Abstract
The zinc finger transcription factor GATA6 is believed to have important roles in the development of several organs including the liver, gastrointestinal tract and heart. However, analyses of the contribution of GATA6 toward organogenesis have been hampered because Gata6-/- mice fail to develop beyond gastrulation due to defects in extraembryonic endoderm function. We have therefore generated a mouse line harbouring a conditional loss-of-function allele of Gata6 using Cre/loxP technology.
LoxP elements were introduced into introns flanking exon 2 of the Gata6 gene by homologous recombination in ES cells. Mice containing this altered allele were bred to homozygosity and were found to be viable and fertile. To assess the functional integrity of the loxP sites and to confirm that we had generated a Gata6 loss-of-function allele, we bred Gata6 'floxed' mice to EIIa-Cre mice in which Cre is ubiquitously expressed, and to Villin-Cre mice that express Cre in the epithelial cells of the intestine. We conclude that we have generated a line of mice in which GATA6 activity can be ablated in a cell type specific manner by expression of Cre recombinase. This line of mice can be used to establish the role of GATA6 in regulating embryonic development and various aspects of mammalian physiology.
Similar content being viewed by others
Background
The mouse Gata6 gene encodes a 45 kD protein containing two highly conserved zinc-finger DNA binding domains with a Cys-X2-Cys-X17-Cys-X2-Cys motif that directs binding to the nucleotide sequence element (A/T)GATA(A/G) 1. GATA4, 5 and 6 make up a subset of GATA factors that have been implicated in the development of several organs including the heart, lung, gastrointestinal tract and liver [1–4]. Development of GATA6 null embryos arrests during gastrulation as a consequence of defects in extraembryonic endoderm function [2, 5]. This early embryonic lethality can be rescued by complementing the GATA6 null embryos with a wild type extraembryonic visceral endoderm using tetraploid embryo complementation, and embryos derived by this process can survive until E10.5 [3]. Analyses of such Gata6-/-ES cell-derived embryos has revealed defects in hepatogenesis, which supports the proposal that GATA6 is an important developmental regulator. Although the ability to generate embryos from Gata6-/-ES cells by tetraploid embryo complementation has provided important insight into the contribution of GATA6 during early embryogenesis, this approach is not compatible with studying the role of GATA6 at later stages in development or its role in controlling differentiation of specific cell types. In the current report, we describe the generation of mice containing a conditional null allele of Gata6 that can be used for cell type-specific removal of GATA6 by Cre-mediated recombination.
Results and discussion
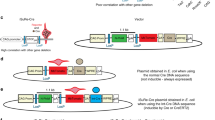
To ensure the elimination of GATA6 activity, we chose to flank Gata6 exon 2 (based on nomenclature described by Brewer et al [6, 7]) with loxP elements because this exon encodes the majority of the GATA6 protein (Fig. 1A). We decided to use a recombineering approach to facilitate the accurate placement of the loxP elements following the procedure described by Lee et al [8]. The final targeting vector contained a loxP element located between exons 2 and 3. In addition, a cassette containing a loxP element lying immediately 5' to a neomycin phosphotransferase (neo) gene, which conferred resistance to G418, was introduced between exons 1 and 2 (Fig. 1A). FRT sites flanked the neo gene, in order to allow removal of neo by Flp recombinase. Two novel restriction endonuclease cleavage sites (EcoRI and PacI) were also introduced into the Gata6 targeting vector to allow the identification of correctly modified Gata6 alleles by Southern blot analyses. The targeting vector was introduced into R1 ES cells [9] by electroporation, and G418-resistant ES cell clones were tested for homologous recombination at the Gata6 locus by Southern blot analysis. Fig. 1B shows an example of a correctly targeted ES (Gata6loxP(FRTneoFRT)/+) cell line. Following digestion of ES cell genomic DNA with EcoRI, a probe that lies 3' to the short arm of homology used in the targeting vector identified an expected 8.1 kb wild type Gata6 DNA fragment, while correctly targeted cells contain an additional 3.2 kb fragment due to the introduction of a new EcoRI site. When NdeI digested DNA was probed with a DNA fragment lying 5' to the expected position of the introduced loxP site, it identified the predicted 13.2 kb wild type fragment and a novel 15.2 kb fragment, which resulted from introducing the neo cassette into the Gata6 locus. Five correctly targeted ES cell clones were recovered and three of these cell lines were used to generate chimeric mice by morula aggregation [10]. These chimeric animals were then mated with CD-1 mice and successful germline transmission of the Gata6loxP(FRTneoFRT) allele was confirmed by Southern blot analysis of tail DNA (Fig. 1B). Mice carrying a single Gata6loxP(FRTneoFRT)/+ allele were viable and fertile. However, we were unable to obtain mice that were homozygous for this allele, which suggested that the inclusion of the neo cassette disrupted GATA6 function and resulted in embryonic lethality as observed for Gata6-/- embryos [2, 5]. We therefore established a mouse line, Gata6loxP/+, that lacked the neo cassette by inducing recombination between the FRT sites in vivo [11]. This was achieved by mating Gata6loxP(FRTneoFRT)/+ mice to a transgenic mouse, B6;SJL-Tg(ACTFLPe)9205Dym/J, in which Flp recombinase is widely expressed from the human beta actin gene promoter [11]. Correct excision of the neo cassette was confirmed in offspring by Southern blot and PCR analyses (Fig. 1B & C). Gata6loxP/+ mice were finally interbred to generate Gata6loxP/loxPmice, which are healthy, fertile and have reached maturity.
Generation of a Gata6 conditional null allele. (A) Schematic showing a map of the Gata6 genomic locus and the targeting vector with exons represented by open boxes. The relative position of Southern blot probes (lines), PCR primers (small arrowheads), loxP (large arrowheads) and FRT (ovals) sites, as well as cassettes encoding neomycin phosphotransferase (neo) and diphtheria toxin (DT), are included. Sizes of relevant EcoRI (e), NdeI (n), and PacI (p) restriction endonuclease fragments are shown in kilobase pairs (kb). (B) Southern blot analysis of genomic DNA isolated from ES cells (lanes 1, 2, 8 and 9) or mouse tails (lanes 3–7 and 10). An example of an ES cell line containing a correctly targeted Gata6loxP(FRTneoFRT)allele is shown in lanes 2 and 8. Mice harbouring the modified Gata6 allele were generated from these ES cells (lanes 3 and 10). Exon 2 of Gata6 was deleted (Gata6del) or the neo cassette alone was deleted, leaving Gata6 exon 2 flanked by loxP elements (Gata6loxP), by breeding Gata6loxP(FRTneoFRT)mice to transgenic mice expressing either Cre (lane 6 and 7) or Flp (lane 4) recombinases, respectively. The size of restriction fragments identified by 5' and 3' probes (Fig. 1A) was deduced from their position relative to standard DNA fragments. (C) The genotypes of mice and embryos were also determined by PCR amplification of genomic DNA. Primers were designed that differentiated between the Gata6+ (gt4F/4R; 159 bp), GataloxP(gt4F/4R; 250 bp) and Gata6del(k/o2F/2R; 568 bp) alleles, as well as neo (Neo A/B; 225 bp), flp (Flp F/R; 750 bp), and cre (Cre 1/2; 300 bp) transgenes.
We next addressed whether removal of GATA6 could be induced in vivo by expression of Cre recombinase. We chose to disrupt the Gata6 gene in the gastrointestinal tract because it is highly expressed in the epithelium and may have roles in controlling gut physiology or development. To produce Gata6loxP/+Villin-Cre mice, we bred Gata6loxP/loxPmice with transgenic mice (Tg(Vil-cre)997Gum) in which Cre expression was directed to the epithelial cells of the small intestine by the Villin (Vil1) promoter [12]. Using this transgenic strain of mice, Cre activity can be detected in the epithelium of the small intestine from E14.5 [12]. Gata6loxP/+Villin-Cre males were mated with Gata6loxP/loxPfemales to obtain Gata6loxP/+Villin-Cre control and Gata6loxP/loxPVillin-Cre experimental offspring. In previous experiments, we observed that the efficiency through which Cre mediates recombination between loxP sites varies between different transgenic male mice [13]. We noted a similar situation when using various Gata6loxP/+Villin-Cre males. In the majority of cases (4/5 stud males) we were unable to obtain Gata6loxP/loxPVillin-Cre offspring (n = 95 mice genotyped), and this lethality associated with presumptive loss of GATA6 in the intestine is currently under investigation. However, one Gata6loxP/+Villin-Cre male did produce offspring (12/90) whose genotype was determined to be Gata6loxP/loxPVillin-Cre by PCR analysis of tail DNA. We, therefore, used RT-PCR with Gata6 primers corresponding to nucleotide sequences predicted to be deleted after recombination between Gata6 loxP elements, to compare the steady-state levels of Gata6 mRNA in intestines isolated from control and experimental mice generated by this particular Gata6loxP/+Villin-Cre male. Fig. 2A shows that Gata6 mRNA could be detected in control intestine but not in intestine isolated from a subset of Gata6loxP/loxPVillin-Cre offspring. The remaining Gata6loxP/loxPVillin-Cre mice were found to continue to express Gata6 mRNA at varying levels (data not shown). We also examined expression of GATA6 protein in the intestines of Gata6loxP/loxPVillin-Cre offspring derived from the same Gata6loxP/+Villin-Cre male. Fig. 2B shows that GATA6 was detected as an abundant nuclear protein in the epithelial cells of control intestines. However, Gata6loxP/loxPVillin-Cre mice displayed a marked reduction in the abundance of intestinal GATA6 (Fig. 2B). From these results, we conclude that the Gata6 gene can be conditionally disrupted in Gata6loxP/loxPmice by cell-type specific expression of Cre recombinase.
Gata6 can be successfully ablated in the intestine of Gata6loxP/loxPmice by expression of Cre from the Villin promoter. (A) The steady-state level of Gata6 mRNA was compared in intestines isolated from four control Gata6loxP/+Villin-Cre (lanes 2–5) or experimental Gata6loxP/loxPVillin-Cre (lanes 6–9) mice. Amplification of Pol2 (Polr2a) mRNA showed that similar levels of starting material were utilized in each reaction. (B) Immunohistochemistry showing that GATA6 was detected as an abundant nuclear protein in epithelial cells (arrows) of control intestines, but was detected at greatly reduced levels in Gata6loxP/loxPVillin-Cre intestines.
Cre-mediated deletion of exon2 in Gata6loxP/loxPmice was predicted to result in loss of GATA6 function. If this were true, we expected that embryos homozygous for the deleted allele (Gata6del) should undergo developmental arrest, as was described for Gata6-/- embryos [2, 5]. Gata6del/+ mice were produced by mating Gata6loxP(FRTneoFRT)/+ mice with an EIIa-Cre transgenic mouse (B6.FVB-Tg(EIIa-cre)C5379Lmgd/J). The EIIa-Cre mouse expresses Cre recombinase in nearly all tissues, including those of pre-implantation embryos, and has been used previously to mediate recombination between loxP sites in germ cells [14]. Gata6+/delprogeny were identified by Southern blot analyses of NdeI digested tail DNA. As shown in Fig. 1B, Southern blots hybridized with the 5' Gata6 probe (Fig. 1A) revealed the conversion of a 15.2 kb NdeI fragment from the Gata6loxP(FRTneoFRT) allele to an expected 12 kb NdeI fragment from the Gata6delallele. Gata6+/delwere then mated inter se, and the genotype of embryos collected between E8.5 and E11.5 was determined by PCR analysis of genomic DNA. Of 77 embryos recovered, 27 were Gata6+/+ and 50 were Gata6+/del(Table 1). In addition, 23 partially resorbed empty decidual masses were recovered, which we believe likely resulted from the early developmental arrest of Gata6del/delembryos. We finally determined whether the developmental lethality associated with loss of GATA6 could be induced by expression of Cre in pre-implantation embryos. To achieve this, Gata6loxP/loxPmice were bred to Gata6+/delEIIa-Cre mice, which are heterozygous for the EIIa-Cre transgene. The genotype of resulting embryos ranging from E8.5 to E11.5 was again determined by PCR. While we recovered 14 Gata6loxP/+, 18 Gata6loxP/del, and 16 Gata6loxP/+EIIaCre embryos, no Gata6loxP/delEIIaCre embryos were identified although 13 empty resorbing decidual masses were observed (Table 1).
Conclusion
In summary, we conclude that we have generated a line of mice in which GATA6 transcriptional activity can be ablated by expression of Cre. We believe that the availability of this mouse will be useful to elucidate the contribution of GATA6 during organogenesis as well as its physiological role in adult mice.
Methods
Plasmid construction
A bacterial artificial chromosome (BAC # RP23-410I12) that contained Gata6 genomic DNA was obtained from BACPAC Resources, Children Hospital Research Institute, Oakland, CA. Plasmid pNeb-DT-GATA6 was generated by cloning a 12.2 kb EcoRV/PacI genomic fragment containing Gata6 exons 1 and 2 into the PacI/HincII site of a vector, pNEB193-DT, which contained a diphtheria toxin (DT) expression cassette to enrich against random integration of the targeting plasmid in ES cells. A cassette containing loxP-neo-loxP was amplified by PCR from the plasmid pSV-LNL (modified from Zhang et al)[15] using the following primers: Gata6ET1: GCTTGCTGTTTGAGTCTACCCCATTTCTGCCTGTTTCTTGACATCCCTTCGAATTCTGGTACCGGCGCGCCTAGTCGAC, Gata6ET2: ATCCATTATTGTCAATGTCTAAAGATGGAATTGTCTCTGCACAAGCTATCTTCTCAACTCGAGCCCTTAATTAACCGGT. These oligonucleotides contained 55 bp of sequence from Gata6 intron 2 (underlined). This amplicon was introduced into Gata6 intron 2 sequence in pNEB-DT-GATA6 by homologous recombination in E.coli following the procedure described by Lee et al [8] The loxPneoloxP cassette was then converted to a single loxP site by expression of Cre recombinase [8]. A loxP(FRTneoFRT) cassette was then amplified from pSV-LFNF using primers Gata6ET3: CACGCTGGTGGTTGTAAGGCGGTTTGTGTTTAAGGTGTGCGGTTGGCCTGGACGTGTGGTACCGGCGCGCCTAGTCGAC, Gata6ET4: GAAAAAGTTACCTAGCCCAGAGAAAGTGAGATGCCAGGAAAGGCATAAGGATATCAACTCGAGCCCTTAATTAACCGGT. These oligonucleotides contain 56 bp (Gata6ET3) and 52 bp (Gata6ET4) of sequence from Gata6 intron 1, respectively. This cassette was introduced into Gata6 intron 1, again using homologous recombination in E.coli. to generate the final targeting vector (Fig. 1A).
ES cell targeting and animals
Linear targeting vector (100μg) was introduced into R1 ES cells by electroporation, and the genotype of colonies resistant to 350μg/ml of Geneticin (Gibco BRL) was determined by Southern blot (Fig. 1B). Chimeric mice were generated by aggregation of ES cells with CD-1 morulae as described previously [10] and the modified allele was passed through the germline by breeding chimeras to CD1 mice. Gata6loxP/loxPmice were produced by breeding Gata6loxp(FRTneoFRT)/+ mice to B6;SJL-Tg(ACTFLPe)9205Dym/J mice [16] (Jackson Labs) to delete the FRTneoFRT cassette by Flp-mediated recombination in the germline. The ACTFLPe transgene was removed by breeding F1 Gata6loxP/loxPmice into CD-1 mice. Gata6+/delmice were generated by mating Gata6loxp(FRTneoFRT)/+ animals with B6.FVB-Tg(EIIa-cre)C5379Lmgd/J transgenic mice [17] (Jackson Labs) to allow Cre-mediate recombination between loxP elements in the germline. The EIIa-Cre transgene was removed by breeding Gata6+/delF1 mice with CD1 mice. The MCW IACUC committee approved all procedures using animals.
Southern blot, PCR and RT-PCR
Southern blot analyses were performed using standard conditions with probes indicated in Fig. 1. Genotypes were determined by PCR using the following oligonucleotide primer pairs: Gata6 gt4F/4R, GTGGTTGTAAGGCGGTTTGT, ACGCGAGCTCCAGAAAAAGT; Gata6 k/o2F/2R, AGTCTCCCTGTCATTCTTCCTGCTC, TGATCAAACCTGGGTCTACACTCCTA; Flp F/R, GGTCCAACTGCAGCCCAAGCTTCC, GTGGATCGATCCTACCCCTTGCG [16]; Cre 1/2, GTTCGCAAGAACCTGATGGACA, CTAGAGCCTGTTTTGCACGTTC [18]; Neo A/B, GCCAACGCTATGTCCTGATAGCGGT, AGCCGGTCTTGTCGATCAGGATGAT. RT-PCR was performed as described previously [19] with the following primer pairs: Gata6 9/10; AGTTTTCCGGCAGAGCAGTA, AGTCAAGGCCATCCACTGTC, Pol2 F/R; CTGATGCGGGTGCTGAGTGAGAAGG, GCGGTTGACCCCATGACGAGTG.
Immunohistochemistry
Immunohistochemistry was performed using antigen retrieval in citrate buffer as described previously [13] using an anti-GATA6 antibody (AF1700 R&D Systems,1/1000 dilution).
Abbreviations
- Neo:
-
neomycin phosphotransferase
References
Morrisey EE, Ip HS, Lu MM, Parmacek MS: GATA-6: a zinc finger transcription factor that is expressed in multiple cell lineages derived from lateral mesoderm. Dev Biol. 1996, 177: 309-322. 10.1006/dbio.1996.0165.
Morrisey EE, Tang Z, Sigrist K, Lu MM, Jiang F, Ip HS, Parmacek MS: GATA6 regulates HNF4 and is required for differentiation of visceral endoderm in the mouse embryo. Genes Dev. 1998, 12: 3579-3590.
Zhao R, Watt AJ, Li J, Luebke-Wheeler J, Morrisey EE, Duncan SA: GATA6 is essential for embryonic development of the liver but dispensable for early heart formation. Mol Cell Biol. 2005, 25: 2622-2631. 10.1128/MCB.25.7.2622-2631.2005.
Laverriere AC, MacNeill C, Mueller C, Poelmann RE, Burch JB, Evans T: GATA-4/5/6, a subfamily of three transcription factors transcribed in developing heart and gut. J Biol Chem. 1994, 269: 23177-23184.
Koutsourakis M, Langeveld A, Patient R, Beddington R, Grosveld F: The transcription factor GATA6 is essential for early extraembryonic development. Development. 1999, 126: 723-732.
Brewer A, Gove C, Davies A, McNulty C, Barrow D, Koutsourakis M, Farzaneh F, Pizzey J, Bomford A, Patient R: The human and mouse GATA-6 genes utilize two promoters and two initiation codons. J Biol Chem. 1999, 274: 38004-38016. 10.1074/jbc.274.53.38004.
Patient RK, McGhee JD: The GATA family (vertebrates and invertebrates). Curr Opin Genet Dev. 2002, 12: 416-422. 10.1016/S0959-437X(02)00319-2.
Lee EC, Yu D, Martinez de Velasco J, Tessarollo L, Swing DA, Court DL, Jenkins NA, Copeland NG: A highly efficient Escherichia coli-based chromosome engineering system adapted for recombinogenic targeting and subcloning of BAC DNA. Genomics. 2001, 73: 56-65. 10.1006/geno.2000.6451.
Nagy A, Rossant J, Nagy R, Abramow-Newerly W, Roder JC: Derivation of completely cell culture-derived mice from early-passage embryonic stem cells. Proc Natl Acad Sci U S A. 1993, 90: 8424-8428.
Wood SA, Allen ND, Rossant J, Auerbach A, Nagy A: Non-injection methods for the production of embryonic stem cell-embryo chimaeras. Nature. 1993, 365: 87-89. 10.1038/365087a0.
Rodriguez CI, Buchholz F, Galloway J, Sequerra R, Kasper J, Ayala R, Stewart AF, Dymecki SM: High-efficiency deleter mice show that FLPe is an alternative to Cre-loxP. Nat Genet. 2000, 25: 139-140. 10.1038/75973.
Madison BB, Dunbar L, Qiao XT, Braunstein K, Braunstein E, Gumucio DL: Cis elements of the villin gene control expression in restricted domains of the vertical (crypt) and horizontal (duodenum, cecum) axes of the intestine. J Biol Chem. 2002, 277: 33275-33283. 10.1074/jbc.M204935200.
Parviz F, Matullo C, Garrison WD, Savatski L, Adamson JW, Ning G, Kaestner KH, Rossi JM, Zaret KS, Duncan SA: Hepatocyte nuclear factor 4alpha controls the development of a hepatic epithelium and liver morphogenesis. Nat Genet. 2003, 34: 292-296. 10.1038/ng1175.
Lakso M, Pichel JG, Gorman JR, Sauer B, Okamoto Y, Lee E, Alt FW, Westphal H: Efficient in vivo manipulation of mouse genomic sequences at the zygote stage. Proc Natl Acad Sci U S A. 1996, 93: 5860-5865. 10.1073/pnas.93.12.5860.
Zhang Y, Buchholz F, Muyrers JP, Stewart AF: A new logic for DNA engineering using recombination in Escherichia coli. Nat Genet. 1998, 20: 123-128. 10.1038/2417.
Dymecki SM: Flp recombinase promotes site-specific DNA recombination in embryonic stem cells and transgenic mice. Proc Natl Acad Sci U S A. 1996, 93: 6191-6196. 10.1073/pnas.93.12.6191.
Holzenberger M, Lenzner C, Leneuve P, Zaoui R, Hamard G, Vaulont S, Bouc YL: Cre-mediated germline mosaicism: a method allowing rapid generation of several alleles of a target gene. Nucleic Acids Res. 2000, 28: E92-10.1093/nar/28.21.e92.
Chen J, Kubalak SW, Minamisawa S, Price RL, Becker KD, Hickey R, Ross JJ, Chien KR: Selective requirement of myosin light chain 2v in embryonic heart function. J Biol Chem. 1998, 273: 1252-1256. 10.1074/jbc.273.2.1252.
Li J, Ning G, Duncan SA: Mammalian hepatocyte differentiation requires the transcription factor HNF-4alpha. Genes Dev. 2000, 14: 464-474.
Acknowledgements
We would like to thank Drs. Robert Burgess and Francis Stewart for providing plasmids and Dr. Neal Copeland for providing bacterial strains used in recombineering. Pregnant mare serum gonadotrophin (PMSG) used in superovulation was obtained from Dr. A.F. Parlow at the National Hormone and Peptide Program (Torrance, CA). We are also grateful to Dr. Michele Battle for critically evaluating the manuscript and for providing plasmids. This work was supported by an NIH training grant from the MCW cardiovascular research centre to C.P.S and NIH grants to S.A.D.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
C.P.S. generated the targeting vector and carried out analyses of conditional knockout mice, as well as contributed to experimental design and draft of the manuscript. J. L. generated aggregation chimeras. S.A.D. conceived of the study, contributed to experimental design and interpretation of results, and coordinated the project and writing of the manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
Open Access This article is published under license to BioMed Central Ltd. This is an Open Access article is distributed under the terms of the Creative Commons Attribution License ( https://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Sodhi, C.P., Li, J. & Duncan, S.A. Generation of mice harbouring a conditional loss-of-function allele of Gata6. BMC Dev Biol 6, 19 (2006). https://doi.org/10.1186/1471-213X-6-19
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-213X-6-19