medium was inoculated to an OD600 of 0.15, and the cells were grown at 37°C until they reached an OD600 of 0.5. E. coli cells were treated with 0.5 mM IPTG and grown for 3 h (to an OD600 of 0.8). Cells were harvested, washed once with ice-cold water and once with transport buffer (50 mM KH2PO4/K2HPO4, 50 mM MgCl2 pH 7.0). The cells were finally resuspended in transport buffer (10 OD600 ml-1) and stored on ice. Uptake experiments were performed with 500 μl of cells that were vigorously stirred at 30°C. After warming for 2 min, glucose was added to a final concentration of 1 mM, and the assay was started by adding [14C]riboflavin (specific activity, 5.54 MBq/mg; a generous gift of R. Krämer, Köln, Germany) to a final concentration of 2 μM. Several aliquots were removed at minute intervals, filtered on 0.45-μm GN-6 membrane filters (Pall, Dreieich, Germany), washed with an excess of water, and analysed by liquid scintillation counting. The transport activity was determined without further additions or after adding carbonyl cyanide m-chlorophenylhydrazone (CCCP; 130 μM), unlabeled riboflavin, FMN, FAD or roseoflavin (MP Biomedicals, Montreal, Canada) 3 min before addition of the labeled substrate. The latter assay was done only once.
RNA preparation, Northern (RNA) blotting and hybridisation
The preparation of total RNA from B. subtilis, the transfer of RNA to a solid support and the hybridisation to a ribM opt specific digoxigenin-labelled DNA probe was carried out according to standard procedures [27]. The detection of the digoxigenin labelled probe was performed as suggested by the supplier of the "DIG DNA Labeling and Detection Kit" (Roche Diagnostics, Mannheim, Germany).
Preparation of B. subtilis membranes and Western blot analysis
Stationary phase cultures (50 ml) were harvested by centrifugation and washed twice with cold 50 mM Tris-HCl (pH 8.0). Pellets were dissolved in 1 ml TMS buffer (50 mM Tris-HCl, pH 8.0; 16 mM MgCl2; 33% sucrose (w/v); 300 μg/ml lysozyme; 1 mM phenylmethanesulfonylfluoride, PMSF) and incubated for 60 min at 37°C. Protoplasts were harvested by centrifugation (10 min at 7,500 × g at 4°C). Pellets were suspended in 1 ml lysis buffer (50 mM Tris-HCl, pH 8.0; 5 mM MgSO4; 2 mM PMSF). Cell free extracts were produced by sonication (1 min at 60% of the maximal power and 50% interval) on ice. Membranes were collected by ultracentrifugation for 30 min at 100,000 × g and washed once with 50 mM Tris-HCl (pH 8.0). Pellets were dissolved in 50 mM Tris-HCl (pH 8.0). The protein concentration was estimated by the method of Bradford [35]. The samples were analyzed by SDS-PAGE on 4-20% gradient polyacrylamide gels using 50 μg of protein per lane. After transfer to nitrocellulose membranes his6-tagged RibM was immunologically detected (mouse anti-penta-his primary antibodies/goat anti-mouse IgG alkaline phosphatase(AP)-coupled secondary antibodies; Novagen, Darmstadt, Germany). AP was detected with the "AP Detection Reagent Kit" (Novagen) using 3-bromo-4-chloro-5-indolyl phosphate and nitro blue tetrazolium chloride.
Monitoring of riboflavin production
Riboflavin synthesis by B. subtilis in the time course of fermentation was monitored as follows. An aliquot from the culture containing cells and medium (500 μl) was combined with 465 μl 4 N NaOH and vigorously mixed for 1 min. The mixture was neutralized by adding potassium phosphate (1 M pH 8.0) and centrifuged for 5 min at 13,000 × g at room temperature. Riboflavin in the supernatant was determined by using a standard procedure employing HPLC [21]. Using the above described protocol the total amount of riboflavin (intracellular riboflavin and riboflavin present in the fermentation broth) was measured. For the statistical analysis of the data student's t test was applied comparing two unknown means based on independent samples.
References
Fischer M, Bacher A: Biosynthesis of flavocoenzymes. Nat Prod Rep. 2005, 22 (3): 324-350. 10.1039/b210142b.
Abbas CA, Sibirny AA: Genetic control of biosynthesis and transport of riboflavin and flavin nucleotides and construction of robust biotechnological producers. Microbiol Mol Biol Rev. 2011, 75 (2): 321-360. 10.1128/MMBR.00030-10.
Vogl C, Grill S, Schilling O, Stulke J, Mack M, Stolz J: Characterization of riboflavin (vitamin B2) transport proteins from Bacillus subtilis and Corynebacterium glutamicum. J Bacteriol. 2007, 189 (20): 7367-7375. 10.1128/JB.00590-07.
Burgess CM, Slotboom DJ, Geertsma ER, Duurkens RH, Poolman B, van Sinderen D: The riboflavin transporter RibU in Lactococcus lactis: molecular characterization of gene expression and the transport mechanism. J Bacteriol. 2006, 188 (8): 2752-2760. 10.1128/JB.188.8.2752-2760.2006.
Duurkens RH, Tol MB, Geertsma ER, Permentier HP, Slotboom DJ: Flavin binding to the high affinity riboflavin transporter RibU. J Biol Chem. 2007, 282 (14): 10380-10386. 10.1074/jbc.M608583200.
Vitreschak AG, Rodionov DA, Mironov AA, Gelfand MS: Regulation of riboflavin biosynthesis and transport genes in bacteria by transcriptional and translational attenuation. Nucleic Acids Res. 2002, 30 (14): 3141-3151. 10.1093/nar/gkf433.
Rodionov DA, Hebbeln P, Eudes A, ter Beek J, Rodionova IA, Erkens GB, Slotboom DJ, Gelfand MS, Osterman AL, Hanson AD, et al: A novel class of modular transporters for vitamins in prokaryotes. J Bacteriol. 2009, 191 (1): 42-51. 10.1128/JB.01208-08.
Zhang P, Wang J, Shi Y: Structure and mechanism of the S component of a bacterial ECF transporter. Nature. 2010, 468 (7324): 717-720. 10.1038/nature09488.
ter Beek J, Duurkens RH, Erkens GB, Slotboom DJ: Quaternary structure and functional unit of energy coupling factor (ECF)-type transporters. J Biol Chem. 2011, 286 (7): 5471-5475. 10.1074/jbc.M110.199224.
Neubauer O, Alfandega A, Schoknecht J, Sternberg U, Pohlmann A, Eitinger T: Two essential arginine residues in the T components of energy-coupling factor transporters. J Bacteriol. 2009, 191 (21): 6482-6488. 10.1128/JB.00965-09.
Eitinger T, Rodionov DA, Grote M, Schneider E: Canonical and ECF-type ATP-binding cassette importers in prokaryotes: diversity in modular organization and cellular functions. FEMS Microbiol Rev. 2011, 35 (1): 3-67. 10.1111/j.1574-6976.2010.00230.x.
Lee ER, Blount KF, Breaker RR: Roseoflavin is a natural antibacterial compound that binds to FMN riboswitches and regulates gene expression. RNA Biol. 2009, 6 (2):
Grill S, Yamaguchi H, Wagner H, Zwahlen L, Kusch U, Mack M: Identification and characterization of two Streptomyces davawensis riboflavin biosynthesis gene clusters. Arch Microbiol. 2007, 188 (4): 377-387. 10.1007/s00203-007-0258-1.
Mack M, Grill S: Riboflavin analogs and inhibitors of riboflavin biosynthesis. Appl Microbiol Biotechnol. 2006, 71 (3): 265-275. 10.1007/s00253-006-0421-7.
Stüber D, Matile H, Garotta G: System for high-level production in Escherichia coli and rapid purification of recombinant proteins: application to epitope mapping, preparation of antibodies, and structure-function analysis. Immunological Methods. 1990, 4: 121-152.
Bandrin SV, Beburov M, Rabinovich PM, Stepanov AI: [Riboflavin auxotrophs of Escherichia coli]. Genetika. 1979, 15 (11): 2063-2065.
Perkins J, Pero J: Biosynthesis of riboflavin, biotin, folic acid, and cobalamin. Bacillus subtilis and Its Closest Relatives: from Genes to Cells. Edited by: Sonenshein A, Hoch J, Losick R. 2002, Washington DC: ASM Press, 271-286.
Perkins J, Sloma A, Hermann T, Theriault K, Zachgo E, Erdenberger T, Hannett N, Chatterjee NP, Williams V, Rufo GA, Hatch R, Pero J: Genetic engineering of Bacillus subtilis for the commercial production of riboflavin. Journal of Industrial Microbiology and Biotechnology. 1999, 22 (1): 8-18. 10.1038/sj.jim.2900587.
Demain AL: Riboflavin oversynthesis. Annu Rev Microbiol. 1972, 26: 369-388. 10.1146/annurev.mi.26.100172.002101.
Coquard D, Huecas M, Ott M, van Dijl JM, van Loon AP, Hohmann HP: Molecular cloning and characterisation of the ribC gene from Bacillus subtilis: a point mutation in ribC results in riboflavin overproduction. Mol Gen Genet. 1997, 254 (1): 81-84. 10.1007/s004380050393.
Mack M, van Loon AP, Hohmann HP: Regulation of riboflavin biosynthesis in Bacillus subtilis is affected by the activity of the flavokinase/flavin adenine dinucleotide synthetase encoded by ribC. J Bacteriol. 1998, 180 (4): 950-955.
Stahmann KP, Revuelta JL, Seulberger H: Three biotechnical processes using Ashbya gossypii, Candida famata, or Bacillus subtilis compete with chemical riboflavin production. Appl Microbiol Biotechnol. 2000, 53 (5): 509-516. 10.1007/s002530051649.
Forster C, Revuelta JL, Kramer R: Carrier-mediated transport of riboflavin in Ashbya gossypii. Appl Microbiol Biotechnol. 2001, 55 (1): 85-89. 10.1007/s002530000483.
Diesveld R, Tietze N, Furst O, Reth A, Bathe B, Sahm H, Eggeling L: Activity of exporters of Escherichia coli in Corynebacterium glutamicum, and their use to increase L-threonine production. J Mol Microbiol Biotechnol. 2009, 16 (3-4): 198-207. 10.1159/000142530.
Vrljic M, Garg J, Bellmann A, Wachi S, Freudl R, Malecki MJ, Sahm H, Kozina VJ, Eggeling L, Saier MH: The LysE superfamily: topology of the lysine exporter LysE of Corynebacterium glutamicum, a paradyme for a novel superfamily of transmembrane solute translocators. J Mol Microbiol Biotechnol. 1999, 1 (2): 327-336.
Vrljic M, Sahm H, Eggeling L: A new type of transporter with a new type of cellular function: L-lysine export from Corynebacterium glutamicum. Mol Microbiol. 1996, 22 (5): 815-826. 10.1046/j.1365-2958.1996.01527.x.
Sambrook J, Fritsch E, Maniatis T: Molecular cloning: a laboratory manual. 1989, Cold Spring Harbor, NY: Cold Spring Harbor Laboratory, 2
Bertani G: Lysogeny at mid-twentieth century: P1, P2, and other experimental systems. J Bacteriol. 2004, 186 (3): 595-600. 10.1128/JB.186.3.595-600.2004.
Studier FW, Moffatt BA: Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J Mol Biol. 1986, 189 (1): 113-130. 10.1016/0022-2836(86)90385-2.
Spizizen J: Transformation of Biochemically Deficient Strains of Bacillus Subtilis by Deoxyribonucleate. Proc Natl Acad Sci USA. 1958, 44 (10): 1072-1078. 10.1073/pnas.44.10.1072.
Saito H, Shibata T, Ando T: Mapping of genes determining nonpermissiveness and host-specific restriction to bacteriophages in Bacillus subtilis Marburg. Mol Gen Genet. 1979, 170 (2): 117-122. 10.1007/BF00337785.
Hümbelin M, Griesser V, Keller T, Schurter W, Haiker M, Hohmann HP, Ritz H, Richter G, Bacher A, van Loon APGM: GTP cyclohydrolase II and 3,4-dihydroxy-2-butanone 4-phosphate synthase are rate-limiting enzymes in riboflavin synthesis of an industrial Bacillus subtilis strain used for riboflavin production. J Ind Microbiol Biotechnol. 1999, 22: 1-7. 10.1038/sj.jim.2900590.
Maciag IE, Viret JF, Alonso JC: Replication and incompatibility properties of plasmid pUB110 in Bacillus subtilis. Mol Gen Genet. 1988, 212 (2): 232-240. 10.1007/BF00334690.
Chang S, Cohen SN: High frequency transformation of Bacillus subtilis protoplasts by plasmid DNA. Mol Gen Genet. 1979, 168 (1): 111-115. 10.1007/BF00267940.
Bradford MM: A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976, 72: 248-254. 10.1016/0003-2697(76)90527-3.
Acknowledgements
This work was supported by the state of Baden-Württemberg (Germany) ("Ideenwettbewerb"). We thank Dr. E. Huebener for critically reading the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
SH was involved in plasmid and strain construction, performed the Northern blot experiments and generated the membrane preparation/Western blot data. DP performed the growth experiments in the presence of flavin analogs. JS and CV did the riboflavin uptake experiments in Escherichia coli. ML carried out the experiments with the riboflavin production strain. MM coordinated the experiments and drafted the manuscript. All authors have read and approved the final version of the manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
Open Access This article is published under license to BioMed Central Ltd. This is an Open Access article is distributed under the terms of the Creative Commons Attribution License ( https://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Hemberger, S., Pedrolli, D.B., Stolz, J. et al. RibM from Streptomyces davawensis is a riboflavin/roseoflavin transporter and may be useful for the optimization of riboflavin production strains. BMC Biotechnol 11, 119 (2011). https://doi.org/10.1186/1472-6750-11-119
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1472-6750-11-119