Abstract
Background
Heme is a preferred iron source of bacterial pathogens. Streptococcus equi subspecies equi is a bacterial pathogen that causes strangles in horses. Whether S. equi has a heme acquisition transporter is unknown.
Results
An S. equi genome database was blasted with the heme binding proteins Shp and HtsA of Streptococcus pyogenes, and found that S. equi has the homologue of Shp (designated SeShp) and HtsA (designated SeHtsA). Tag-free recombinant SeShp and SeHtsA and 6xHis-tagged SeHtsA (SeHtsAHis) were prepared and characterized. Purified holoSeShp and holoSeHtsA bind Fe(II)-protoporphyrin IX (heme) and Fe(III)-protoporphyrin IX (hemin) in a 1:1 stoichiometry, respectively, and are designated hemoSeShp and hemiSeHtsA. HemiSeShp and hemiSeHtsAHis can be reconstituted from apoSeShp and apoSeHtsAHis and hemin. HemoSeShp is stable in air and can be oxidized to hemiSeShp by ferricyanide. HemiSeHtsA can be reduced into hemoSeHtsA, which autoxidizes readily. HemoSeShp rapidly transfers its heme to apoSeHtsAHis. In addition, hemoSeShp can also transfer its heme to apoHtsA, and hemoShp is able to donate heme to apoSeHtsAHis.
Conclusion
The primary structures, optical properties, oxidative stability, and in vitro heme transfer reaction of SeShp and SeHtsA are very similar to those of S. pyogenes Shp and HtsA. The data suggest that the putative cell surface protein SeShp and lipoprotein SeHtsA are part of the machinery to acquire heme in S. equi. The results also imply that the structure, function, and functional mechanism of the heme acquisition machinery are conserved in S. equi and S. pyogenes.
Similar content being viewed by others
Background
Iron is an essential nutrient that facilitates a wide variety of vital biological functions in almost all organisms. The ability of almost all pathogenic bacteria to acquire iron from the host environment is essential to establish infection [1]. Under the physiological conditions the predominant species of free iron, Fe3+, has a very low solubility and is of insufficient quantity to sustain bacterial growth. During infection, the mammalian host further deprives invading microorganisms of available iron through a process known as the hypoferraemic response [2]. Consequently, pathogenic bacteria must acquire iron from alternate sources and/or overcome the low solubility of Fe3+ by producing siderophores [3], whose complexes with Fe3+ is taken up by specific transporters.
Heme bound to hemoproteins constitutes the major iron reservoir in mammalian hosts and is the preferred iron source of bacterial pathogens [4]. Heme may be an even more important iron source for some streptococci. Unlike many other pathogenic bacteria, no streptococci have been reported to produce siderophores [5], though it was recently shown that the ABC transporter FtsABCD of Streptococcus pyogenes takes up ferric ferrichrome [6]. Free heme and hemoproteins, but not Fe-loaded transferrin nor lactoferrin, can be used as the iron source of S. pyogenes [7].
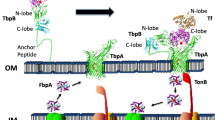
Different mechanisms for heme acquisition have been demonstrated in bacteria. Gram-negative bacteria Serratia marcescens, Pseudomonas aeruginosa, Pseudomonas fluorescens, Haemophilus influenza, and Yersinia pestis secrete proteins called hemophores to extract heme from host hemoproteins and deliver it to the receptor in the outer membrane [8]. The caught heme is then transported across the outer membrane into the periplasm in a TonB-dependent process [9] and finally brought across the cytoplasmic membrane by heme-specific ATP-binding cassette (ABC)-containing transporter. Some Gram-negative bacteria also acquire heme in a hemophore-independent mechanism. Specific outer membrane receptor directly interacts with and extracts heme from hemoproteins. Hemophore has not been found in Gram-positive bacteria. In addition to heme specific ABC transporters [10–12], cell surface proteins binding hemoproteins and heme have been identified in Gram-positive pathogens [13, 14]. It is believed that these surface proteins scavenge and relay heme from host hemoproteins to ABC transporter [15, 16].
The machinery involved in heme acquisition in Gram-positive bacteria is not conserved. S. pyogenes produces a cell surface heme-binding protein Shp [13]. This protein does not show any significant homology to the cell surface heme-binding proteins of Staphylococcus aureus [14]. The shp and htsABC genes, which encode heme-specific ABC transporter, are co-transcribed with the 6 other genes and regulated by MtsR in response to iron levels [11, 17, 18]. We recently demonstrated that Shp rapidly transfers its heme to apoHtsA by a direct and affinity-driven mechanism [15, 16].
Streptococcus equi subspecies equi, a member of Gram-positive group C streptococci, causes strangles in horses, one of the most common infectious diseases of equine [19]. A metal binding protein (MBL) has been detected in S. equi [20], which is also a homologue of MtsA of Streptococcus pyogenes manganese and iron transporter MtsABC [21]. However, very little is known about iron acquisition in S. equi. S. equi has the homologue of Shp and HtsABC (designated SeShp and SeHtsABC, respectively. For simplicity, Shp and HtsA refer to the S. pyogenes proteins in this report.). As an initial effort to understand the iron acquisition processes in S. equi, we prepared and characterized SeShp and SeHtsA. The results suggest that the machinery and mechanism of heme acquisition are conserved in S. equi and S. pyogenes.
Results
S. equi shp and htsA genes
A blast search of available S. equi genome [22] was conducted to find out whether S. equi has a heme acquisition system homologous to the one in S. pyogenes. S. equi homologues of shp and htsABC are present. SeShp and Shp encoded by spy1796 are highly conserved in the N-terminal half of the mature form, sharing 75% identity and 91% similarity in amino acid sequence in the region of amino acid residues 28 through 173 amino acids. In contrast, the secretion signal sequences and the C-terminal half share less than 25% identity (Fig. 1A), suggesting that the N-terminal half of SeShp and Shp is important to their function. Unlike SeShp, SeHtsA shares with HtsA, which is encoded by spy1795, 76% identity in amino acid sequence throughout the whole proteins (Fig. 1B). The 5 downstream genes cotranscribed with shp and htsABC [13] also have their homologues next to S. equi shp and htsABC, indicating that the locus is conserved in S. pyogenes and S. equi.
Recombinant SeShp and SeHtsA proteins
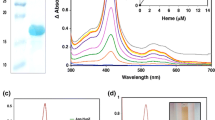
The existence of the shp and htsABC homologues in S. equi suggests that the pathogen may have a heme acquisition system similar to the one in S. pyogenes. If this is true, SeShp and SeHtsA should bind heme. To test this idea, recombinant mature SeShp, without the C-terminal transmembrane domain and charged tail, and SeHtsA, without lipid moiety, were prepared. Tag-free SeShp was overexpressed in E. coli, while tag-free SeHtsA was produced at a much lower level (Fig. 2). Fusion of the 6xHis tag to SeHtsA (SeHtsAHis) at the amino terminus greatly enhanced yield of SeHtsA. SeShp, SeHtsA, and SeHtsAHis were purified to more than 90% purity based on SDS-PAGE (Fig. 2).
SDS-PAGE analysis of recombinant holoSeShp, apoSeShp, SeHtsA, and SeHtsHis. SeShp lacks its secretion signal sequence and C-terminal transmembrane domain and charged tail, and SeHtsA and SeHtsHis lacks the secretion signal sequence and lipid moiety. The gel was stained with GelCode blue. Samples shown are E. coli with vector (control) (lane 1), E. coli lysate containing SeShp (2); SeHtsA (3), and SeHtsAHis (4), and purified holoSeShp (5), apoSeShp (6), SeHtsA (7), and SeHtsAHis (8).
Expression of SeShp, SeHtsA, or SeHtsAHis turns E. coli pellet red, the same phenomenon observed in the production of Shp and HtsA in E. coli, suggesting that SeShp and SeHtsA also bind heme (Fe(II)-protoporphyrin IX). Purification of SeShp results in two forms, one with red color and another without color. The red SeShp fraction has a spectrum with absorption peaks at 428, 530 and 560 nm in the visible region (Fig. 3A). This spectrum is identical to that of heme-containing Shp, suggesting that SeShp binds heme. A pyridine hemochrome assay was used to further determine whether the chromophore associated with SeShp is heme. The spectrum of SeShp in the assay is identical to that of the pyridine hemochrome derived from Shp, displaying peaks at 418, 524, and 556 nm (Fig. 3B), confirming that SeShp binds heme. Measurements of heme and protein contents found that each SeShp binds 0.9 heme in the red SeShp sample. These results indicate that SeShp binds heme in a 1:1 stoichiometry.
SeShp and SeHtsA bind heme. (A) UV-visible absorption spectra of 9 μM SeShp (dashed line) and SeHtsA (solid line) in 20 mM Tris-HCl, pH 8.0. (B) Spectra of reduced pyridine hemochrome derived from Shp (positive control) (solid line), SeShp (open circles), and SeHtsA (crosses). The spectra were normalized for comparison.
Both SeHtsA and SeHtsAHis purified are red, and the spectrum of SeHtsA shows an absorption peak at 410 nm (Fig. 3A), which is similar to the spectrum of hemiHtsA. HemiHtsA binds the hexacoordinate hemin (Fe(III)-protoporphyrin IX), suggesting that SeHtsA also bind hemin. The spectrum of reduced pyridine hemochrome derived from SeHtsA was identical to that derived from Shp, confirming that SeHtsA indeed binds hemin (Fig. 3B). Measurements of hemin and protein contents found that purified SeHtsA binds 0.97 heme per SeHtsA molecule, indicating a 1:1 stoichiometry for hemin binding. SeHtsA displays a much higher ratio of the Soret absorption peak over the absorbance at 280 nm (Fig. 3A) than SeHtsAHis (Fig. 4B) at a same protein concentration, indicating that not all SeHtsHis had bound hemin. Hemin- and protein-content measurements found that the molar ratio of bound hemin over SeHtsAHis protein was 0.18. These results indicate that 82% of SeHtsAHis was in a heme-free or apo-protein form.
Reconstitution of holoSeShp (A) and holoSeHtsAHis (B) by hemin. Purified apoSeShp and 82% apoSeHtsHis were incubated with hemin and separated from free hemin by gel filtration using a Sephadex G-25 column. Displayed are the optical absorption spectra of the untreated (dashed curves) and treated (solid curves) protein samples. Concentrations of SeShp and SeHtsAHis were 10 μM and 13 μM, respectively.
Reconstituted SeShp- and SeHtsAHis-hemin complexes
If SeShp and SeHtsA bind heme and hemin, holoSeShp and holoSeHtsA should be reconstituted from apo-proteins and hemin. ApoSeShp and SeHtsAHis containing 82% apo-form were incubated with hemin, and passed through a G-25 column to separate proteins from free hemin. ApoSeShp does not have absorption in the visible region, while the treated SeShp has an absorption peak at 418 nm, the Soret absorption peak of bound hemin (Fig. 4A). Measurements of protein and hemin contents found that the treated SeShp bound 1.08 hemin per protein molecule. The absorbance of reconstituted SeHtsAHis at 408 nm was 4.2 times as that of untreated SeHtsAHis at the same protein concentration (Fig. 4B). As assessed by the pyridine hemochrome and protein concentration assays, reconstituted SeHtsAHis bound 1.01 hemin per protein molecule. These results indicate that hemiSeShp and hemiSeHtsAHis are reconstituted from hemin and apo-proteins, demonstrating that both proteins bind hemin in a 1:1 stoichiometry.
Oxidation states of protoporphyrin Fe in SeShp and SeHtsA
The spectrum of purified holoSeShp has a dominant α peak at 560 nm and resembles the spectra of reduced b-type cytochrome with a 6-coordinate heme iron and hemochrome, indicating that purified holoSeShp is in the hexacoordinate heme or hemochrome form (designated hemoSeShp). In contrast, holoSeShp reconstituted from apoSeShp and hemin shows a typical spectrum of hemichrome or oxidized b-type cytochrome with a 6-coordiate hemin iron, implying that the reconstituted holoSeShp is in the hemichrome form (designated hemiSeShp). These results indicate that purified holoSeShp is in the reduced state, which is resistant to autoxidation. The following results further confirm this conclusion. The treatment of hemoSeShp with ferricyanide results in the hemichrome visible spectrum, and hemoSeShp is formed upon the addition of dithionite to hemiSeShp and remains in the hemochrome form after the removal of excess dithionite (Fig. 5A). The results demonstrate that the ferric to ferrous and ferrous to ferric transition can be readily achieved by the treatments with the reducing and oxidizing agents, respectively.
Conversion between and stability of oxidation states of SeShp and SeHtsAHis. (A) Optical absorption spectra of 6 μM hemiSeShp from the oxidation of hemoSeShp with ferricyanide (dashed curve) and hemoSeShp obtained from the reduction of hemiSeShp with dithionite (solid curve). The hemoSeShp spectrum was recorded after the removal of excess dithionite. (B) Optical absorption spectra of 5 μM dithionite-treated holoSeHtsAHis in the presence of excess dithionite (dashed line) and after the removal of dithionite (solid line).
Both purified SeHtsA and reconstituted SeHtsAHis are in the hemichrome form. The addition of excess dithionite to hemiSeHtsHis instantly leads to the formation of hemoSeHtsAHis, as evidenced by the shift of the Soret peak from 410 to 424 nm and the replacement of the absorption peak of hemichrome at 530 nm with the hemochrome specific peaks at 526 and 558 nm (Fig. 5B). However, the removal of excess dithionite by gel filtration results in only hemiSeHtsAHis, indicating that hemoSeHtsA is oxidized by O2.
Heme transfer from hemoSeShp to apoSeHtsAHis
The spectral changes are associated with the heme transfer from Shp to apoHtsA and the subsequent autoxidation of the resulting hemoHtsA [15, 16]. The changes have been used to examine the mechanism of the heme transfer. To determine whether hemoSeShp also transfer its heme to apoSeHtsAHis, the spectrum of a hemoSeShp/apoSeHtsA mixture was recorded with time after mixing. Upon mixing, the Soret absorption peak instantly shifts from 428 to 424 nm. Following the rapid initial shift, a slower spectral change occurs during which the Soret absorption peak gradually shifts towards 410 nm, and, at the same time, the 524 nm and 556 nm peaks decrease (Fig. 6A). Similar rapid and slow spectral changes are observed after mixing hemoShp and apoHtsA and are due to the formation and subsequent autoxidation of hemoHtsA [16]. If the slower change in Fig. 6A is due to the autoxidation of hemoSeHtsAHis, mixing the proteins under anaerobic conditions should lack the slow spectral change. These were what happened when hemoSeShp and apoSeHtsAHis were mixed in the presence of excess sodium dithionite (Fig. 6B). The data indicate that the heme transfer from hemoSeShp to apoSeHtsAHis follows the same mechanism of the heme transfer from hemoShp to apoHtsA.
Spectral evidence for heme transfer from hemoSeShp to apoSeHtsAHis. (A) The shift over time in the visible spectrum of a mixture containing 4 μM hemoSeShp and 6 μM SeHtsAHis. The spectra (solid curves) were taken at 0, 90, 165, 315, 540, 990 seconds after mixing. The dashed line is the sum of the spectra of 4 μM hemoSeShp and 6 μM SeHtsAHis. (B) The spectral change in the visible region after mixing 4 μM hemoSeShp and 6 μM SeHtsAHis with excess dithionite. The spectra were recorded immediately (solid curve) and 6 min after mixing (dotted grey curve). The spectrum of hemoSeShp is included as a reference for change (dashed curve). The spectra were normalized for comparison. Insert: the same spectra in the region from 510 to 590 nm. The arrows indicate the directions of the spectral changes with time.
Heme transfer from hemoSeShp to apoHtsA and from hemoShp to apoSeHtsA
If the structures, functions, and functional mechanism of the heme acquisition machinery are well conserved in S. equi and S. pyogenes, hemoSeShp and hemoShp should be able to transfer their heme to apoHtsA and apoSeHtsA, respectively. To test this idea, the heme transfer experiment was performed for the combinations of hemoSeShp/apoHtsA and hemoShp/apoSeHtsA. In both cases, the rapid shift in the Soret absorption peak and the subsequent slow change are observed (Fig. 7), indicating that hemoSeShp and hemoShp each can transfer heme to both apoSeHtsA and apoHtsA.
Heme transfer from hemoSeShp to apoHtsA and from hemoShp to apoSeHtsAHis. Presented are the shift in the visible spectra of the mixtures of 4 μM hemoShp/6 μM apoSeHtsAHis (A) and 4 μM hemoSeShp/6 μM apoHtsA (B) immediately and 90, 165, 315, 540, and 990 seconds after mixing. The dashed line in each panel is the sum of the spectra of the two individual proteins at the same concentrations. The arrows indicate the directions of the spectral changes with time.
Discussion
In this study, we have identified the putative cell surface protein SeShp and lipoprotein SeHtsA as heme binding proteins. We have demonstrated that hemoSeShp can rapidly transfer its heme to apoSeHtsA. The results suggest that the structures, functions, and functional mechanisms of the heme acquisition machinery in S. equi and S. pyogenes are conserved.
Three evidences support the conclusion that SeShp and SeHtsA bind heme and hemin. First, the purified holoSeShp and holoSeHtsA possess the absorption spectra similar to those of the reduced and oxidized b-type cytochrome or the hemochrome and hemichrome, respectively [23, 24]. Secondly, pyridine hemochrome analysis of SeShp and SeHtsA results in reduced pyridine hemochrome. Thirdly, the hemichrome complexes are formed from the reaction of apoSeShp or apoSeHtsAHis and hemin. It is interesting that the purified holoSeShp shows the dominant α absorbance band at 560 nm, which is only seen for reduced b-type cytochrome and hemochrome. The treatment of hemoSeShp with ferricyanide converts it into hemiSeShp, and reduction of hemiSeShp by dithionite turns it back to hemoSeShp. Thus, hemoSeShp is unusually stable in air, though the basis for the stability is unknown. In contrast to hemoSeShp, hemoSeHtsA is readily oxidized by oxygen.
The initial rapid and subsequent slower spectral changes upon mixing hemoShp and apoHtsA have been shown to be associated with the formation of hemoHtsA and its autoxidation, respectively [16]. The spectral changes observed after mixing hemoSeShp and apoSeHtsA are very similar to those that occurred when mixing hemoShp and apoHtsA, indicating that hemoSeShp also rapidly transfers its heme to apoSeHtsA and the resulting hemoSeHtsA autoxidizes. This interpretation is supported by the fact that the slower spectral change in the reaction of hemoSeShp and apoSeHtsAHis is absent when dithionite is present.
Our data suggest that S. equi and S. pyogenes use similar machinery to acquire heme. SeShp and Shp or SeHtsA and HtsA share high homology in amino acid sequence. These homologous proteins have similar absorption spectra associated with bound heme and hemin and similar stability in oxidation states in air. More importantly, like Shp, SeShp can rapidly transfer its heme to apoHtsA. These data suggest that the heme acquisition machinery and mechanism in both pathogens are conserved. This assertion is further supported by the fact that heme can be transferred from hemoSeShp to apoHtsA and from hemoShp to apoSeHtsA.
The high homology between SeShp and Shp is located in the N-terminal half of the proteins. We propose that the N-terminal half is involved in heme binding and transfer while the C-terminal half functions as a spacer to place the N-terminal part beyond the cell wall. Blast of GenBank does not reveal other proteins homologous to SeShp and Shp, suggesting that SeShp and Shp contain a novel heme-binding pocket, which renders the stability of hemochrome in air. In contrast, SeHtsA also share homology with FatB encoded by the genomes of Listeria monocytogenes, Staphylococcus aureus, Bacillus anthracis, and Clostridium perfringens. FatB is annotated as a putative ferric ferrichrome binding protein of ABC transporters. The fact that SeHtsA and HtsA bind heme suggests that these SeHtsA homologues may be heme-binding proteins and part of the transporters for heme acquisition.
Methods
Materials and strain
Sephadex G-25, SP sepharose and DEAE sepharose were purchased from Amersham (Piscataway, NJ). Nickel-nitrilotriacetic acid agarose (Ni-NTA) was obtained from QIAGEN (Valencia, CA). Bovine hemin chloride was from Sigma (St. Louis, MO). S. equi strain SEM1 was isolated from a horse suffering strangles in Montana, USA in 2003.
Sequence alignment
Blast search of the Streptococcus equi genome [25] and GenBank with the Shp and HtsA amino acid sequences were carried out to identify their homologues. Sequence alignment was performed using multiple sequence alignment by CLUSTALW under default parameters [26].
Gene cloning
S. equi shp and htsA were cloned from strain S. equi strain SEM1 using primer pairs 5'-AccatggATAAGGGTGAGCTTTATTCC-3'/5'-CgaattcTAATTATTTACTGGCTTTTGGGAC-3' and 5'-AccatggGTGTGGATCAGCACCCTAG-3'/5'-CgaattcTTACTTGACTTCCTTTGCTCC-3', respectively. The ccatgg and gaattc in lower case in the primers are introduced NcoI and EcoRI sites, respectively, and the sequences underlined correspond to the respective genes. The PCR products were digested with Nco I and Eco RI and ligated into pET-21d at the Nco I and Eco RI sites to yield pSEHTSA and pSESHP for S. equi htsA and shp genes, respectively. SeHtsA produced from pSEHTSA is tag-free mature protein with its first residue Cys20 replaced with Gly. SeShp produced from pSESHP is tag-free protein without the secretion signal sequence (residues 1–29) and lacks the transmembrane domain and charged tail (amino acids 260 to 293) located at the carboxyl terminus. The S. equi htsA PCR product was also cloned into pET-His2 [11] at the Nco I and Eco RI sites to yield pSEHTSA2, which encodes SeHtsAHis with a 6xHis tag (MHHHHHHLETMG) fused to the second amino acid residue of mature SeHtsA. The cloned genes have the identical sequences with those in the S. equi genome database [22].
Protein expression
SeShp, SeHtsA, and SeHtsAHis were expressed in Escherichia coli BL21(DE3) containing the corresponding plasmid. E. coli cells were grown at 37°C in 3 L Luria-Bertani broth supplemented with 100 mg of ampicillin/L. When the optical density at 600 nm reached approximately 1.0, the expression was induced with 0.5 mM isopropyl β-D-thiogalactopyranoside for 6 hours and harvested by centrifugation.
Protein purification
All solutions were buffered with 20 mM Tris-HCl, pH 8.0 (Tris-HCl). To prepare samples for column chromatography, bacterial pellets containing expressed protein were resuspended in 40 ml Tris-HCl, sonicated on ice for 15 min, and centrifuged at 20,000 × g for 10 min to remove cell debris. All dialysis was performed against 3 l Tris-HCl at 4°C for 12 hours with one buffer change. Purified proteins were concentrated using Centricon Plus 20 filtration devices (Millipore, Bedford, MA). Shp and HtsA of S. pyogenes were prepared as previously described [11, 13].
To purify holoSeShp, the lysate containing SeShp was loaded onto a DEAE column (2.5 × 6 cm), washed with 100 ml Tris-HCl, and bound SeShp was eluted with a 100-ml gradient of 0 – 0.3 M NaCl in Tris-HCl. Eluted fractions were analyzed using SDS-PAGE and those containing holoSeShp and apoSeShp were separately pooled and dialyzed. The holoSeShp pool was loaded onto an SP sepharose column (1.5 × 20 cm) and washed with 80 ml Tris-HCl, and the column was eluted with a 100-ml gradient of 0–0.15 M NaCl in Tris-HCl. Fractions containing holoSeShp with >90% of purity were pooled, dialyzed, concentrated, and stored at -20°C prior to use. To prepare apoSeShp, the apoSeShp pool was loaded onto a Q sepharose column (1.5 × 20 cm), and the column was eluted with a 0–0.3 M gradient of NaCl. The fractions containing SeShp were pooled, adjusted to 1.0 M (NH4)2SO4, and loaded onto a 1.5 × 5.0-cm phenyl sepharose column. The column was washed with 50 ml 1.0 M (NH4)2SO4 and eluted with an 80-ml linear gradient of 1.0 to 0.25 M gradient of (NH4)2SO4.
To prepare SeHtsA, the lysate containing SeHtsA was loaded onto a 2.5 × 10-cm column of DEAE sepharose. The column was washed with Tris-HCl and eluted with a 150-ml linear gradient of 0–0.1 M NaCl. Fractions containing relatively pure SeHtsA were pooled, adjusted to 1.0 M (NH4)2SO4, and loaded onto a 1.5 × 6-cm column of phenyl sepharose. The column was washed with 50 ml 1.0 M (NH4)2SO4 and eluted with a 40-ml linear gradient of 1.0–0.6 M (NH4)2SO4. Fractions containing SeHtsA were pooled, dialyzed, and subjected to the DEAE Sepharose chromatography again as described above. Fractions containing SeHtsA with >90% purity were pooled, dialyzed, and concentrated using a Centricon Plus 20 filter device.
To purify SeHtsAHis, the lysate containing SeHtsAHis was loaded onto a Ni-NTA column (2.5 × 6 cm), and the column was washed with 100 ml Tris-HCl and eluted with 0.15 M imidazole in Tris-HCl. Fractions positive with SeHtsAHis, as assessed by SDS-PAGE, were pooled and dialyzed. The dialyzed sample was loaded onto a DEAE sepharose column (2.5 × 5 cm), and the column was washed with 80 ml Tris-HCl and eluted with 0.10 M NaCl Tris-HCl. Fractions containing SeHtsAHis were pooled, dialyzed, concentrated, and stored at -20°C prior to use.
Assays for protein and heme or hemin contents
Protein concentrations were determined using a modified Lowry protein assay kit (Pierce, Rockford, IL) with bovine serum albumin as a standard. Heme or hemin content was measured using a pyridine hemochrome assay as follows [27]. Protein sample was added in 750 μl Tris-HCl and mixed with 75 μl 1 N NaOH, 175 μl pyridine, and approximately 2 mg sodium dithionite. The optical absorption spectrum was immediately recorded. The heme or hemin content was determined using the extinction coefficient ε418 = 191.5 mM-1 cm-1.
Reconstitution of hemiSeShp and hemiSeHtsAHis
HemiSeShp and hemiSeHtsAHis were reconstituted from apoSeShp and 82% apoSeHtsHis, respectively, with bovine hemin chloride as described previously [11]. Briefly, each protein was incubated at room temperature with hemin in 1.0 ml Tris-HCl at a protein:hemin ratio of 1:2 for 15 minutes. The sample was then loaded onto a Sephadex G-25 column (1.5 × 20 cm), and the holoprotein was eluted with Tris-HCl.
Reduction and oxidation of holoSeShp and holoSeHtsAHis
Reduction of hemiSeShp and hemiSeHtsAHis was achieved by adding approximately 1 mg sodium dithionite to 200 μl protein. In the specified cases, excess dithionite was removed from the proteins by passing the samples through a Sephadex G-25 column (1.5 × 20 cm). Oxidation of hemoSeShp was achieved by the reaction with ferricyanide. Ferricyanide (1 mM) was added to 200 μl hemoSeShp, and hemiSeShp formed was separated from excess ferricyanide using a Sephadex G-25 column (1.5 × 20 cm).
Heme transfer
Transfer of heme from hemoSeShp or hemoShp to SeHtsAHis (82% apo form and 18% holo form) or apoHtsA was followed by monitoring the spectral changes, as described previously [16]. Briefly, the holo- and apo-proteins were mixed at the indicated concentrations in 100 μl Tris-HCl at 25°C, and the optical absorption spectra of the samples were recorded at indicated times. Heme transfer under reduced conditions was achieved by addition of excess sodium dithionite to both apo and holo proteins prior to mixing. All absorption spectra were obtained using a SPECTRAmax 384 Plus spectrophotometer (Molecular Devices, Sunnyvale, CA).
Nucleotide sequence accession numbers
The GenBank accession numbers for the nucleotide sequences of the Streptococcus equi shp and htsA genes are DQ975461 and DQ975462, respectively.
References
Wooldridge KG, Williams PH: Iron uptake mechanisms of pathogenic bacteria. FEMS Microbiol Rev. 1993, 12: 325-348. 10.1111/j.1574-6976.1993.tb00026.x.
Weinberg ED: Iron and Infection. Microbiol Rev. 1978, 42: 45-66.
Neilands JB: Microbial iron compounds. Ann Rev Biochem. 1981, 50: 715-731. 10.1146/annurev.bi.50.070181.003435.
Rouault TA: Pathogenic bacteria prefer heme. Science. 2004, 305: 1577-1578. 10.1126/science.1102975.
Heinrichs DE, Rahn A, Dale SE, Sebulsky MT: Staphylococcus, Streptococcus, and Bacillus. Iron Transport in Bacteria. Edited by: Crosa JH, Mey AR, Payne SM. 2004, Washington DC: ASM Press, 394-296.
Hanks TS, Liu M, McClure MJ, Lei B: ABC transporter FtsABCD of Streptococcus pyogenes mediates uptake of ferric ferrichrome. BMC Microbiol. 2005, 5: 62-10.1186/1471-2180-5-62.
Eichenbaum Z, Muller E, Morse SA, Scott JR: Acquisition of iron from host proteins by the group A Streptococcus. Infect Immun. 1996, 64: 5428-5429.
Debarbieux L, Wandersman C: Hemophore-dependent heme acquisition systems. Iron Transport in Bacteria. Edited by: Crosa JH, Mey AR, Payne SM. 2004, Washington DC: ASM Press, 38-47.
Klebba PE, Rutz JM, Liu J, Murphy CK: Mechanisms of TonB-catalyzed iron transport through the enteric bacterial cell envelope. J Bioenerget Biomemb. 1993, 25: 603-611.
Drazek ES, Hammack CA, Schmitt MP: Corynebacterium diphtheriae genes required for acquisition of iron from haemin and haemoglobin are homologous to ABC haemin transporters. Mol Microbiol. 2000, 36: 68-84. 10.1046/j.1365-2958.2000.01818.x.
Lei B, Liu M, Voyich JM, Prater CI, Kala SV, Deleo FR, Musser JM: Identification and characterization of HtsA, a second heme-binding protein made by Streptococcus pyogenes. Infect Immun. 2003, 71: 5962-5969. 10.1128/IAI.71.10.5962-5969.2003.
Skaar EP, Humayun M, Bae T, DeBord KL, Schneewind O: Iron-source preference of Staphylococcus aureus infections. Science. 2004, 305: 1626-1628. 10.1126/science.1099930.
Lei B, Smoot LM, Menning HM, Voyich JM, Kala SV, DeLeo FR, Reid SD, Musser JM: Identification and characterization of a novel heme-associated cell surface protein made by Streptococcus pyogenes. Infect Immun. 2002, 70: 4494-4500. 10.1128/IAI.70.8.4494-4500.2002.
Mazmanian SK, Skaar EP, Gaspar AH, Humayun M, Gornicki P, Jelenska J, Joachmiak A, Missiakas DM, Schneewind O: Passage of heme-iron across the envelope of Staphylococcus aureus. Science. 2003, 299: 906-909. 10.1126/science.1081147.
Liu M, Lei B: Heme transfer from streptococcal cell surface protein Shp to HtsA of transporter HtsABC. Infect Immun. 2005, 73: 5086-5092. 10.1128/IAI.73.8.5086-5092.2005.
Nygaard TK, Blouin GC, Liu M, Fukumura M, Olson JS, Fabian M, Dooley DM, Lei B: The mechanism of direct heme transfer from the Streptococcal cell surface protein Shp to HtsA of the HtsABC transporter. J Biol Chem. 2006, 281: 20761-20771. 10.1074/jbc.M601832200.
Bates CS, Toukoi C, Neely MN, Eichenbaum Z: Characterization of MtsR, a new metal regulator in group a Streptococcus, involved in iron acquisition and virulence. Infect Immun. 2005, 73: 5743-5753. 10.1128/IAI.73.9.5743-5753.2005.
Hanks TS, Liu M, McClure MJ, Fukumura M, Duffy A, Lei B: Differential Regulation of Iron/Manganese-Specific MtsABC and Heme-Specific HtsABC Transporters by Metalloregulator MtsR of Group A Streptococcus. Infect Immun. 2006, 74: 5132-5139. 10.1128/IAI.00176-06.
Timoney JF: The pathogenic equine streptococci. Vet Res. 2004, 35: 397-409. 10.1051/vetres:2004025.
Harrington DJ, Greated JS, Chanter N, Sutcliffe IC: Identification of Lipoprotein Homologues of Pneumococcal PsaA in the Equine Pathogens Streptococcus equi and Streptococcus zooepidemicus. Infect Immun. 2000, 68: 6048-6051. 10.1128/IAI.68.10.6048-6051.2000.
Janulczyk R, Ricci S, Bjorck L: MtsABC is important for manganese and iron transport, oxidative stress resistance, and virulence of Streptococcus pyogenes. Infect Immun. 2003, 71: 2656-2664. 10.1128/IAI.71.5.2656-2664.2003.
Website title.http://www.sanger.ac.uk/Projects/S_equi/
Trevaskis B, Watts RA, Andersson CR, Llewellyn DJ, Hargrove MS, Olson JS, Dennis ES, Peacock J: Two hemoglobin genes in Arabidopsis thaliana: the evolutionary origins of leghemoglobins. Proc Natl Acad Sci USA. 1997, 94: 12230-12234. 10.1073/pnas.94.22.12230.
Hargrove MS, Brucker EA, Stec B, Sarath G, Arrendondo-Peter R, Klucas RV, Olson JS, Philips GN: Crystal structure of a nonsymbiotic plant hemoglobin. Structure. 2000, 8: 1005-1014. 10.1016/S0969-2126(00)00194-5.
Website title.http://www.sanger.ac.uk/cgi-bin/blast/submitblast/s_equi
Website title.http://align.genome.jp
Fuhrhop JH, Smith KM: Laboratory methods. Porphyrins and Metalloporphyrins. Edited by: Smith KM. 1975, New York: Elsevier Publishing Co, 804-807.
Acknowledgements
This work was supported by grants 2004-35204-14637 from the NRI Competitive Grants Program of the U.S. Department of Agriculture, K22 AI057347 from National Institutes of Health, and P20 RR-020185 from the National Center for Research Resources and the Montana State University Agricultural Experimental Station.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
TKN purified hemoSeShp and SeHtsAHis, characterized the proteins, and drafted part of the manuscript. ML cloned the genes. MJM purified SeHtsA. BL designed the study and prepared the manuscript. All authors have read and approved the final manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
Open Access This article is published under license to BioMed Central Ltd. This is an Open Access article is distributed under the terms of the Creative Commons Attribution License ( https://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Nygaard, T.K., Liu, M., McClure, M.J. et al. Identification and characterization of the heme-binding proteins SeShp and SeHtsA of Streptococcus equi subspecies equi. BMC Microbiol 6, 82 (2006). https://doi.org/10.1186/1471-2180-6-82
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2180-6-82